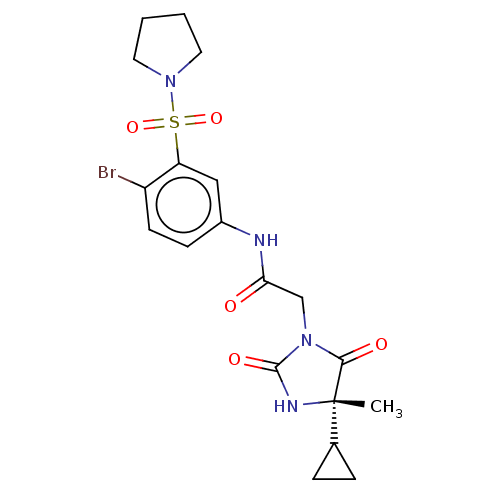
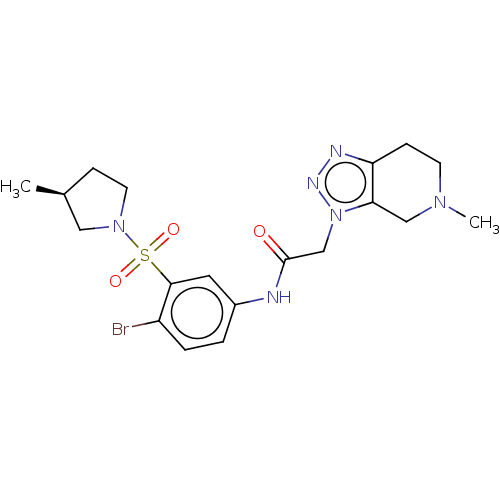
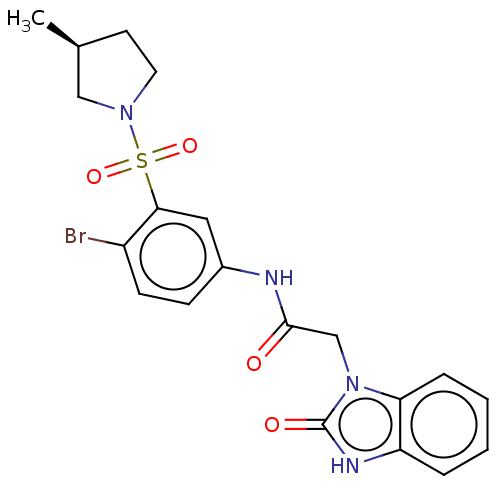
Report error Found 179 Enz. Inhib. hit(s) with all data for entry = 50008013
Affinity DataIC50: 6.30nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
Affinity DataKd: 7.90nMAssay Description:Binding affinity to human partial length CECR2 (P423 to D543 residues) expressed in bacterial expression system by BROMOscan assayMore data for this Ligand-Target Pair
Affinity DataKd: 10nMAssay Description:Binding affinity to CECR2 (unknown origin) by ITC analysisMore data for this Ligand-Target Pair
Affinity DataKd: 10nMAssay Description:Binding affinity to CECR2 (unknown origin) by ITC analysisMore data for this Ligand-Target Pair
TargetATPase family AAA domain-containing protein 2(Human)
University of Strathclyde
Curated by ChEMBL
University of Strathclyde
Curated by ChEMBL
Affinity DataKd: 10nMAssay Description:Binding affinity to human partial length ATAD2A (Q981 to R1108 residues) expressed in bacterial expression system by BROMOscan assayMore data for this Ligand-Target Pair
Affinity DataKd: 10nMAssay Description:Binding affinity to human partial length CECR2 (P423 to D543 residues) expressed in bacterial expression system by BROMOscan assayMore data for this Ligand-Target Pair
TargetBromodomain adjacent to zinc finger domain protein 2A(Human)
University of Strathclyde
Curated by ChEMBL
University of Strathclyde
Curated by ChEMBL
Affinity DataKd: 16nMAssay Description:Binding affinity to human partial length BAZ2A (M1792 to L1905 residues) expressed in bacterial expression system by BROMOscan assayMore data for this Ligand-Target Pair
TargetTranscription initiation factor TFIID subunit 1(Human)
University of Strathclyde
Curated by ChEMBL
University of Strathclyde
Curated by ChEMBL
