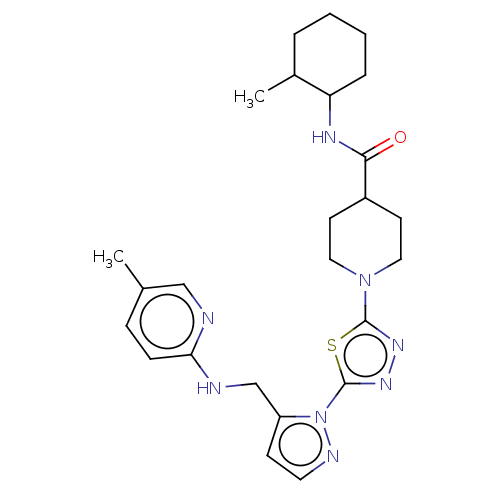
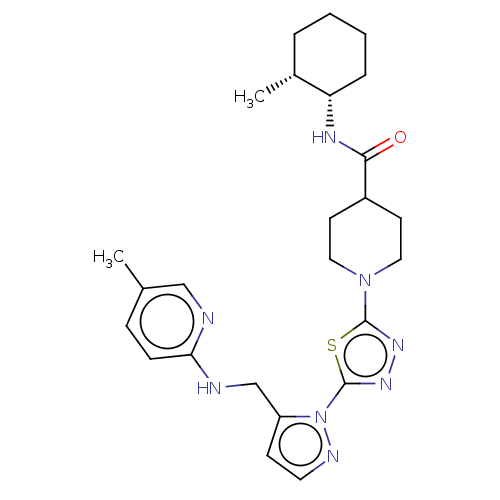
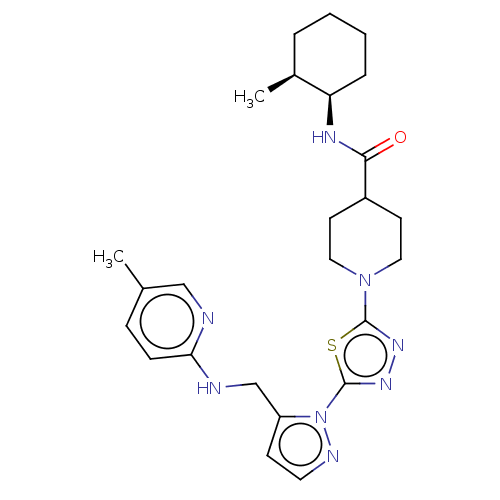
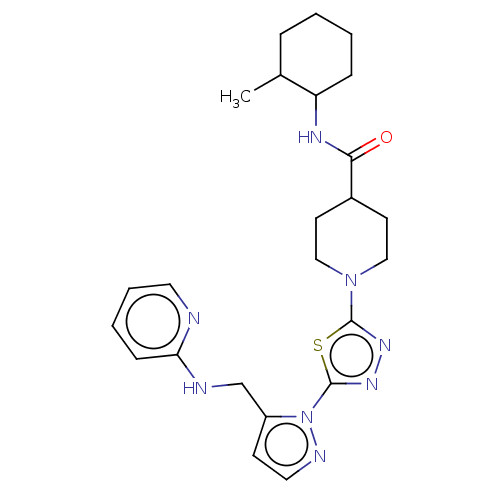
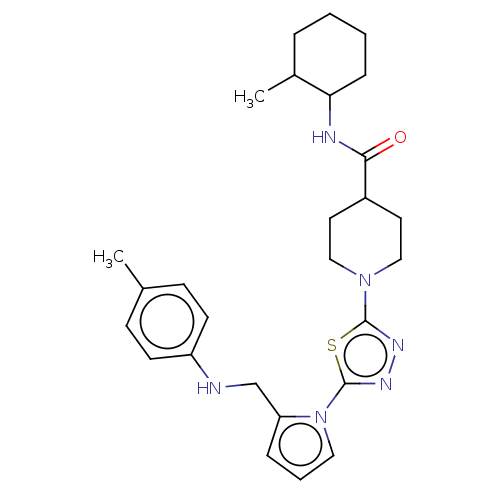
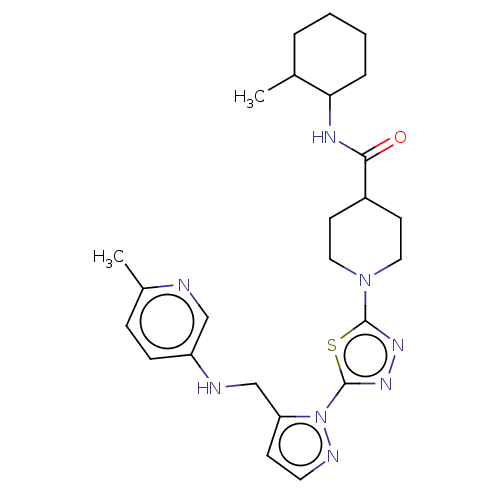
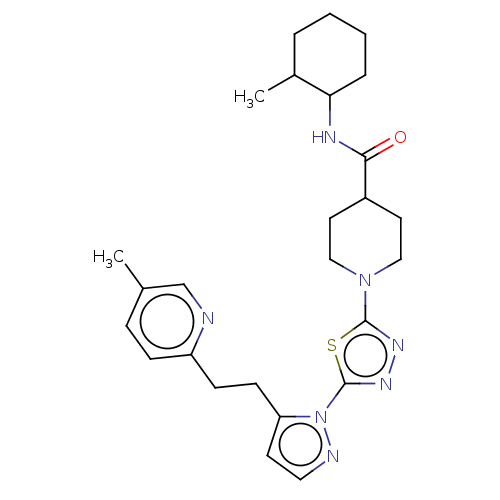
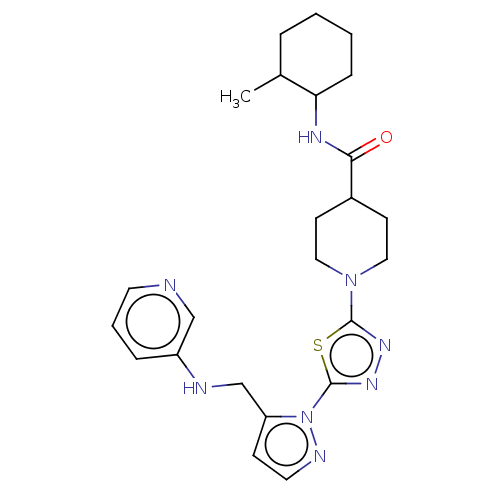
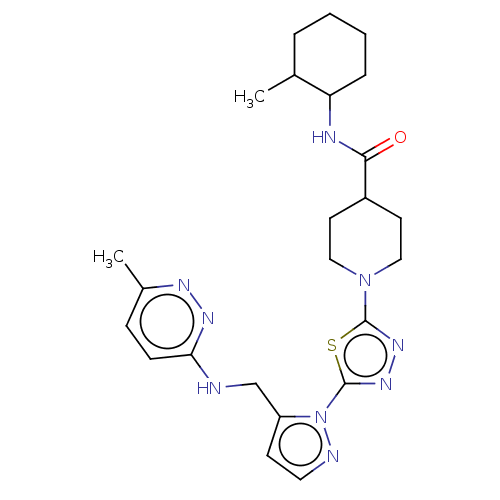
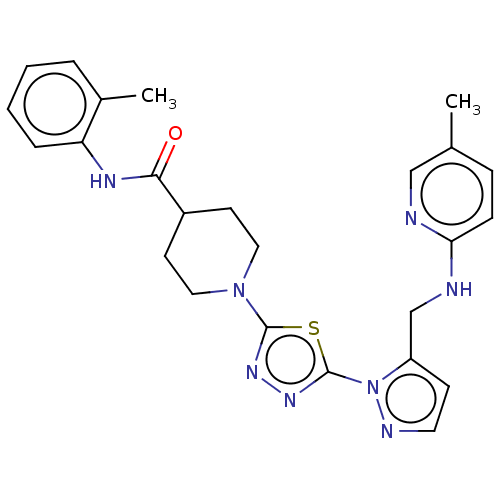
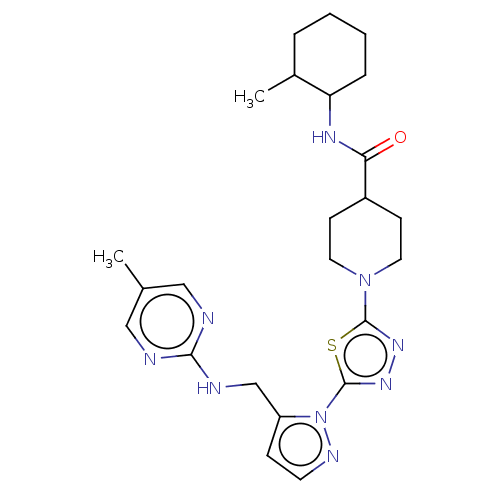
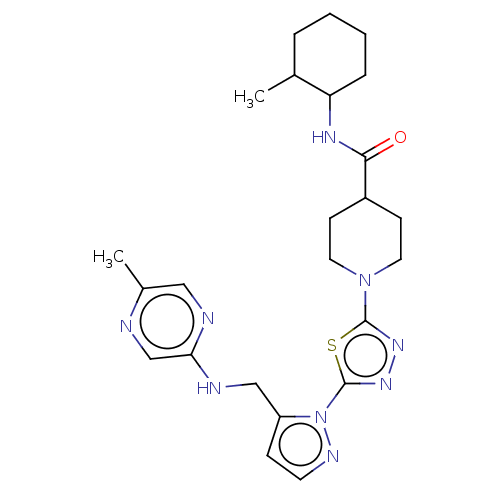
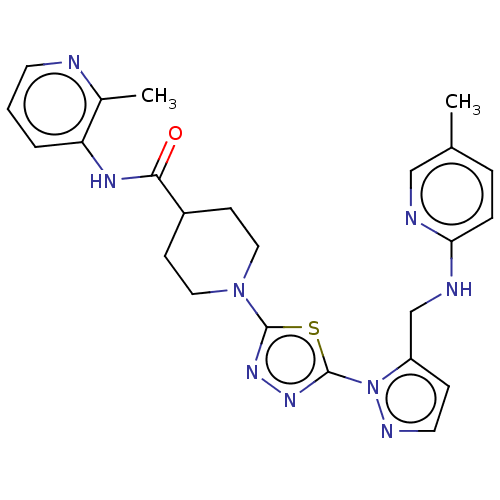
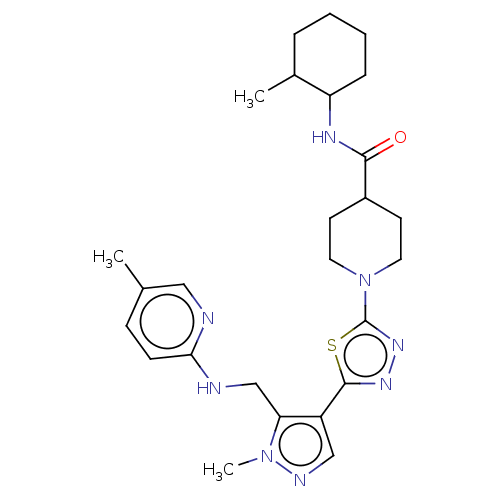
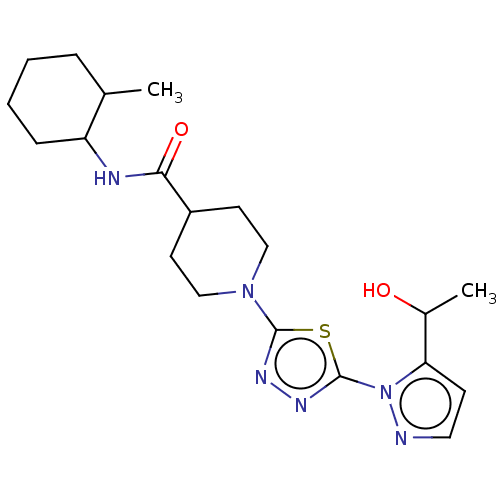
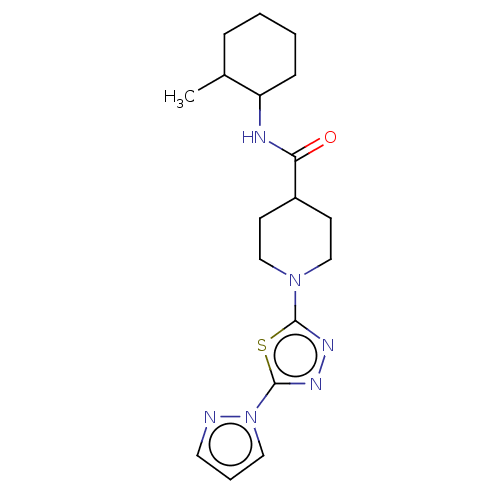
Report error Found 39 Enz. Inhib. hit(s) with all data for entry = 50008982
Affinity DataIC50: 3.90nMAssay Description:Displacement of BODIPY-Cyclopamine from human HA-tagged Smo receptor expressed in human U2OS cells at 1 to 10000 nM incubated for 2 hrs by DAPI stain...More data for this Ligand-Target Pair
Affinity DataIC50: 7.10nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.10nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.10nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.30nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.40nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:Displacement of BODIPY-Cyclopamine from human HA-tagged Smo receptor expressed in human U2OS cells at 1 to 10000 nM incubated for 2 hrs by DAPI stain...More data for this Ligand-Target Pair
Affinity DataIC50: 42nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 46nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 46nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 47nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 47nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 51nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 57nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 69nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 74nMAssay Description:Displacement of BODIPY-Cyclopamine from human HA-tagged Smo receptor expressed in human U2OS cells at 1 to 10000 nM incubated for 2 hrs by DAPI stain...More data for this Ligand-Target Pair
Affinity DataIC50: 81nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 140nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 141nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 180nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 220nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 280nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 290nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 410nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 460nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 550nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.70E+3nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.72E+3nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated ...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Soochow University
Curated by ChEMBL
Soochow University
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of human ERG by patch clamp methodMore data for this Ligand-Target Pair