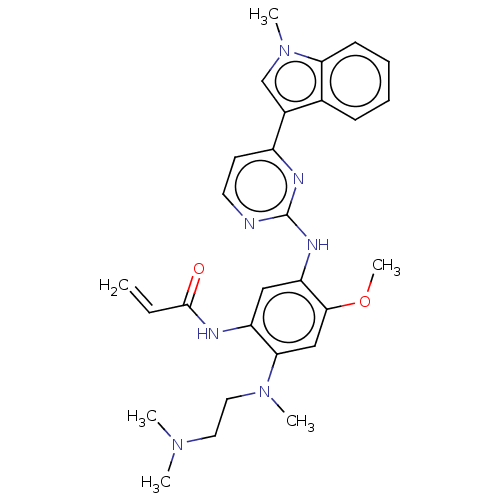
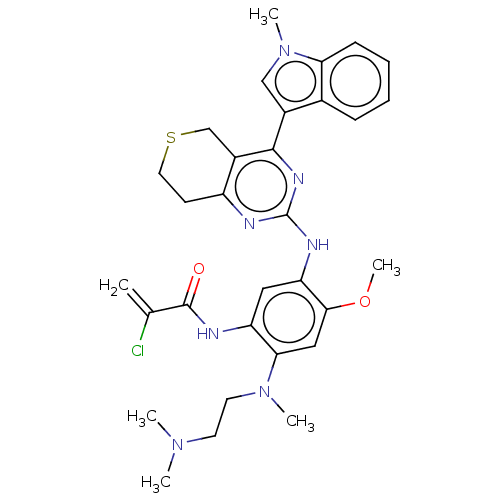
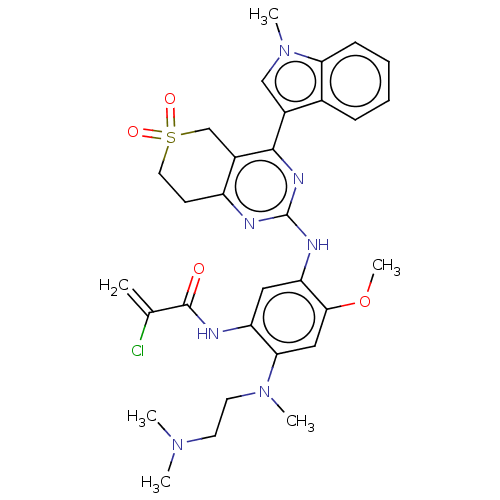
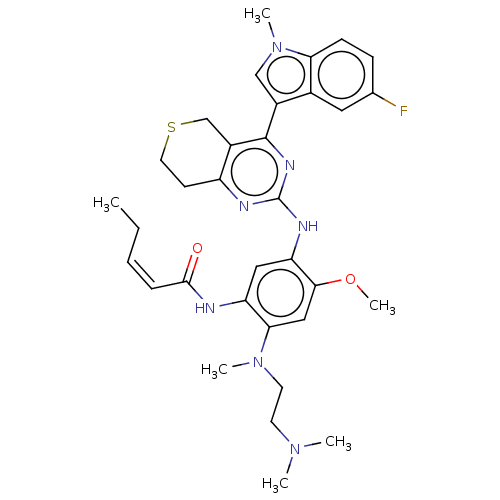
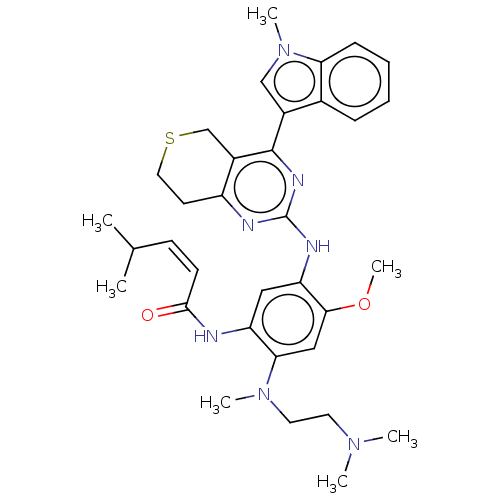
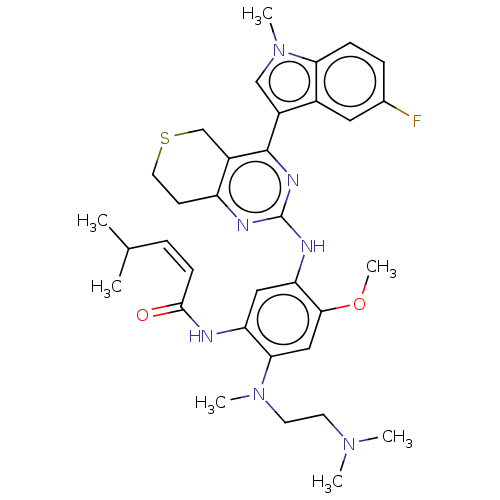
Report error Found 22 Enz. Inhib. hit(s) with all data for entry = 50006918
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 8nMAssay Description:Inhibition of EGFR T790M/L858R double mutant (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 16nMAssay Description:Inhibition of EGFR T790M/L858R double mutant (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 16nMAssay Description:Inhibition of wildtype EGFR (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 31nMAssay Description:Inhibition of EGFR T790M/L858R double mutant (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 35nMAssay Description:Inhibition of EGFR T790M/L858R double mutant (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 104nMAssay Description:Inhibition of wildtype EGFR (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 201nMAssay Description:Inhibition of wildtype EGFR (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 511nMAssay Description:Inhibition of wildtype EGFR (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 570nMAssay Description:Inhibition of wildtype EGFR (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 990nMAssay Description:Inhibition of EGFR T790M/L858R double mutant (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 1.13E+3nMAssay Description:Inhibition of EGFR T790M/L858R double mutant (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 1.25E+3nMAssay Description:Inhibition of EGFR T790M/L858R double mutant (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 1.62E+3nMAssay Description:Inhibition of EGFR T790M/L858R double mutant (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 1.69E+3nMAssay Description:Inhibition of EGFR T790M/L858R double mutant (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 1.75E+3nMAssay Description:Inhibition of EGFR T790M/L858R double mutant (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of EGFR T790M/L858R double mutant (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of wildtype EGFR (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of wildtype EGFR (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of wildtype EGFR (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of wildtype EGFR (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of wildtype EGFR (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Jiangxi Science & Technology Normal University
Curated by ChEMBL
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of wildtype EGFR (unknown origin) using biotin as substrate in presence of ATP by mobility shift assay based ELISAMore data for this Ligand-Target Pair