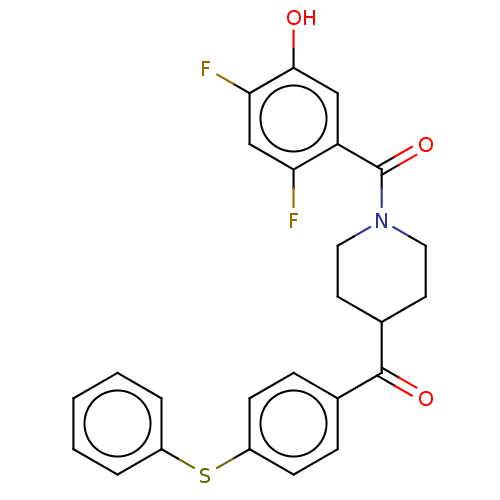
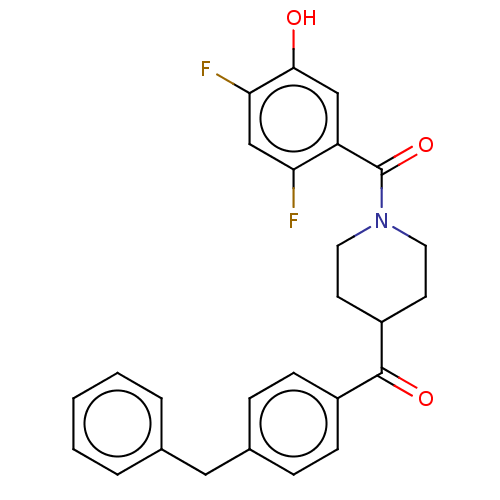
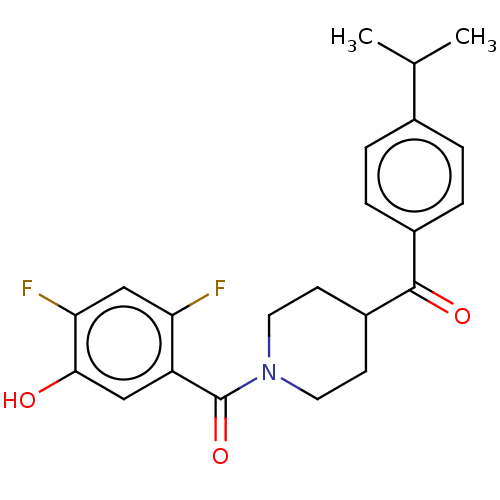
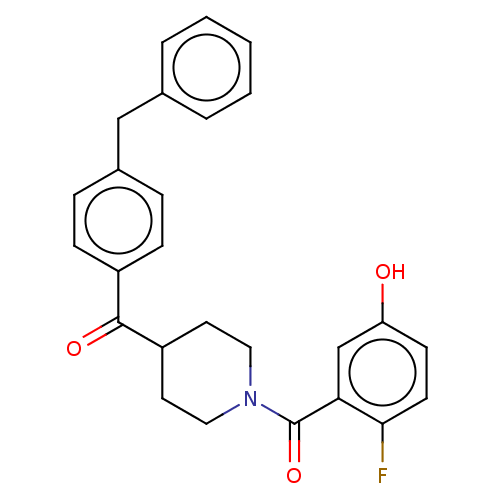
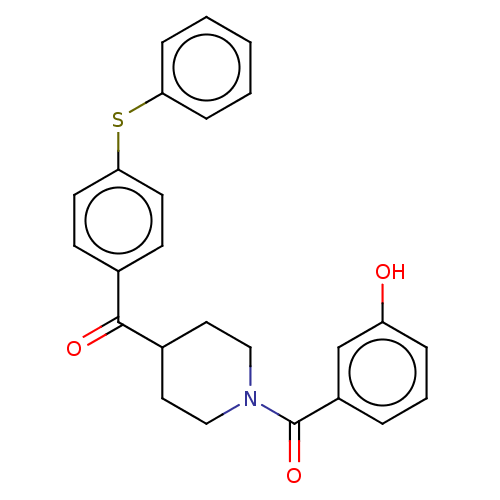
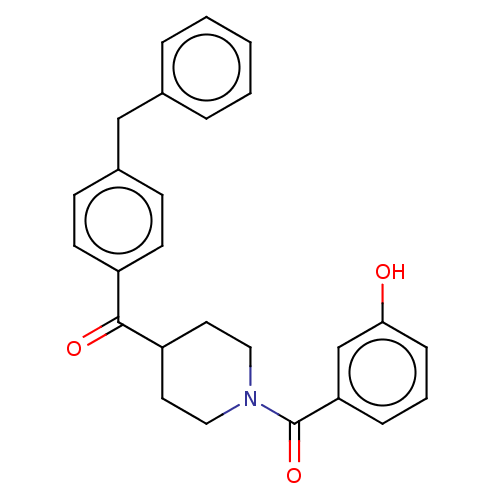
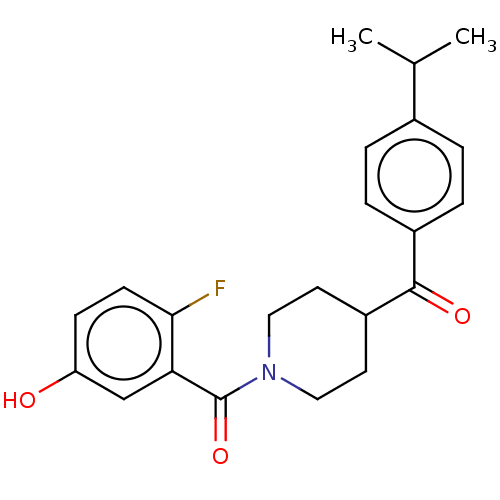
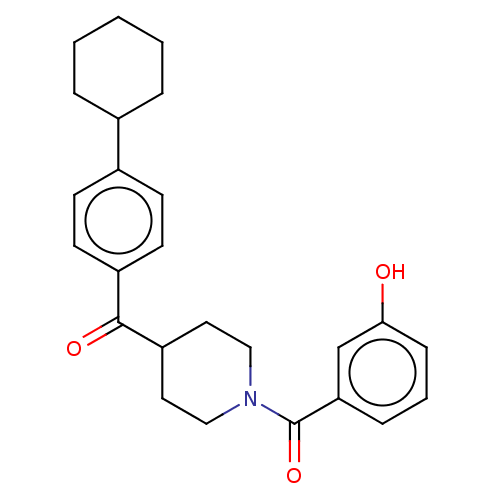
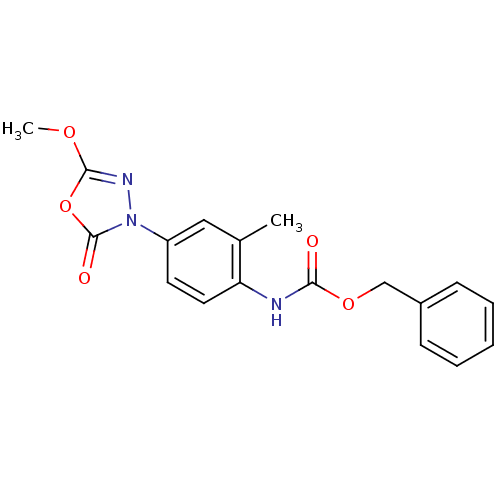
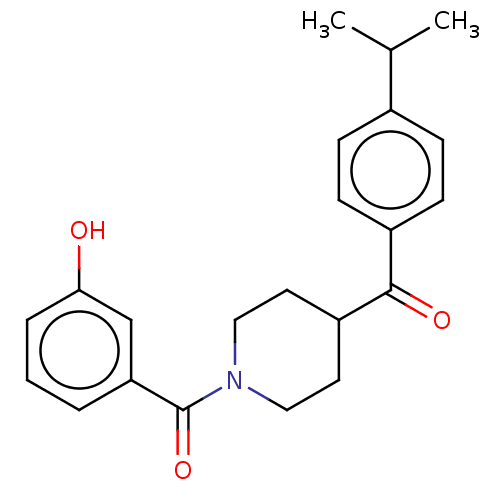
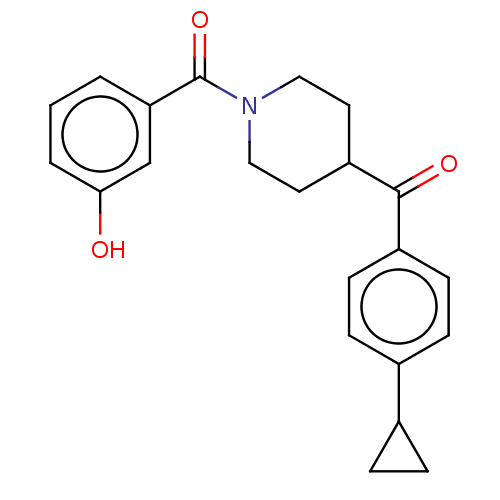
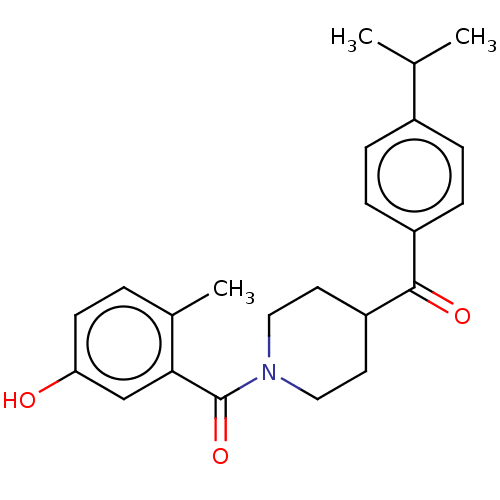
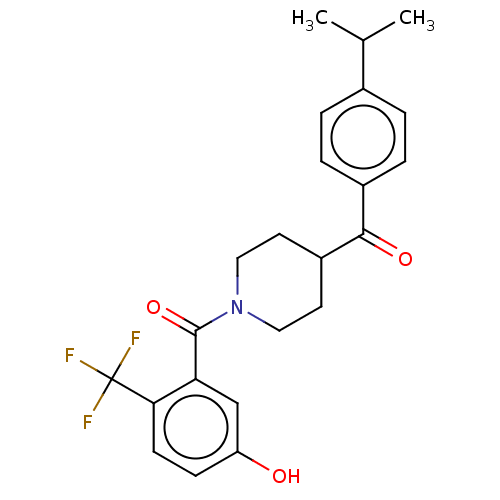
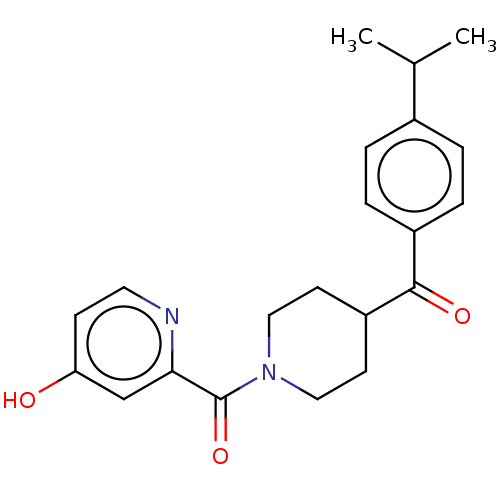
Report error Found 55 Enz. Inhib. hit(s) with all data for entry = 50013016
Affinity DataKi: 11nMAssay Description:Competitive inhibition of human recombinant MAGL using varying concentration of 4-nitrophenylacetate as substrate incubated for 30 mins by microplate...More data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataKi: 27nMAssay Description:Competitive inhibition of human recombinant MAGL using varying concentration of 4-nitrophenylacetate as substrate incubated for 30 mins by microplate...More data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:Competitive inhibition of human recombinant MAGL using varying concentration of 4-nitrophenylacetate as substrate incubated for 30 mins by microplate...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 31nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 36nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataKi: 37nMAssay Description:Competitive inhibition of human recombinant MAGL using varying concentration of 4-nitrophenylacetate as substrate incubated for 30 mins by microplate...More data for this Ligand-Target Pair