Affinity DataKd: 16nMAssay Description:Binding affinity to human partial length TAF1 bromodomain 2 (D1521 to D1656 residues) expressed in bacterial expression system by BROMOscan assayMore data for this Ligand-Target Pair
TargetATPase family AAA domain-containing protein 2B(Human)
University of Strathclyde
Curated by ChEMBL
University of Strathclyde
Curated by ChEMBL
Affinity DataKd: 20nMAssay Description:Binding affinity to human partial length ATAD2B (Q955 to R1082 residues) expressed in bacterial expression system by BROMOscan assayMore data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
TargetTranscription initiation factor TFIID subunit 1-like(Human)
University of Strathclyde
Curated by ChEMBL
University of Strathclyde
Curated by ChEMBL
Affinity DataKd: 25nMAssay Description:Binding affinity to human partial length TAF1L bromodomain 2 (Q1523 to D1654 residues) expressed in bacterial expression system by BROMOscan assayMore data for this Ligand-Target Pair
TargetATPase family AAA domain-containing protein 2(Human)
University of Strathclyde
Curated by ChEMBL
University of Strathclyde
Curated by ChEMBL
Affinity DataIC50: 25nMAssay Description:Displacement of Alexa Fluor 488-labeled ligand from FLAG-6His-Tev-fused ATAD2 (981 to 1121 residue) (unknown origin) measured after 30 mins by TR-FRE...More data for this Ligand-Target Pair
Affinity DataKd: 25nMAssay Description:Binding affinity to human partial length CECR2 (P423 to D543 residues) expressed in bacterial expression system by BROMOscan assayMore data for this Ligand-Target Pair
Affinity DataIC50: 32nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:Inhibition of BSP-BODIPY binding to Nanoluc-tagged CECR2 (unknown origin) expressed in HEK293 cells measured after 4 to 6 hrs by NanoBRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Inhibition of BSP-BODIPY binding to Nanoluc-tagged CECR2 (unknown origin) expressed in HEK293 cells measured after 4 to 6 hrs by NanoBRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
TargetTranscription initiation factor TFIID subunit 1(Human)
