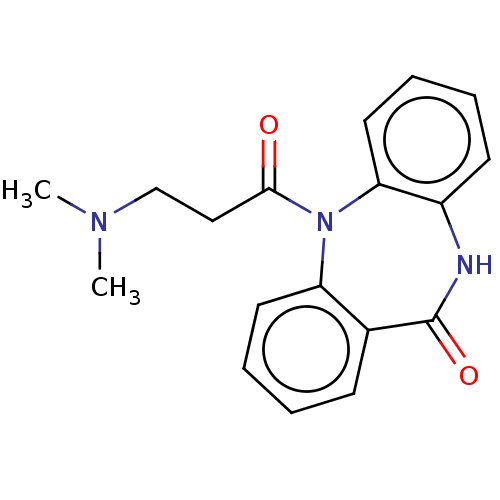
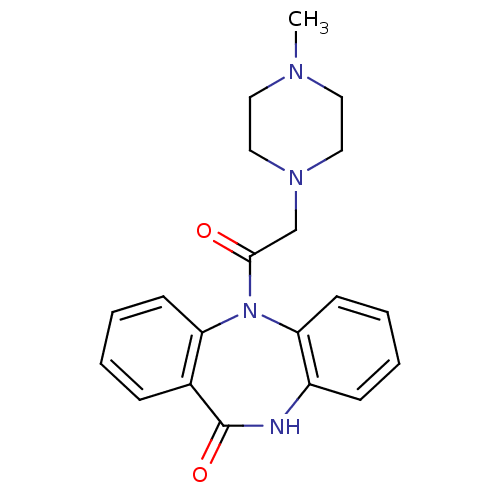
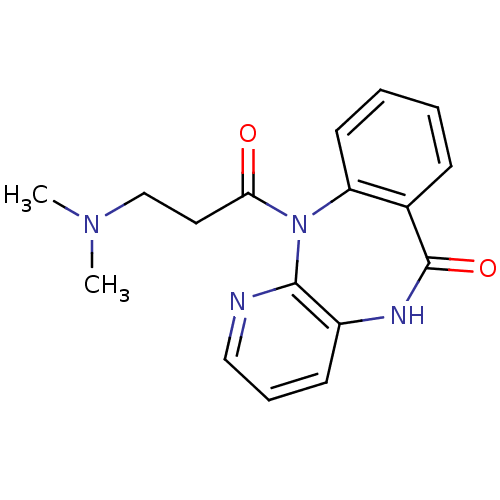
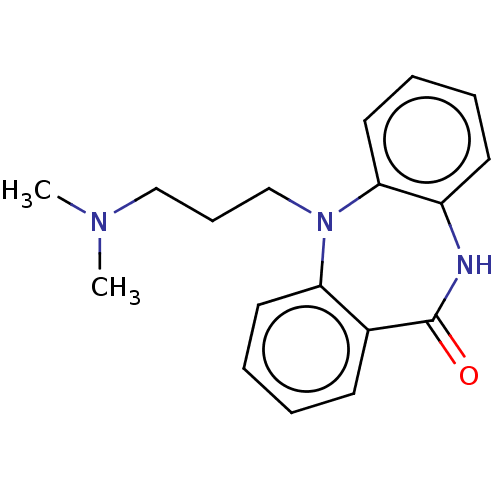
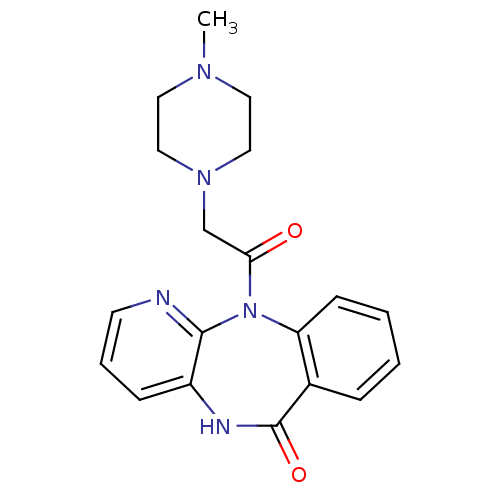
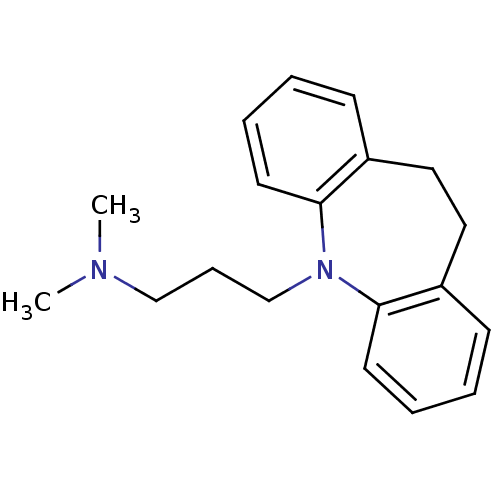
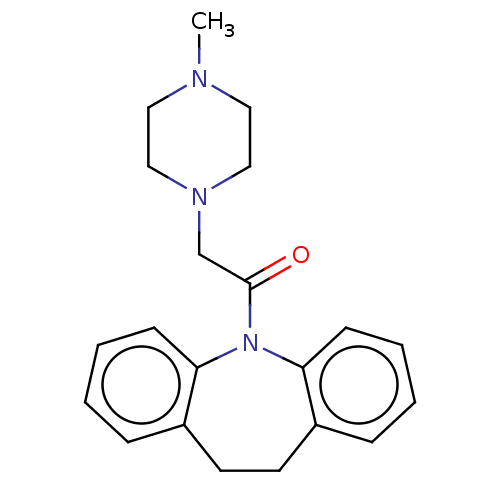
Report error Found 9 Enz. Inhib. hit(s) with all data for entry = 50048757
Affinity DataKi: 6nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor was determined in homogenized rat cortex tissue using [3H]N-methylscopolamine as radioliga...More data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor was determined in homogenized rat cortex tissue using [3H]N-methylscopolamine as radioliga...More data for this Ligand-Target Pair
Affinity DataKi: 45nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor was determined in homogenized rat cortex tissue using [3H]N-methylscopolamine as radioliga...More data for this Ligand-Target Pair
Affinity DataKi: 50nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor was determined in homogenized rat cortex tissue using [3H]N-methylscopolamine as radioliga...More data for this Ligand-Target Pair
Affinity DataKi: 60nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor was determined in homogenized rat cortex tissue using [3H]N-methylscopolamine as radioliga...More data for this Ligand-Target Pair
Affinity DataKi: 65nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor was determined in homogenized rat cortex tissue using [3H]N-methylscopolamine as radioliga...More data for this Ligand-Target Pair
Affinity DataKi: 200nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor was determined in homogenized rat cortex tissue using [3H]N-methylscopolamine as radioliga...More data for this Ligand-Target Pair
Affinity DataKi: 200nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor was determined in homogenized rat cortex tissue using [3H]N-methylscopolamine as radioliga...More data for this Ligand-Target Pair
Affinity DataKi: 240nMAssay Description:Binding affinity against Muscarinic acetylcholine receptor was determined in homogenized rat cortex tissue using [3H]N-methylscopolamine as radioliga...More data for this Ligand-Target Pair