Affinity DataIC50: 49nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 74nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 78nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 81nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 129nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 142nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 278nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 368nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 568nMAssay Description:Inhibition of MAGL in human U937 cells assessed as reduction in [3H]glycerol formation using [1,2,3-3H]2-OG as substrate incubated for 15 mins by liq...More data for this Ligand-Target Pair
Affinity DataIC50: 758nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 844nMAssay Description:Inhibition of MAGL in human U937 cells assessed as reduction in [3H]glycerol formation using [1,2,3-3H]2-OG as substrate incubated for 15 mins by liq...More data for this Ligand-Target Pair
Affinity DataIC50: 868nMAssay Description:Inhibition of MAGL in human U937 cells assessed as reduction in [3H]glycerol formation using [1,2,3-3H]2-OG as substrate incubated for 15 mins by liq...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.23E+3nMAssay Description:Inhibition of MAGL in human U937 cells assessed as reduction in [3H]glycerol formation using [1,2,3-3H]2-OG as substrate incubated for 15 mins by liq...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.30E+3nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+3nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.40E+3nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.30E+3nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.40E+3nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.50E+3nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 8.20E+3nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Displacement of [3H]CP55940 from human CB1 receptor incubated for 90 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Displacement of [3H]CP55940 from human CB1 receptor incubated for 90 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Displacement of [3H]CP55940 from human CB1 receptor incubated for 90 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Displacement of [3H]CP55940 from human CB1 receptor incubated for 90 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Displacement of [3H]CP55940 from human CB2 receptor incubated for 90 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Displacement of [3H]CP55940 from human CB2 receptor incubated for 90 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Displacement of [3H]CP55940 from human CB2 receptor incubated for 90 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Displacement of [3H]CP55940 from human CB2 receptor incubated for 90 mins by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human ABHD6 expressed in HEK293 cells using [3H]2-OG as substrate preincubated for 30 mins followed by substrate addition and measured ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human ABHD6 expressed in HEK293 cells using [3H]2-OG as substrate preincubated for 30 mins followed by substrate addition and measured ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human ABHD6 expressed in HEK293 cells using [3H]2-OG as substrate preincubated for 30 mins followed by substrate addition and measured ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human ABHD6 expressed in HEK293 cells using [3H]2-OG as substrate preincubated for 30 mins followed by substrate addition and measured ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human ABHD12 expressed in HEK293 cells using [3H]2-OG as substrate preincubated for 30 mins followed by substrate addition and measured...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human ABHD12 expressed in HEK293 cells using [3H]2-OG as substrate preincubated for 30 mins followed by substrate addition and measured...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human ABHD12 expressed in HEK293 cells using [3H]2-OG as substrate preincubated for 30 mins followed by substrate addition and measured...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human ABHD12 expressed in HEK293 cells using [3H]2-OG as substrate preincubated for 30 mins followed by substrate addition and measured...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human recombinant MAGL using 4-nitrophenylacetate as substrate incubated for 30 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of FAAH in human U937 cells assessed as reduction in [3H]ethanolamine formation using [ethanolamine-1-3H]AEA as substrate incubated for 15...More data for this Ligand-Target Pair