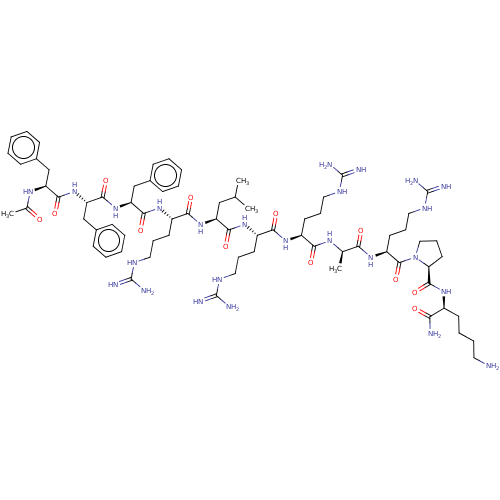
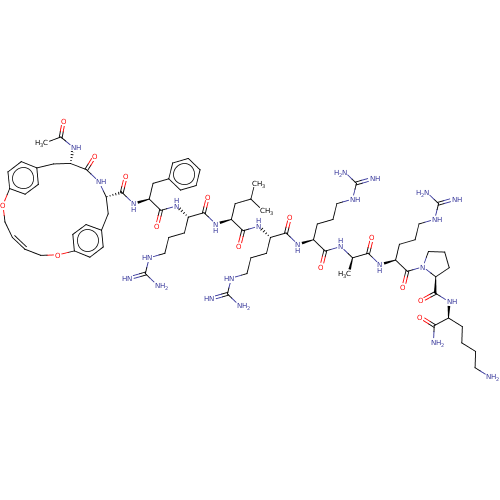
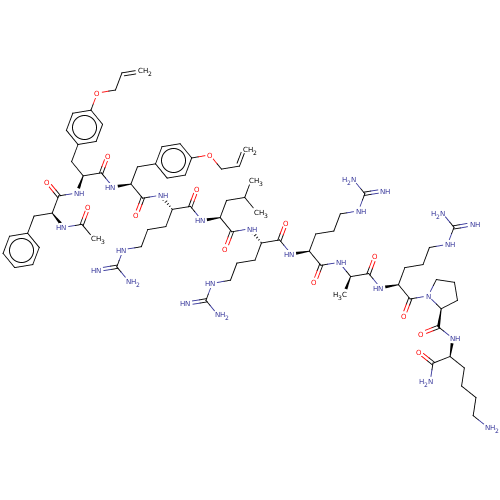
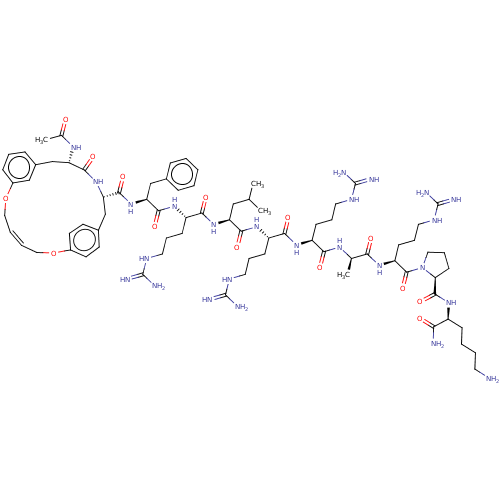
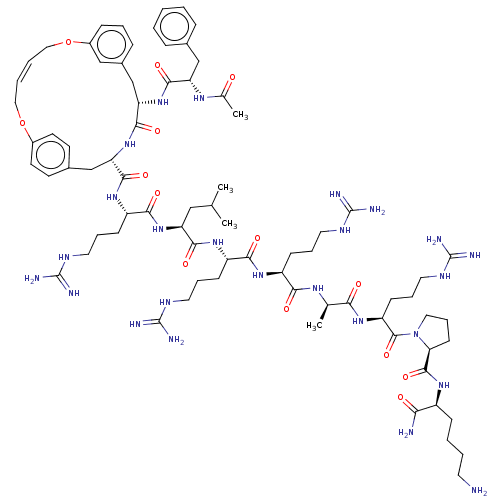
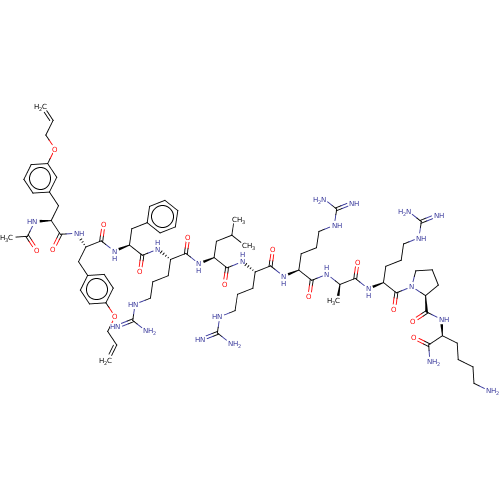
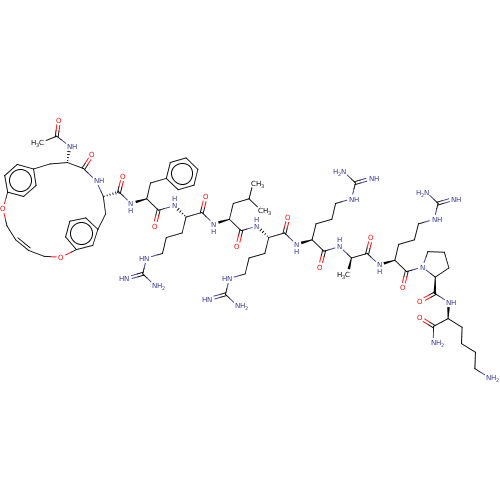
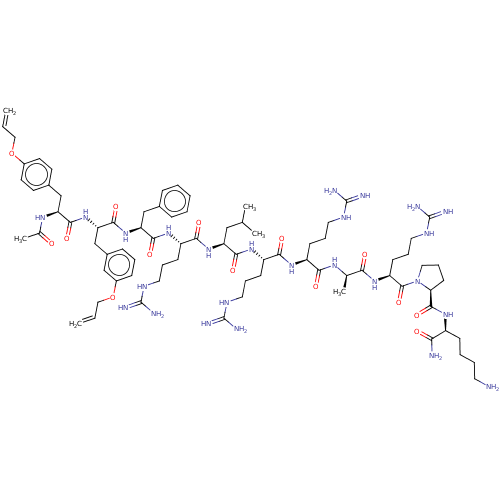
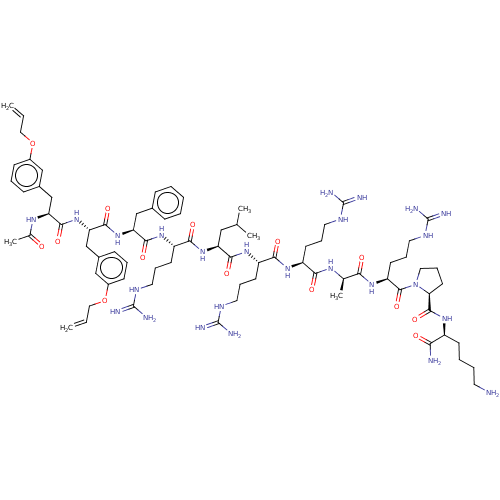
Report error Found 34 Enz. Inhib. hit(s) with all data for entry = 50015489
Affinity DataKi: 3.30nMAssay Description:Displacement of [3H]-Diprenorphine from kappa opiod receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase i...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Displacement of [3H]-Diprenorphine from kappa opiod receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase i...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Displacement of [3H]-Diprenorphine from kappa opiod receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase i...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Displacement of [3H]-Diprenorphine from kappa opiod receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase i...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Displacement of [3H]-Diprenorphine from kappa opiod receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase i...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Displacement of [3H]-Diprenorphine from kappa opiod receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase i...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Displacement of [3H]-Diprenorphine from kappa opiod receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase i...More data for this Ligand-Target Pair
Affinity DataKi: 19nMAssay Description:Displacement of [3H]-Diprenorphine from kappa opiod receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase i...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Displacement of [3H]-Diprenorphine from kappa opiod receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase i...More data for this Ligand-Target Pair
Affinity DataKi: 29nMAssay Description:Displacement of [3H]-Diprenorphine from kappa opiod receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase i...More data for this Ligand-Target Pair
