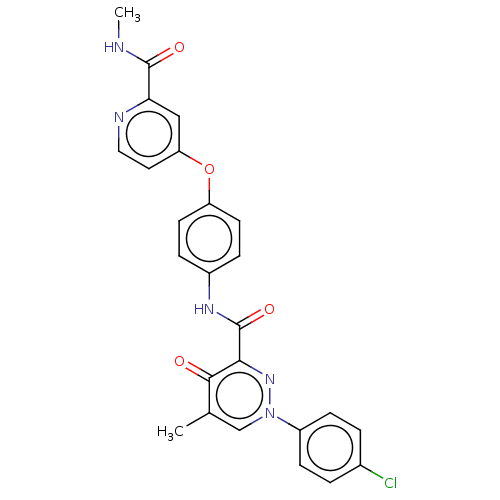
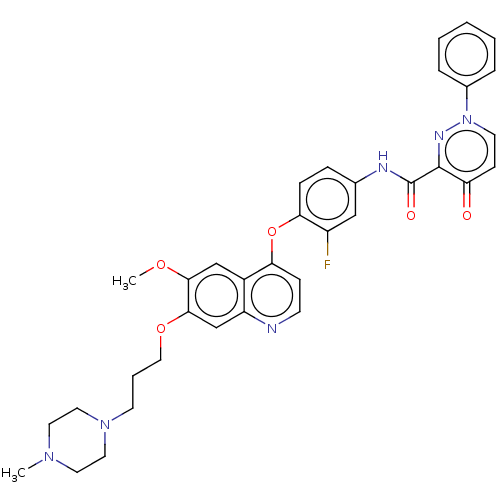
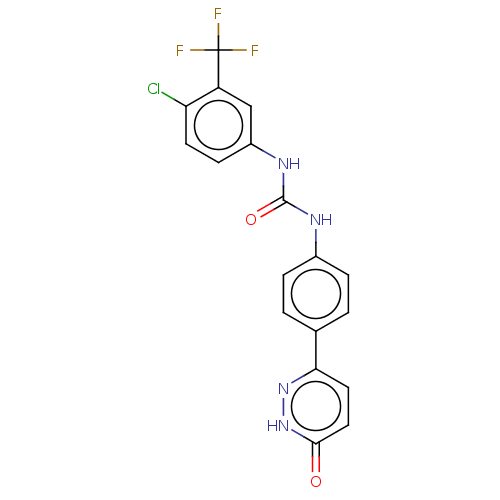
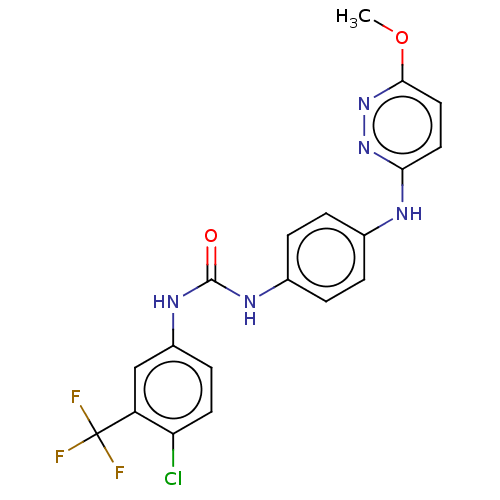
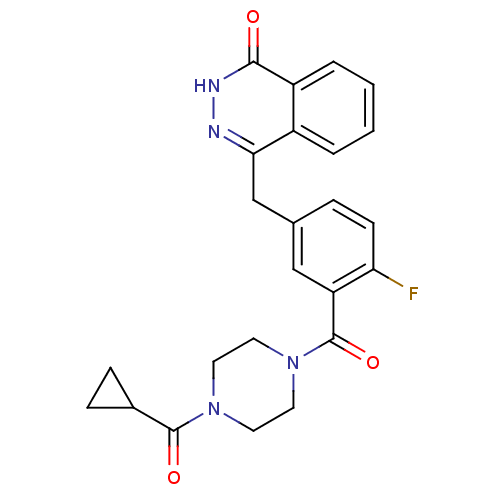
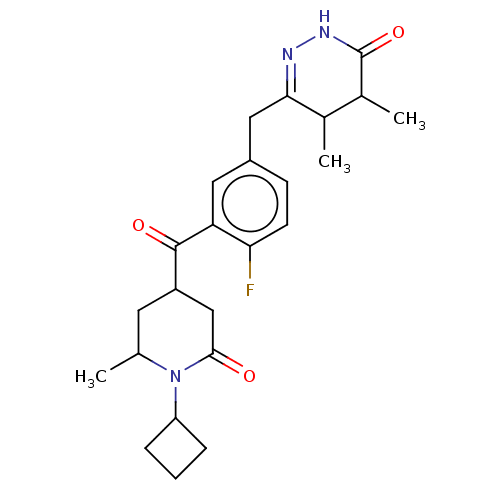
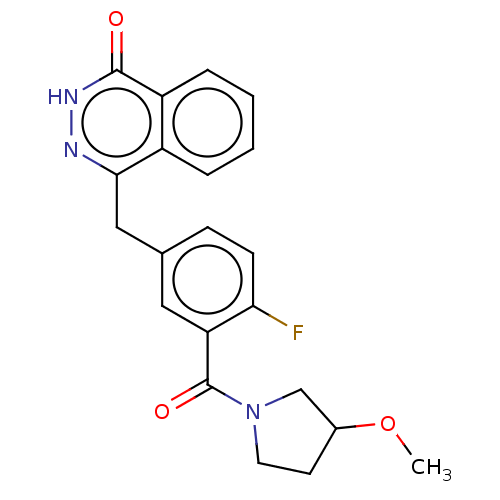
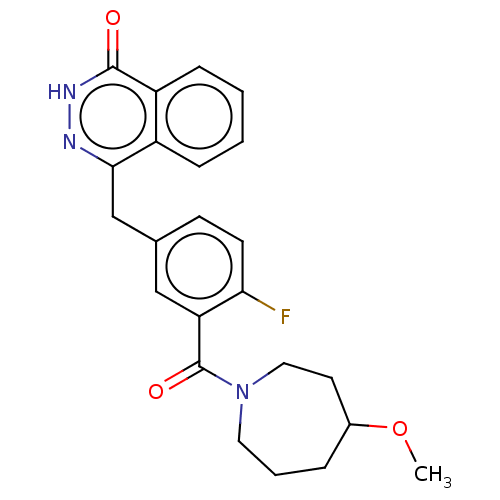
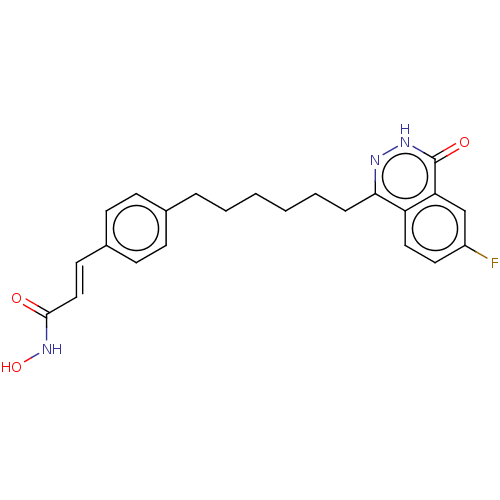
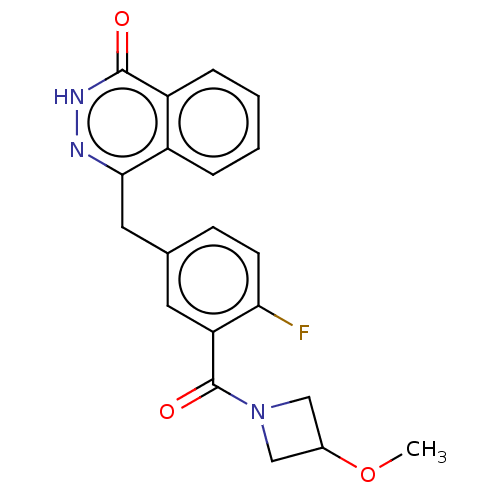
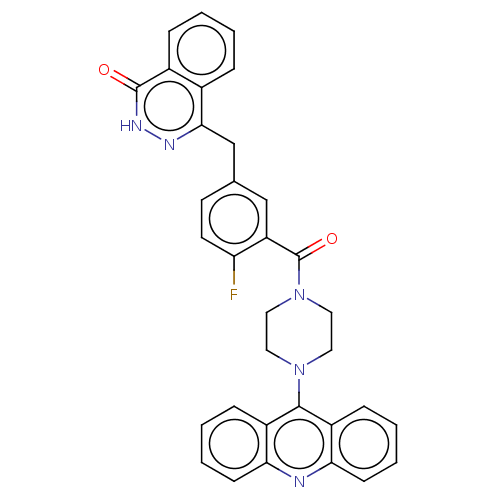
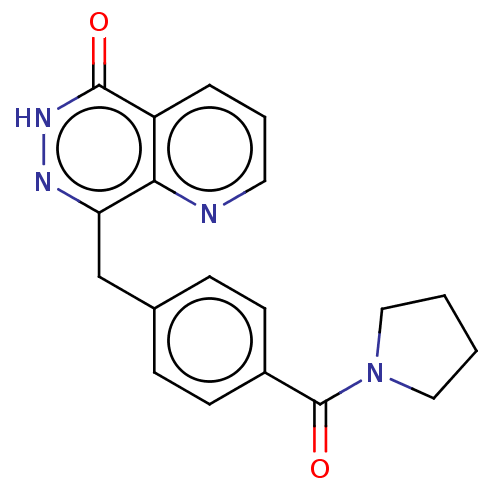
Report error Found 33 Enz. Inhib. hit(s) with all data for entry = 50015716
TargetHepatocyte growth factor receptor(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 0.0190nMAssay Description:Inhibition of human recombinant c-MET in presence of ATP by HTRF assayMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 0.0230nMAssay Description:Inhibition of VEGFR-2 (unknown origin)More data for this Ligand-Target Pair
TargetHepatocyte growth factor receptor(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 0.0350nMAssay Description:Inhibition of human recombinant c-MET in presence of ATP by HTRF assayMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 0.0600nMAssay Description:Inhibition of VEGFR2 (unknown origin) using ULight-4E-BP1 peptide as substrate incubated for 1 hr in presence of ATP by fluorescence based analysisMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 0.120nMAssay Description:Inhibition of VEGFR2 (unknown origin) using ULight-4E-BP1 peptide as substrate incubated for 1 hr in presence of ATP by fluorescence based analysisMore data for this Ligand-Target Pair
TargetEpidermal growth factor receptor(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 0.450nMAssay Description:Inhibition of EGFR (unknown origin)More data for this Ligand-Target Pair