University of Strathclyde
Curated by ChEMBL
University of Strathclyde
Curated by ChEMBL
Affinity DataKd: 63nMAssay Description:Binding affinity to human partial length TAF1 bromodomain 2 (D1521 to D1656 residues) expressed in bacterial expression system by BROMOscan assayMore data for this Ligand-Target Pair
Affinity DataIC50: 63nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
TargetATPase family AAA domain-containing protein 2(Human)
University of Strathclyde
Curated by ChEMBL
University of Strathclyde
Curated by ChEMBL
Affinity DataIC50: 63nMAssay Description:Displacement of Alexa Fluor 488-labeled ligand from FLAG-6His-Tev-fused ATAD2 (981 to 1121 residue) (unknown origin) measured after 30 mins by TR-FRE...More data for this Ligand-Target Pair
Affinity DataKd: 79nMAssay Description:Binding affinity to CECR2 (unknown origin) by ITC analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 79nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
Affinity DataIC50: 79nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
TargetATPase family AAA domain-containing protein 2(Human)
University of Strathclyde
Curated by ChEMBL
University of Strathclyde
Curated by ChEMBL
Affinity DataIC50: 100nMAssay Description:Displacement of Alexa Fluor 488-labeled ligand from FLAG-6His-Tev-fused ATAD2 (981 to 1121 residue) (unknown origin) measured after 30 mins by TR-FRE...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
Affinity DataIC50: 126nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
Affinity DataIC50: 126nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
Affinity DataKd: 126nMAssay Description:Binding affinity to human partial length BRD1 (E556 to A688 residues) expressed in bacterial expression system by BROMOscan assayMore data for this Ligand-Target Pair
TargetBromodomain and PHD finger-containing protein 3(Human)
University of Strathclyde
Curated by ChEMBL
