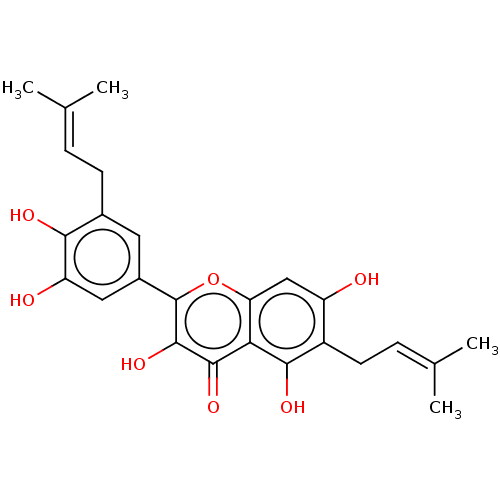
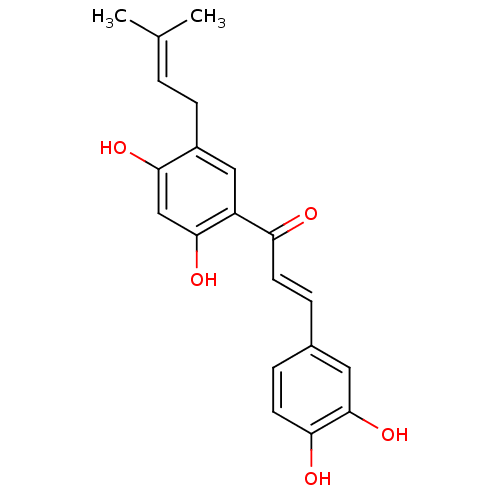
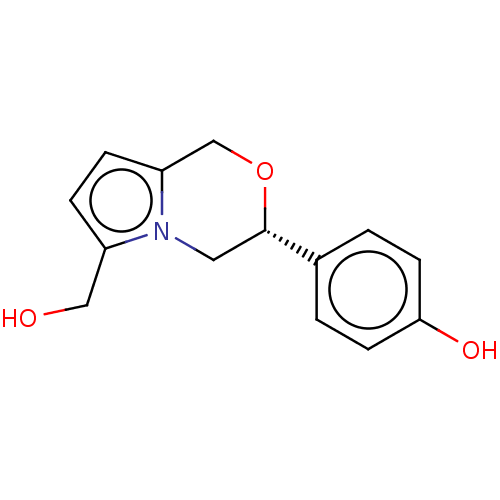
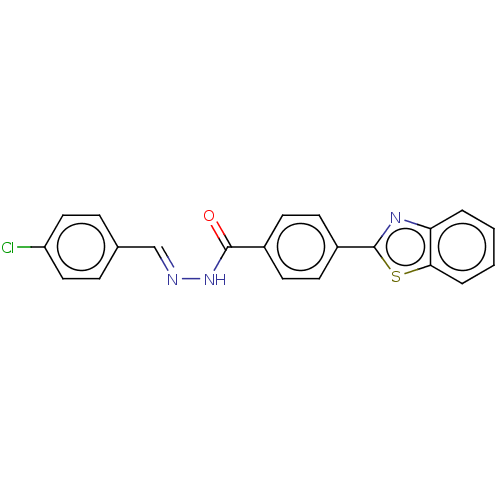
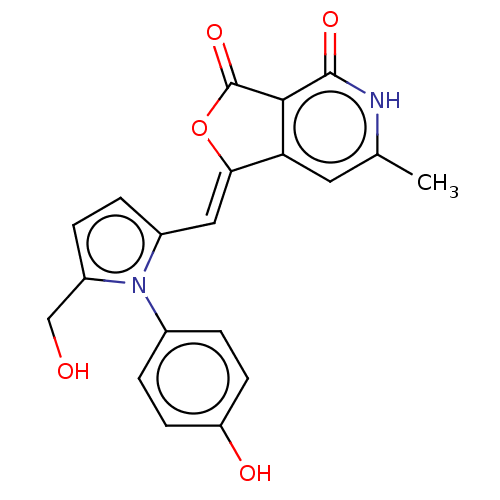
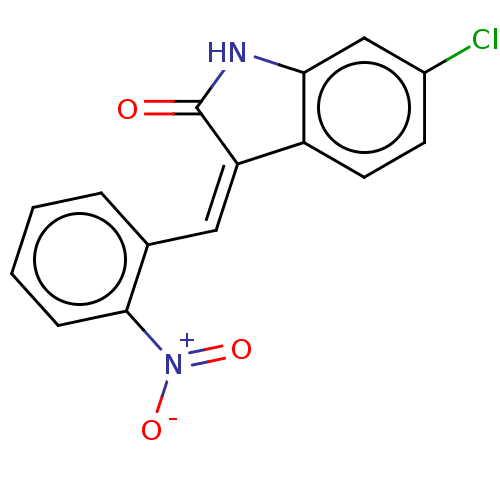
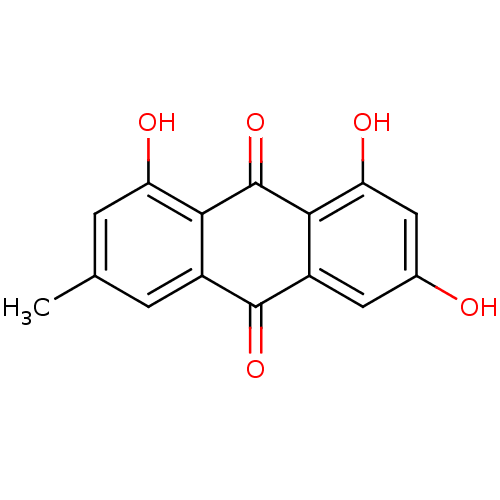
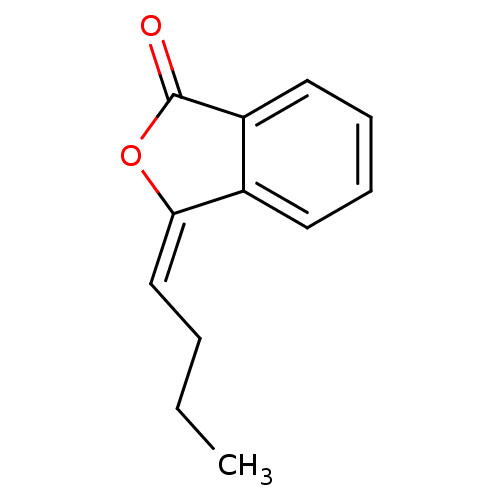
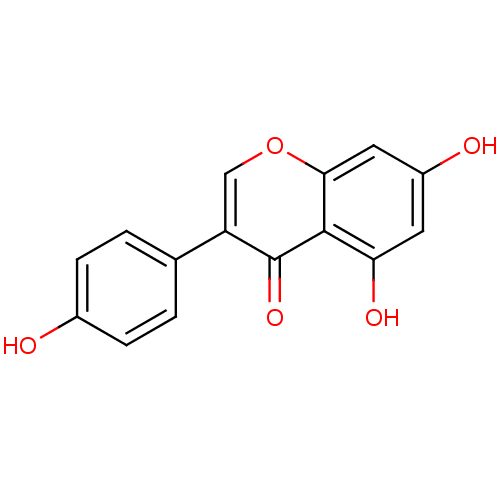
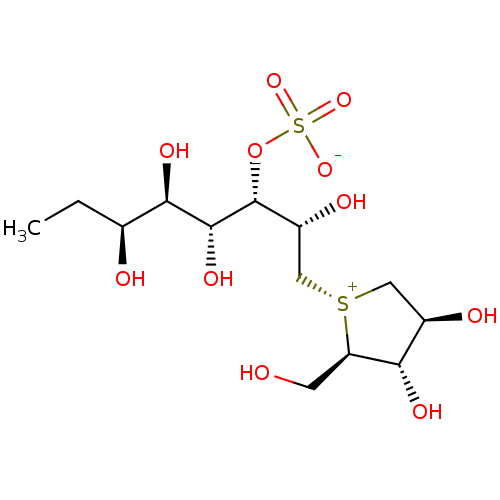
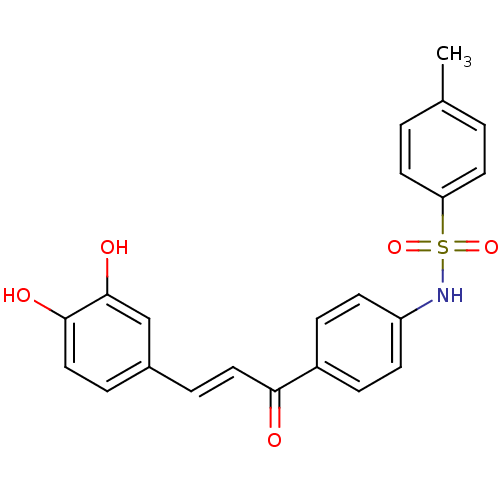
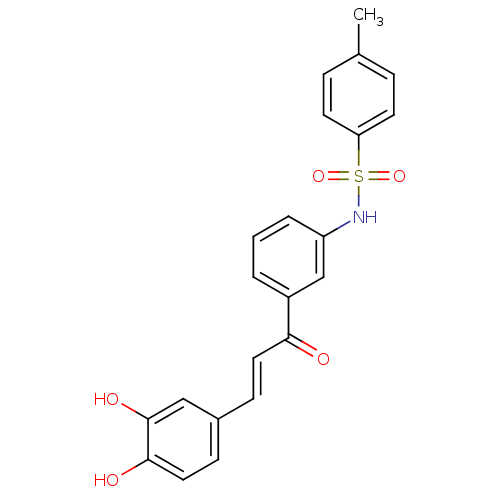
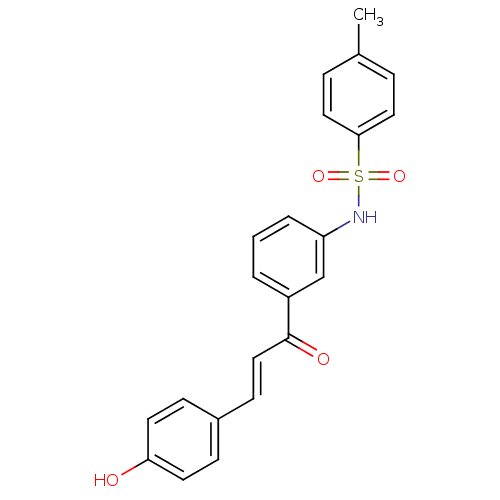
Report error Found 48 Enz. Inhib. hit(s) with all data for entry = 50011775
Affinity DataKi: 2.30E+3nMAssay Description:Mixed type inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl-alpha-D glucopyranoside as substrate incubated for 30 mins by Linewea...More data for this Ligand-Target Pair
Affinity DataKi: 4.20E+3nMAssay Description:Mixed type inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl-alpha-D glucopyranoside as substrate incubated for 30 mins by Linewea...More data for this Ligand-Target Pair
Affinity DataKi: 5.30E+3nMAssay Description:Non-competitive inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl-alpha-D glucopyranoside as substrate incubated for 30 mins by Li...More data for this Ligand-Target Pair
Affinity DataIC50: 6.70E+3nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using sucrose as substrate incubated for 30 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 9.30E+3nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using sucrose as substrate incubated for 30 