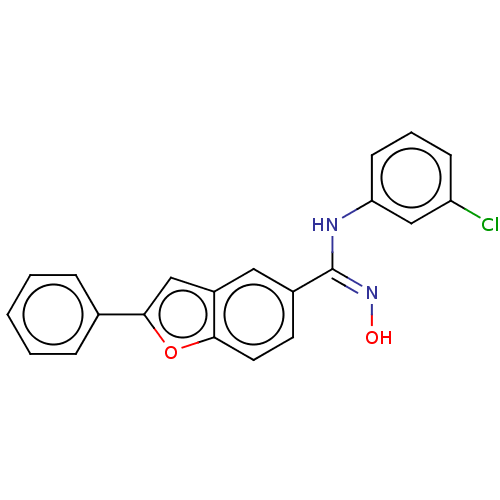
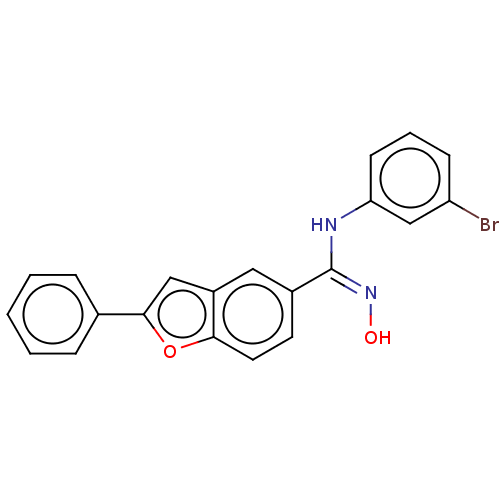
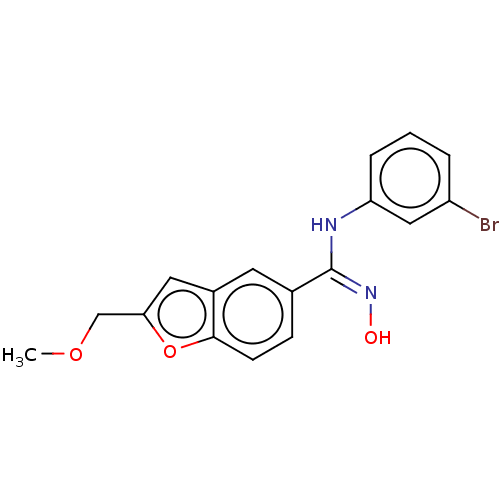
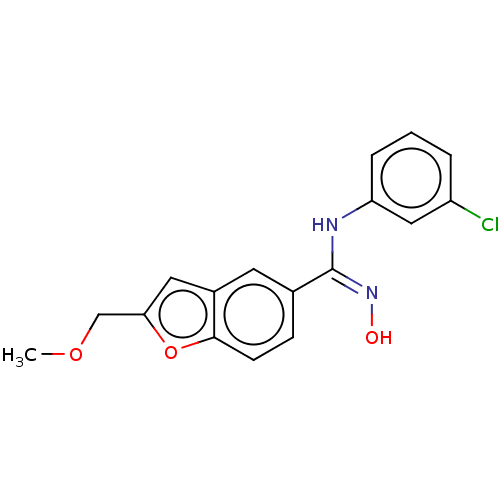
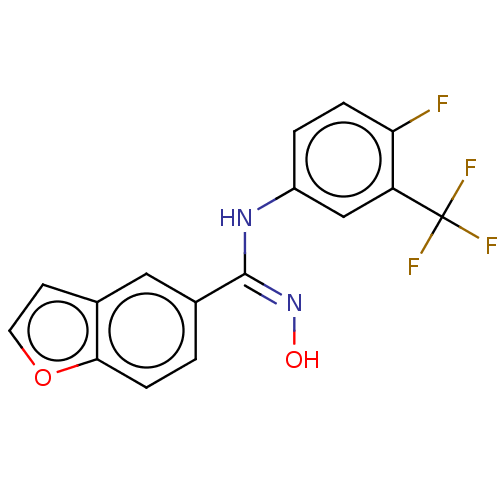
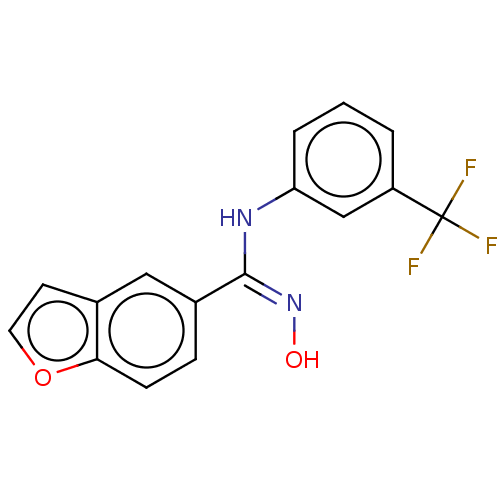
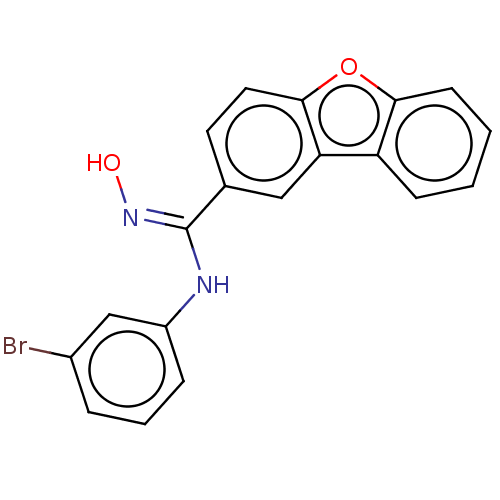
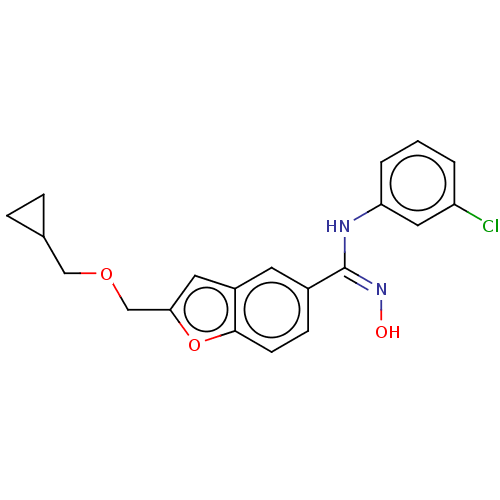
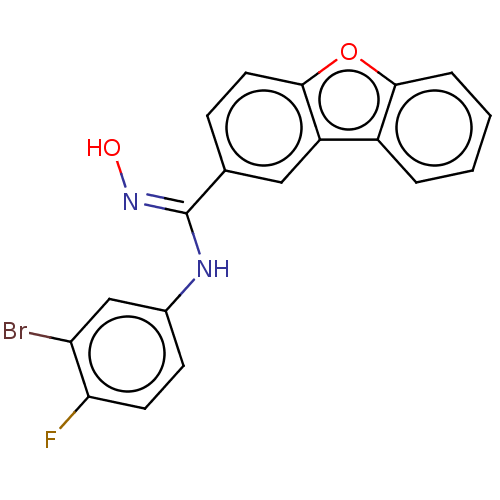
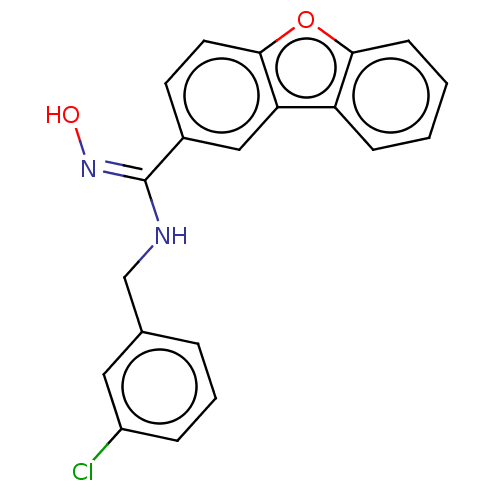
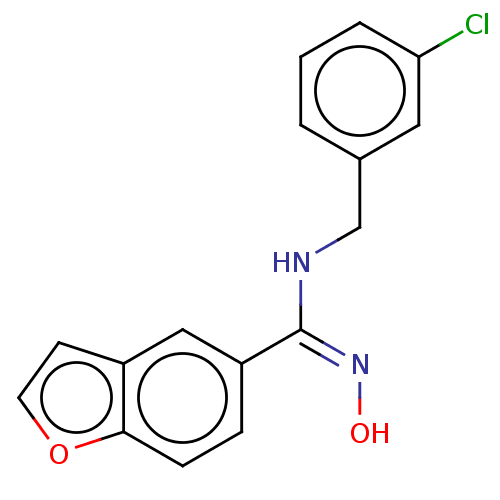
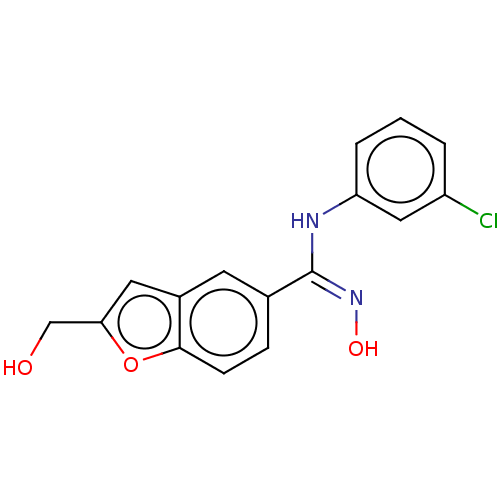
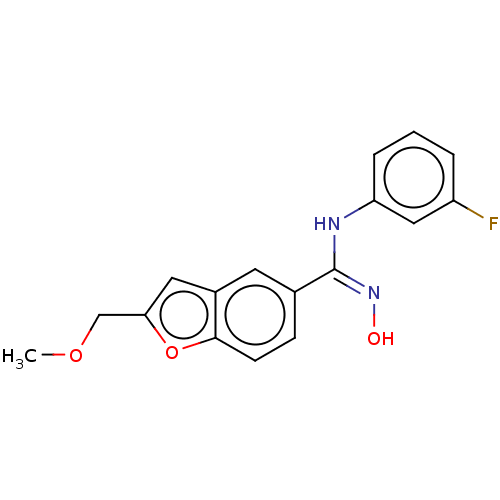
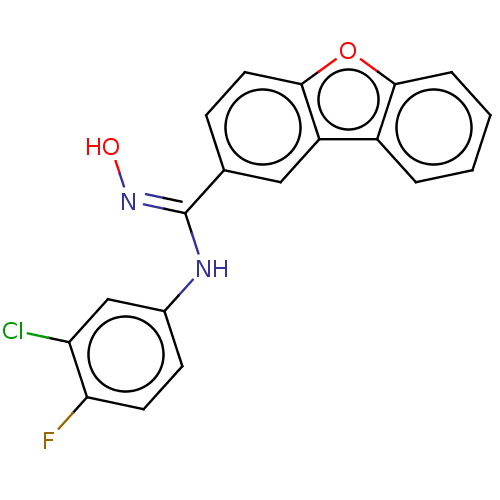
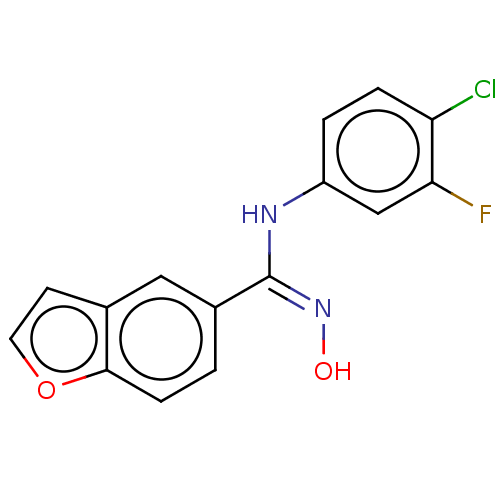
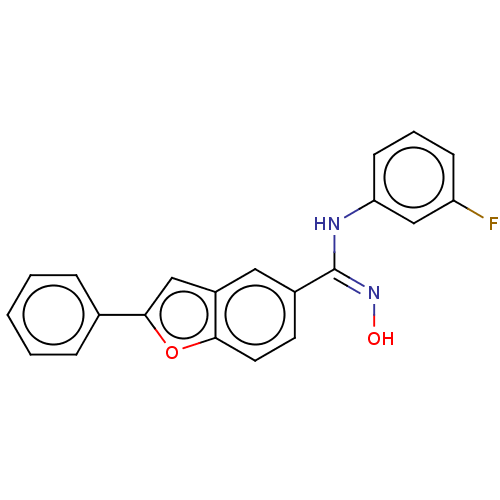
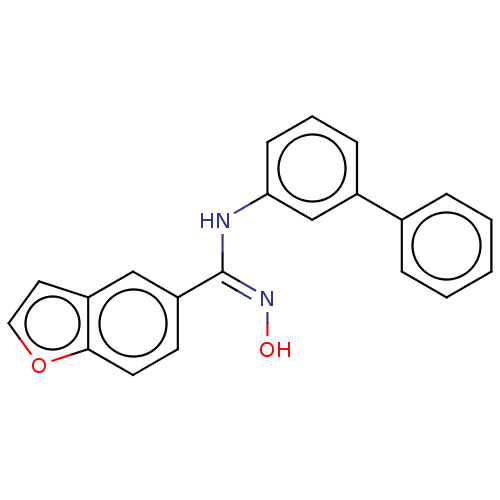
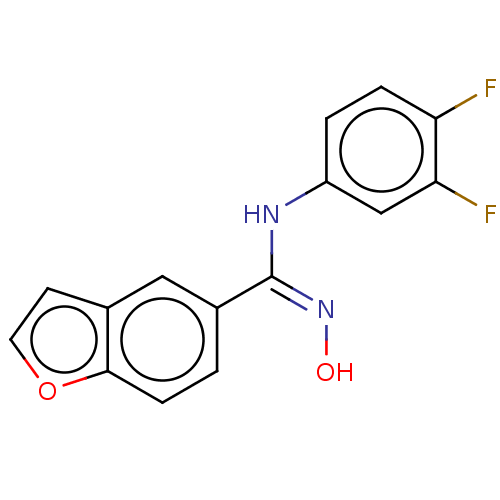
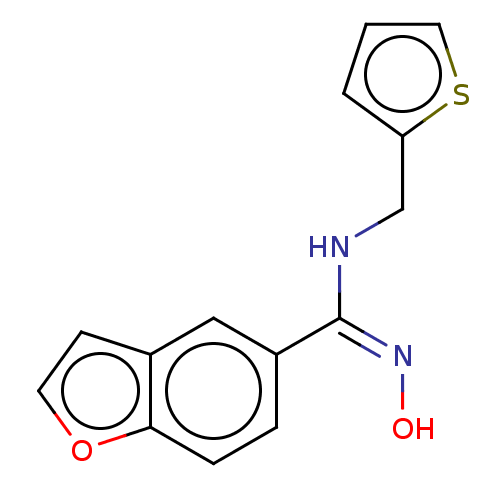
Report error Found 90 Enz. Inhib. hit(s) with all data for entry = 50013876
Affinity DataIC50: 400nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 440nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 690nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 710nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 900nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 900nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 930nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+3nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+3nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+3nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+3nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 2.60E+3nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+3nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 3.60E+3nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 5.20E+3nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 5.80E+3nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 5.90E+3nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 6.10E+3nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 7.10E+3nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 7.60E+3nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+3nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+3nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 8.60E+3nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 9.60E+3nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.03E+4nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.06E+4nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.08E+4nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.08E+4nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+4nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+4nMAssay Description:Inhibition of IDO1 (unknown origin) assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbance based analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.19E+4nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+4nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.62E+4nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.62E+4nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.76E+4nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells assessed as reduction in kynurenine formation using L-tryptophan as substrate by absorbanc...More data for this Ligand-Target Pair