Affinity DataKi: 39nMAssay Description:Displacement of [3H]-Diprenorphine from kappa opiod receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase i...More data for this Ligand-Target Pair
Affinity DataKi: 41nMAssay Description:Displacement of [3H]-Diprenorphine from kappa opiod receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase i...More data for this Ligand-Target Pair
Affinity DataKi: 43nMAssay Description:Displacement of [3H]-Diprenorphine from kappa opiod receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase i...More data for this Ligand-Target Pair
Affinity DataKi: 57nMAssay Description:Displacement of [3H]-Diprenorphine from kappa opiod receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase i...More data for this Ligand-Target Pair
Affinity DataKi: 62nMAssay Description:Displacement of [3H]-Diprenorphine from kappa opiod receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase i...More data for this Ligand-Target Pair
Affinity DataKi: 66nMAssay Description:Displacement of [3H]-Diprenorphine from kappa opiod receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase i...More data for this Ligand-Target Pair
Affinity DataKi: 68nMAssay Description:Displacement of [3H]-Diprenorphine from kappa opiod receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase i...More data for this Ligand-Target Pair
Affinity DataKi: 221nMAssay Description:Displacement of [3H]-DAMGO from mu opioid receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase inhibitorsMore data for this Ligand-Target Pair
Affinity DataKi: 478nMAssay Description:Displacement of [3H]-DAMGO from mu opioid receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase inhibitorsMore data for this Ligand-Target Pair
Affinity DataKi: 490nMAssay Description:Displacement of [3H]-DAMGO from mu opioid receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase inhibitorsMore data for this Ligand-Target Pair
Affinity DataKi: 491nMAssay Description:Displacement of [3H]-DAMGO from mu opioid receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase inhibitorsMore data for this Ligand-Target Pair
Affinity DataKi: 501nMAssay Description:Displacement of [3H]-DAMGO from mu opioid receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase inhibitorsMore data for this Ligand-Target Pair