TargetHepatocyte growth factor receptor(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 0.600nMAssay Description:Inhibition of c-MET (unknown origin) using FAM-labelled peptide as substrate incubated for 10 mins in presence of ATP by mobility shift assayMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 1.30nMAssay Description:Inhibition of VEGFR2 (unknown origin) using ULight-4E-BP1 peptide as substrate incubated for 1 hr in presence of ATP by fluorescence based analysisMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 1.40nMAssay Description:Inhibition of VEGFR2 (unknown origin) using ULight-4E-BP1 peptide as substrate incubated for 1 hr in presence of ATP by fluorescence based analysisMore data for this Ligand-Target Pair
TargetPoly [ADP-ribose] polymerase 1(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 2.5nMAssay Description:Inhibition of human full length PARP1 expressed in Escherichia coli rosetta (DE3) incubated for 20 mins by fluorescence analysisMore data for this Ligand-Target Pair
TargetPoly [ADP-ribose] polymerase 1(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 2.80nMAssay Description:Inhibition of human full length PARP1 expressed in Escherichia coli rosetta (DE3) incubated for 20 mins by fluorescence analysisMore data for this Ligand-Target Pair
TargetPoly [ADP-ribose] polymerase 1(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 2.80nMAssay Description:Displacement of [125I]KX1 from PARP1 in human OVCAR-8 cells incubated for 1 hr by wizard gamma counter analysisMore data for this Ligand-Target Pair
TargetPoly [ADP-ribose] polymerase 1(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 4.20nMAssay Description:Displacement of [125I]KX1 from PARP1 in human OVCAR-8 cells incubated for 1 hr by wizard gamma counter analysisMore data for this Ligand-Target Pair
TargetPoly [ADP-ribose] polymerase 1(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 4.70nMAssay Description:Displacement of [125I]KX1 from PARP1 in human OVCAR-8 cells incubated for 1 hr by wizard gamma counter analysisMore data for this Ligand-Target Pair
TargetPoly [ADP-ribose] polymerase 1(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 7nMAssay Description:Inhibition of PARP1 (unknown origin) by colorimetric assayMore data for this Ligand-Target Pair
