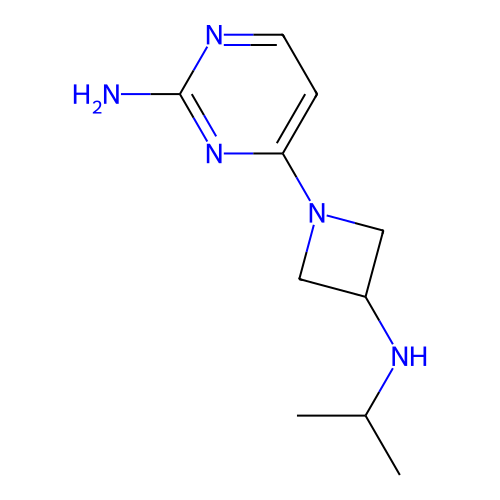
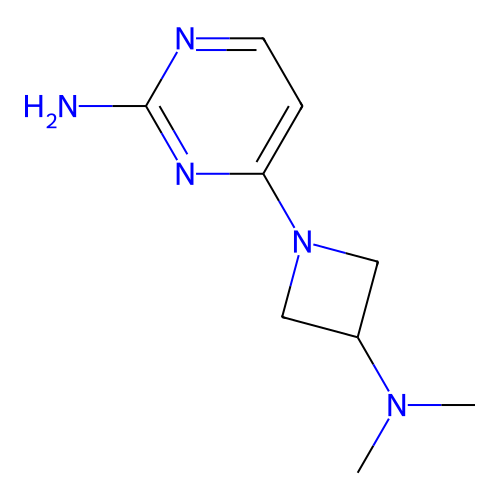
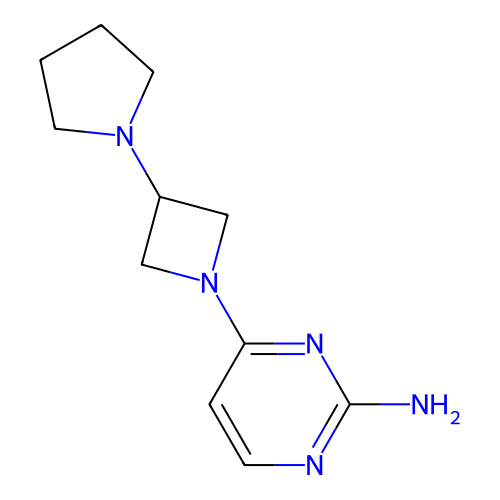
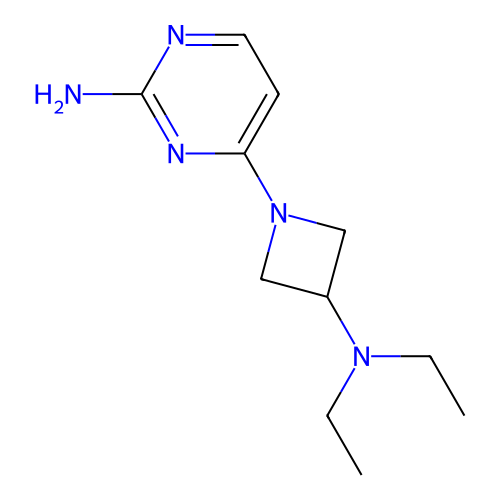
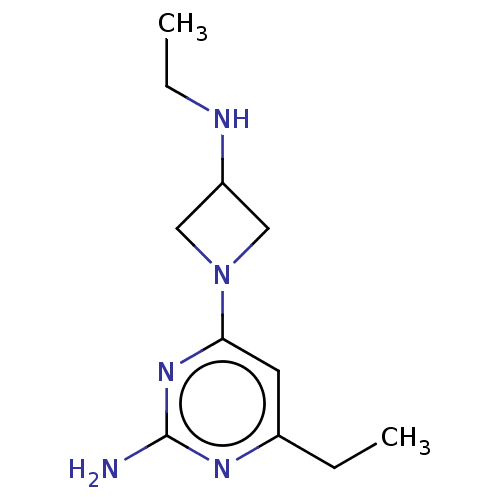
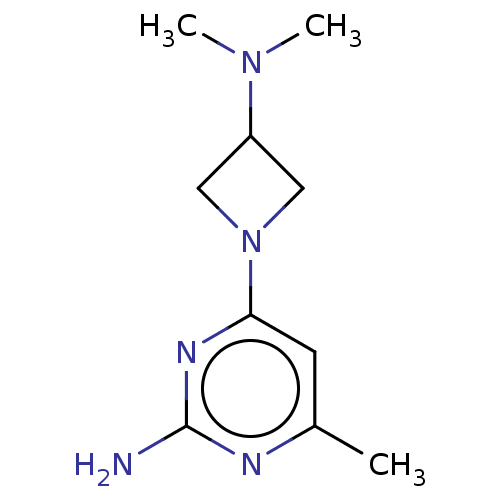
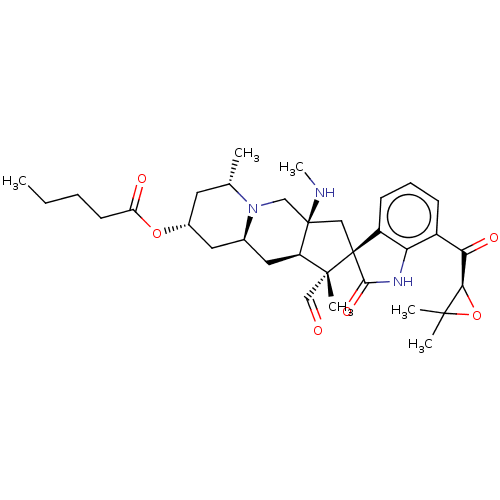
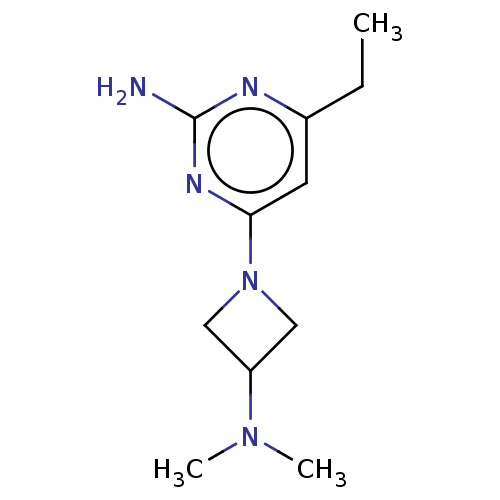
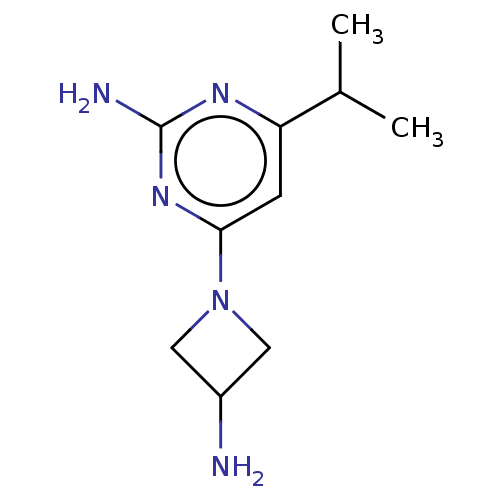
Report error Found 63 Enz. Inhib. hit(s) with all data for entry = 50007933
Affinity DataEC50: 0.100nMAssay Description:Agonist activity at mouse H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataEC50: 0.316nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataEC50: 0.631nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataEC50: 0.794nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Displacement of [3H]NAMH from mouse H3R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataEC50: 1nMAssay Description:Agonist activity at mouse H4R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataEC50: 1.30nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataEC50: 2nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataEC50: 2.5nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataEC50: 3.20nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataKi: 3.20nMAssay Description:Displacement of [3H]NAMH from human H3R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataEC50: 3.20nMAssay Description:Agonist activity at human H4R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataEC50: 4nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells by [35S]GTPgammaS binding assayMore data for this Ligand-Target Pair
Affinity DataEC50: 4nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataEC50: 5nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataEC50: 5nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataEC50: 5nMAssay Description:Agonist activity at human H3R expressed on SK-N-MC cells assessed as inhibition of forskolin-induced beta galactosidase activity preincubated with fo...More data for this Ligand-Target Pair
Affinity DataKi: 6.30nMAssay Description:Displacement of [3H]NAMH from human H3R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataEC50: 7.90nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataEC50: 7.90nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataKi: 7.90nMAssay Description:Displacement of [3H]NAMH from human H4R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Displacement of [3H]NAMH from human H3R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Displacement of [3H]-N-alpha-methylhistamine from human histamine H3 receptor expressed in SK-N-MC cells incubated for 40 min by liquid scintillation...More data for this Ligand-Target Pair