Affinity DataKi: 607nMAssay Description:Displacement of [3H]-DAMGO from mu opioid receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase inhibitorsMore data for this Ligand-Target Pair
Affinity DataKi: 657nMAssay Description:Displacement of [3H]-DAMGO from mu opioid receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase inhibitorsMore data for this Ligand-Target Pair
Affinity DataKi: 828nMAssay Description:Displacement of [3H]-DAMGO from mu opioid receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase inhibitorsMore data for this Ligand-Target Pair
Affinity DataKi: 838nMAssay Description:Displacement of [3H]-DAMGO from mu opioid receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase inhibitorsMore data for this Ligand-Target Pair
Affinity DataKi: 1.22E+3nMAssay Description:Displacement of [3H]-DAMGO from mu opioid receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase inhibitorsMore data for this Ligand-Target Pair
Affinity DataKi: 1.33E+3nMAssay Description:Displacement of [3H]-DAMGO from mu opioid receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase inhibitorsMore data for this Ligand-Target Pair
Affinity DataKi: 1.37E+3nMAssay Description:Displacement of [3H]-DAMGO from mu opioid receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase inhibitorsMore data for this Ligand-Target Pair
Affinity DataKi: 1.38E+3nMAssay Description:Displacement of [3H]-DAMGO from mu opioid receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase inhibitorsMore data for this Ligand-Target Pair
Affinity DataKi: 1.43E+3nMAssay Description:Displacement of [3H]-DAMGO from mu opioid receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase inhibitorsMore data for this Ligand-Target Pair
Affinity DataKi: 1.63E+3nMAssay Description:Displacement of [3H]-DAMGO from mu opioid receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase inhibitorsMore data for this Ligand-Target Pair
Affinity DataKi: 1.74E+3nMAssay Description:Displacement of [3H]-DAMGO from mu opioid receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase inhibitorsMore data for this Ligand-Target Pair
Affinity DataKi: 1.96E+3nMAssay Description:Displacement of [3H]-DAMGO from mu opioid receptor (unknown origin) expressed in CHO cells incubated for 90 mins in presence of peptidase inhibitorsMore data for this Ligand-Target Pair