TargetPoly [ADP-ribose] polymerase 1(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 8.20nMAssay Description:Displacement of [125I]KX1 from PARP1 in human OVCAR-8 cells incubated for 1 hr by wizard gamma counter analysisMore data for this Ligand-Target Pair
TargetPoly [ADP-ribose] polymerase 1(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 12nMAssay Description:Inhibition of PARP1 (unknown origin)More data for this Ligand-Target Pair
TargetPoly [ADP-ribose] polymerase 1(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 16nMAssay Description:Inhibition of PARP1 (unknown origin) by spectrophotometer analysisMore data for this Ligand-Target Pair
TargetTyrosine-protein kinase BTK(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 33nMAssay Description:Inhibition of BTK (unknown origin) incubated for 30 mins in presence of ATP by fluorescence based analysisMore data for this Ligand-Target Pair
TargetPoly [ADP-ribose] polymerase 1(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 34nMAssay Description:Inhibition of PARP1 (unknown origin) by colorimetric assayMore data for this Ligand-Target Pair
TargetPoly [ADP-ribose] polymerase 1(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 37nMAssay Description:Inhibition of PARP1 (unknown origin) by colorimetric assayMore data for this Ligand-Target Pair
TargetPoly [ADP-ribose] polymerase 1(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 44nMAssay Description:Displacement of [125I]KX1 from PARP1 in human OVCAR-8 cells incubated for 1 hr by wizard gamma counter analysisMore data for this Ligand-Target Pair
TargetPoly [ADP-ribose] polymerase 1(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 46nMAssay Description:Inhibition of PARP1 (unknown origin) incubated for 30 mins by ELISA assayMore data for this Ligand-Target Pair
TargetHistone deacetylase 6(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 57nMAssay Description:Inhibition of HDAC6 (unknown origin) measured after 15 mins by fluorescence based assayMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 61nMAssay Description:Inhibition of VEGFR2 (unknown origin) using ULight-4E-BP1 peptide as substrate incubated for 1 hr in presence of ATP by fluorescence based analysisMore data for this Ligand-Target Pair