University of Strathclyde
Curated by ChEMBL
Affinity DataKd: 126nMAssay Description:Binding affinity to human partial length BRPF3 (E588 to G701 residues) expressed in bacterial expression system by BROMOscan assayMore data for this Ligand-Target Pair
Affinity DataIC50: 126nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
Affinity DataKd: 158nMAssay Description:Binding affinity to human partial length GCN5L2 (E726 to K837 residues) expressed in bacterial expression system by BROMOscan assayMore data for this Ligand-Target Pair
Affinity DataIC50: 158nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
TargetTranscription initiation factor TFIID subunit 1(Human)
University of Strathclyde
Curated by ChEMBL
University of Strathclyde
Curated by ChEMBL
Affinity DataKd: 200nMAssay Description:Binding affinity to human partial length TAF1 bromodomain 2 (D1521 to D1656 residues) expressed in bacterial expression system by BROMOscan assayMore data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
Affinity DataIC50: 251nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
Affinity DataIC50: 251nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
Affinity DataIC50: 251nMAssay Description:Inhibition of BSP-BODIPY binding to Nanoluc-tagged CECR2 (unknown origin) expressed in HEK293 cells measured after 4 to 6 hrs by NanoBRET assayMore data for this Ligand-Target Pair
TargetATPase family AAA domain-containing protein 2(Human)
University of Strathclyde
Curated by ChEMBL
University of Strathclyde
Curated by ChEMBL
Affinity DataIC50: 251nMAssay Description:Displacement of Alexa Fluor 488-labeled ligand from FLAG-6His-Tev-fused ATAD2 (981 to 1121 residue) (unknown origin) measured after 30 mins by TR-FRE...More data for this Ligand-Target Pair
Affinity DataIC50: 251nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