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.29E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+4nMAssay Description:Inhibition of jack bean alpha-mannosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMAssay Description:Inhibition of jack bean alpha-mannosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 2.17E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+4nMAssay Description:Inhibition of jack bean alpha-mannosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+4nMAssay Description:Inhibition of jack bean alpha-mannosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+4nMAssay Description:Inhibition of jack bean alpha-mannosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+4nMAssay Description:Inhibition of jack bean alpha-mannosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+4nMAssay Description:Inhibition of jack bean alpha-mannosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 3.04E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 3.84E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+4nMAssay Description:Inhibition of jack bean alpha-mannosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 4.12E+4nMAssay Description:Inhibition of jack bean alpha-mannosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 4.90E+4nMAssay Description:Inhibition of jack bean alpha-mannosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 4.90E+4nMAssay Description:Inhibition of jack bean alpha-mannosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 5.10E+4nMAssay Description:Inhibition of jack bean alpha-mannosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 5.11E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 5.40E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 5.40E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 6.14E+4nMAssay Description:Inhibition of jack bean alpha-mannosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 6.50E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 6.50E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 6.50E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 7.10E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 8.20E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 8.20E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 8.90E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 9.10E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 9.10E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 9.10E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 9.10E+4nMAssay Description:Inhibition of bovine kidney alpha-fucosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 9.23E+4nMAssay Description:Inhibition of jack bean alpha-mannosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.05E+5nMAssay Description:Inhibition of jack bean alpha-mannosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.05E+5nMAssay Description:Inhibition of jack bean alpha-mannosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.05E+5nMAssay Description:Inhibition of jack bean alpha-mannosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.05E+5nMAssay Description:Inhibition of jack bean alpha-mannosidase using PNPG as substrate incubated for 10 mins by spectrophotometric methodMore data for this Ligand-Target Pair