TargetPoly [ADP-ribose] polymerase 1(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 97nMAssay Description:Inhibition of PARP1 (unknown origin) by colorimetric assayMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 110nMAssay Description:Inhibition of VEGFR2 (unknown origin) using poly (Glu,Tyr) 4:1 as substrate measured after 5 mins in presence of ATP by ELISA methodMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 310nMAssay Description:Inhibition of VEGFR2 (unknown origin) using poly (Glu,Tyr) 4:1 as substrate measured after 5 mins in presence of ATP by ELISA methodMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 910nMAssay Description:Inhibition of VEGFR2 (unknown origin) using poly (Glu,Tyr) 4:1 as substrate measured after 5 mins in presence of ATP by ELISA methodMore data for this Ligand-Target Pair
TargetCytochrome c oxidase subunit 2(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 1.40E+3nMAssay Description:Inhibition of COX-2 (unknown origin)More data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 2.50E+3nMAssay Description:Inhibition of VEGFR2 (unknown origin) using Poly (Glu, Tyr) substrate incubated for 40 mins in presence of ATP by ADP-Glo kinase assayMore data for this Ligand-Target Pair
TargetHepatocyte growth factor receptor(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 1.07E+5nMAssay Description:Inhibition of c-Met (unknown origin) incubated for 20 to 30 mins in presence of [33P]-ATP by scintillation counter analysisMore data for this Ligand-Target Pair
TargetCytochrome c oxidase subunit 1(Human)
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Key Laboratory of Technology of Drug Preparation (Zhengzhou University)
Curated by ChEMBL
Affinity DataIC50: 1.15E+5nMAssay Description:Inhibition of COX-1 (unknown origin)More data for this Ligand-Target Pair
 3D Structure (crystal)
3D Structure (crystal)