Affinity DataKd: 251nMAssay Description:Binding affinity to human partial length GCN5L2 (E726 to K837 residues) expressed in bacterial expression system by BROMOscan assayMore data for this Ligand-Target Pair
Affinity DataIC50: 251nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
TargetATPase family AAA domain-containing protein 2(Human)
University of Strathclyde
Curated by ChEMBL
University of Strathclyde
Curated by ChEMBL
Affinity DataIC50: 251nMAssay Description:Displacement of Alexa Fluor 488-labeled ligand from FLAG-6His-Tev-fused ATAD2 (981 to 1121 residue) (unknown origin) measured after 30 mins by TR-FRE...More data for this Ligand-Target Pair
Affinity DataIC50: 251nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
Affinity DataIC50: 316nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
Affinity DataIC50: 398nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
TargetATPase family AAA domain-containing protein 2(Human)
University of Strathclyde
Curated by ChEMBL
University of Strathclyde
Curated by ChEMBL
Affinity DataIC50: 501nMAssay Description:Displacement of Alexa Fluor 488-labeled ligand from FLAG-6His-Tev-fused ATAD2 (981 to 1121 residue) (unknown origin) measured after 30 mins by TR-FRE...More data for this Ligand-Target Pair
Affinity DataIC50: 501nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
Affinity DataIC50: 501nMAssay Description:Displacement of Alexa Fluor 647-labeled GSK tracer from FLAG-6His-Tev-fused CECR2 (424 to 543 residue) (unknown origin) measured after 15 mins by TR-...More data for this Ligand-Target Pair
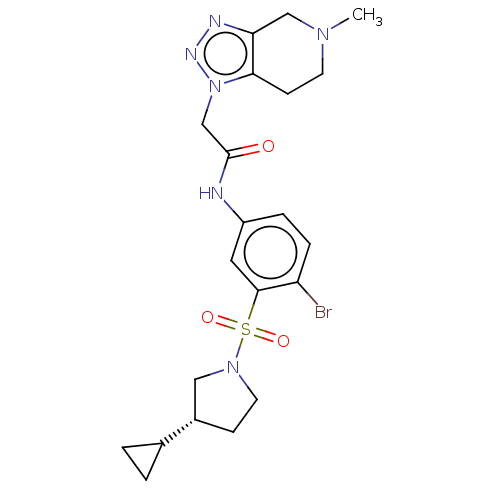
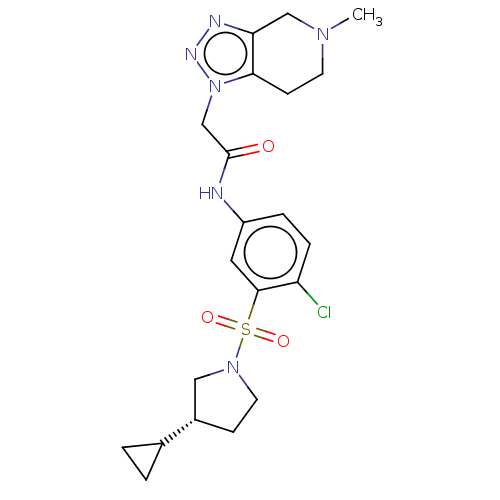


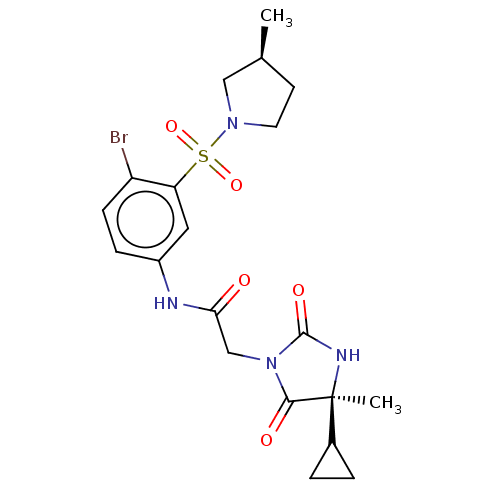
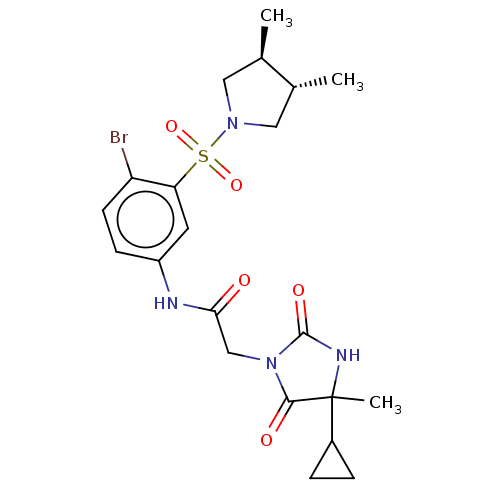
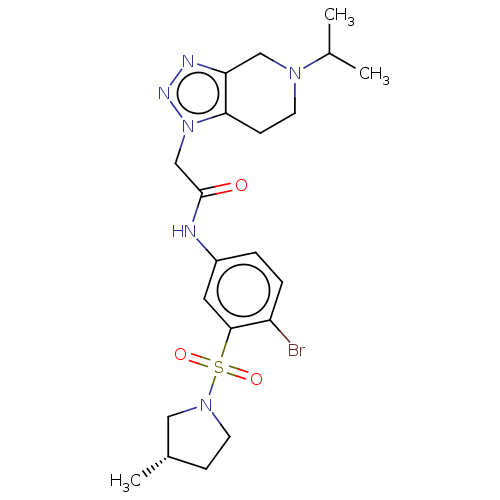
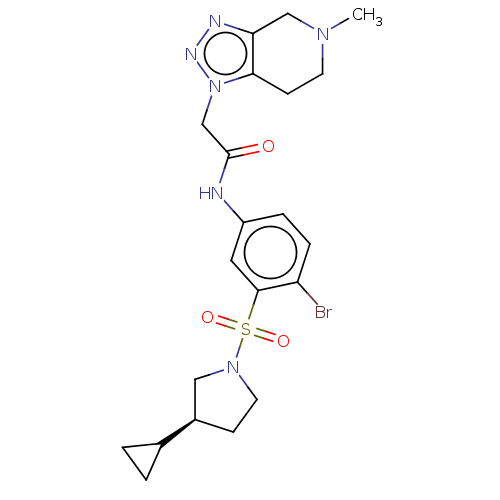


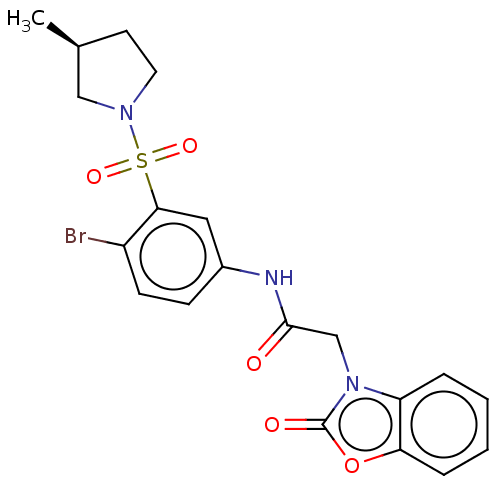
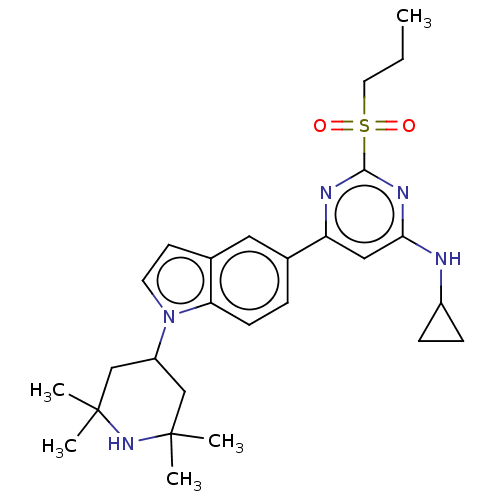
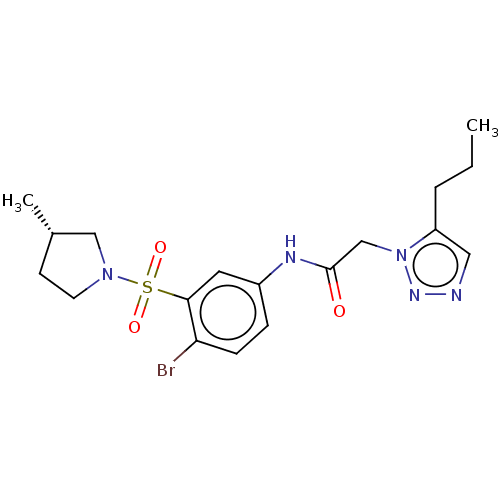
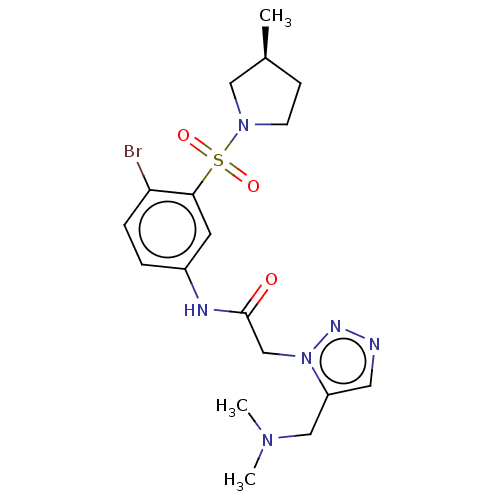

 3D Structure (crystal)
3D Structure (crystal)