Report error Found 23 Enz. Inhib. hit(s) with all data for entry = 50006965
TargetMuscarinic acetylcholine receptor M1(Human)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 1.10nMAssay Description:Agonist activity at human muscarinic M1 receptor expressed in HEK293T cells co-expressing Galpha subunit and PLC-beta3 by split luciferase complement...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Human)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 1.30nMAssay Description:Agonist activity at human muscarinic M1 receptor expressed in HEK293T cells co-expressing Galpha subunit and PLC-beta3 by split luciferase complement...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Human)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 5nMAssay Description:Agonist activity at human muscarinic M1 receptor expressed in HEK293T cells co-expressing Galpha subunit and PLC-beta3 by split luciferase complement...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Human)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 14nMAssay Description:Agonist activity at human muscarinic M1 receptor expressed in HEK293T cells co-expressing Galpha subunit and PLC-beta3 by split luciferase complement...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Human)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 15nMAssay Description:Agonist activity at human muscarinic M1 receptor expressed in HEK293T cells co-expressing Galpha subunit and PLC-beta3 by split luciferase complement...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Human)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 32nMAssay Description:Agonist activity at human muscarinic M1 receptor expressed in HEK293T cells co-expressing Galpha subunit and PLC-beta3 by split luciferase complement...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Human)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 63nMAssay Description:Agonist activity at human muscarinic M1 receptor expressed in HEK293T cells co-expressing Galpha subunit and PLC-beta3 by split luciferase complement...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Human)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 72nMAssay Description:Agonist activity at human muscarinic M1 receptor expressed in HEK293T cells co-expressing Galpha subunit and PLC-beta3 by split luciferase complement...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Human)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 129nMAssay Description:Agonist activity at human muscarinic M1 receptor expressed in HEK293T cells co-expressing Galpha subunit and PLC-beta3 by split luciferase complement...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Human)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 135nMAssay Description:Agonist activity at human muscarinic M1 receptor expressed in HEK293T cells co-expressing Galpha subunit and PLC-beta3 by split luciferase complement...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 151nMAssay Description:Displacement of [3H]-QNB from muscarinic receptor in rat brain membranes incubated for 45 mins under dark conditions by radioligand binding assayMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Human)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 170nMAssay Description:Agonist activity at human muscarinic M1 receptor expressed in HEK293T cells co-expressing Galpha subunit and PLC-beta3 by split luciferase complement...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Human)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 324nMAssay Description:Agonist activity at human muscarinic M1 receptor expressed in HEK293T cells co-expressing Galpha subunit and PLC-beta3 by split luciferase complement...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 363nMAssay Description:Displacement of [3H]-QNB from muscarinic receptor in rat brain membranes incubated for 45 mins under irradiation exposure at 365 nm by radioligand bi...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Human)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 457nMAssay Description:Agonist activity at human muscarinic M1 receptor expressed in HEK293T cells co-expressing Galpha subunit and PLC-beta3 by split luciferase complement...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 479nMAssay Description:Displacement of [3H]-QNB from muscarinic receptor in rat brain membranes incubated for 45 mins under dark conditions by radioligand binding assayMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 1.59E+3nMAssay Description:Displacement of [3H]-QNB from muscarinic receptor in rat brain membranes incubated for 45 mins under irradiation exposure at 365 nm by radioligand bi...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Human)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 2.09E+3nMAssay Description:Agonist activity at human muscarinic M1 receptor expressed in HEK293T cells co-expressing Galpha subunit and PLC-beta3 by split luciferase complement...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Human)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 3.24E+3nMAssay Description:Agonist activity at human muscarinic M1 receptor expressed in HEK293T cells co-expressing Galpha subunit and PLC-beta3 by split luciferase complement...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 3.98E+3nMAssay Description:Displacement of [3H]-QNB from muscarinic receptor in rat brain membranes incubated for 45 mins under irradiation exposure at 365 nm by radioligand bi...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 5.62E+3nMAssay Description:Displacement of [3H]-QNB from muscarinic receptor in rat brain membranes incubated for 45 mins under dark conditions by radioligand binding assayMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 7.08E+3nMAssay Description:Displacement of [3H]-QNB from muscarinic receptor in rat brain membranes incubated for 45 mins under dark conditions by radioligand binding assayMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1/M2/M3/M4/M5(Rat)
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Julius Maximilian University of W£Rzburg
Curated by ChEMBL
Affinity DataIC50: 7.41E+3nMAssay Description:Displacement of [3H]-QNB from muscarinic receptor in rat brain membranes incubated for 45 mins under irradiation exposure at 365 nm by radioligand bi...More data for this Ligand-Target Pair
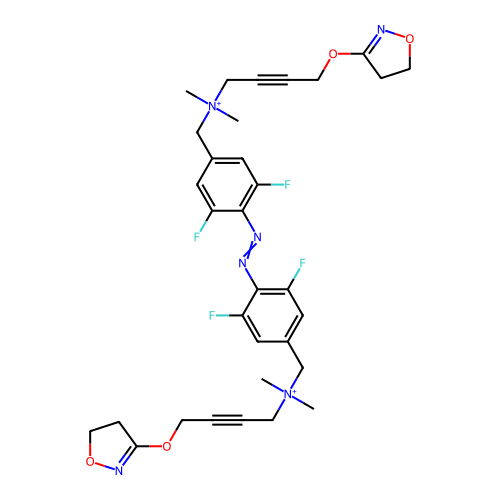
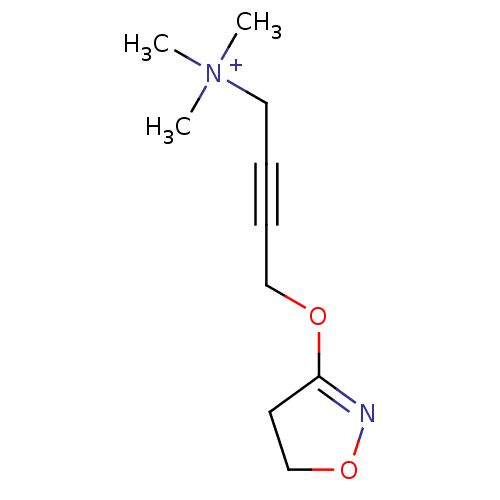
 3D Structure (crystal)
3D Structure (crystal)