Affinity DataEC50: 10nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Displacement of [3H]NAMH from human H3R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataEC50: 13nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Displacement of [3H]NAMH from human H3R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataEC50: 13nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataEC50: 16nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Displacement of [3H]NAMH from human H3R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Displacement of [3H]NAMH from human H3R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Displacement of [3H]histamine from mouse H4R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataEC50: 25nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataKi: 32nMAssay Description:Displacement of [3H]NAMH from human H3R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataEC50: 32nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataEC50: 32nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataKi: 40nMAssay Description:Displacement of [3H]NAMH from human H3R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 40nMAssay Description:Displacement of [3H]NAMH from human H3R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 40nMAssay Description:Displacement of [3H]NAMH from human H3R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataEC50: 40nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataKi: 50nMAssay Description:Displacement of [3H]NAMH from human H3R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 50nMAssay Description:Displacement of [3H]NAMH from human H3R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 50nMAssay Description:Displacement of [3H]NAMH from human H3R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 50nMAssay Description:Displacement of [3H]-N-alpha-methylhistamine from human histamine H3 receptor expressed in CHO cells incubated for 40 min by liquid scintillation cou...More data for this Ligand-Target Pair
Affinity DataEC50: 63nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataKi: 63nMAssay Description:Displacement of [3H]NAMH from human H3R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataEC50: 63nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells by [35S]GTPgammaS binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 79nMAssay Description:Displacement of [3H]NAMH from human H3R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataEC50: 79nMAssay Description:Agonist activity at human H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataKi: 79nMAssay Description:Displacement of [3H]NAMH from human H3R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair