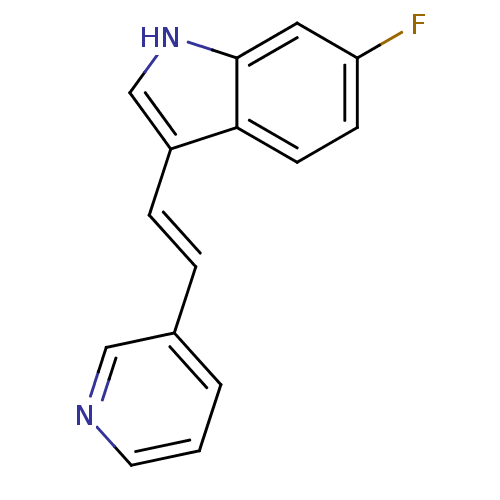
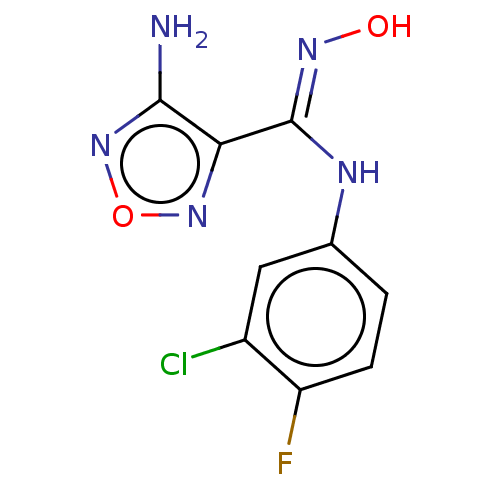
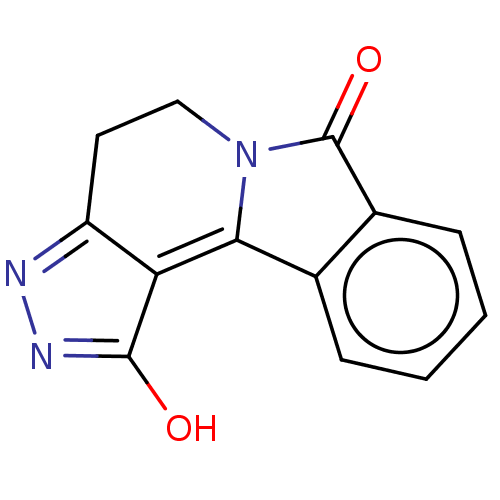
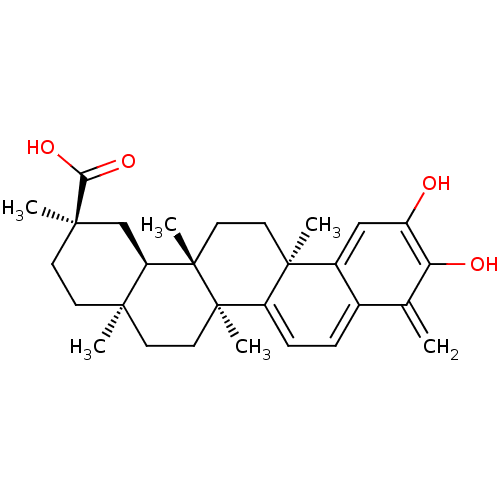
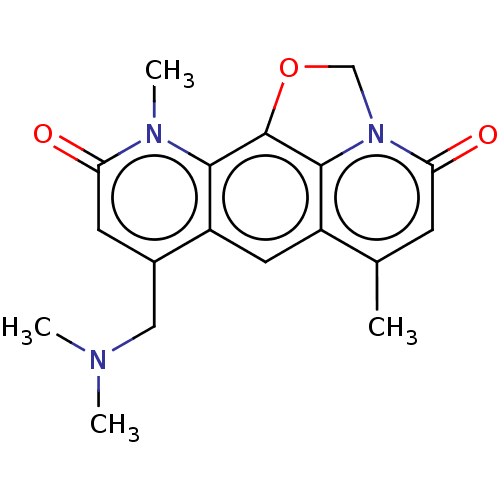
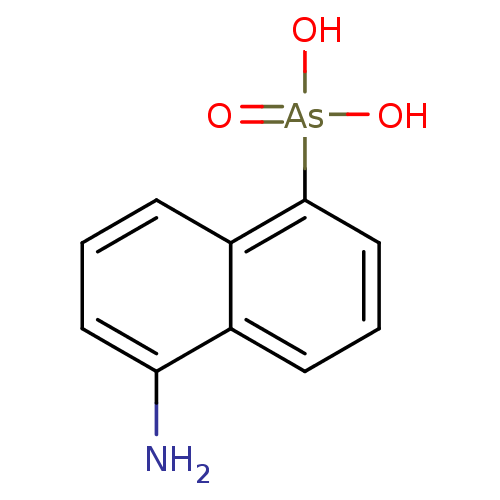
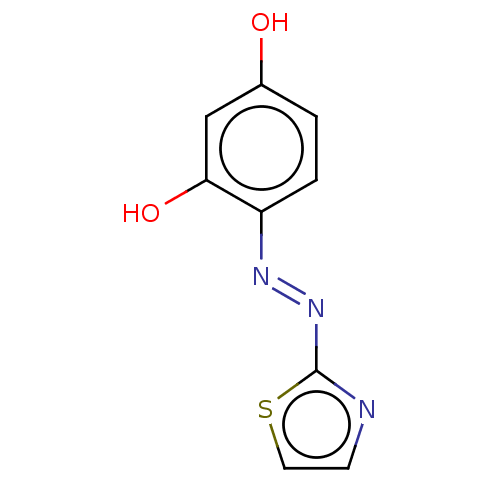
Report error Found 50 Enz. Inhib. hit(s) with all data for entry = 50013607
Affinity DataIC50: 30nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Inhibition of recombinant human IDO1 assessed as reduction in N-formylkynurenine formation using L-tryptophan as substrate incubated for 45 mins by m...More data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Inhibition of TDO2 in rat liver homogenate assessed as reduction in N-formylkynurenine formation using L-tryptophan as substrate incubated for 1 hrMore data for this Ligand-Target Pair
Affinity DataIC50: 60nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 80nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 110nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 110nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 310nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 1.39E+3nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.72E+3nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.12E+3nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 2.79E+3nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.43E+3nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 3.47E+3nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 3.62E+3nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 3.78E+3nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.35E+3nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 4.37E+3nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.41E+3nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.43E+3nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 9.17E+3nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.11E+4nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.13E+4nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 1.18E+4nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.23E+4nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 1.46E+4nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.57E+4nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 1.77E+4nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 1.81E+4nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.23E+4nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 2.27E+4nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 2.38E+4nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+4nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.02E+4nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 3.37E+4nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 3.79E+4nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 4.33E+4nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 4.41E+4nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.50E+4nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.57E+4nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.86E+4nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of recombinant human TDO2 expressed in Escherichia coli BL21 (DE3) assessed as reduction in N-formylkynurenine formation using L-tryptopha...More data for this Ligand-Target Pair
Affinity DataIC50: 3.50E+5nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.78E+5nMAssay Description:Inhibition of recombinant human IDO1 expressed in Escherichia coli EC538 assessed as reduction in N-formylkynurenine formation using L-tryptophan as ...More data for this Ligand-Target Pair
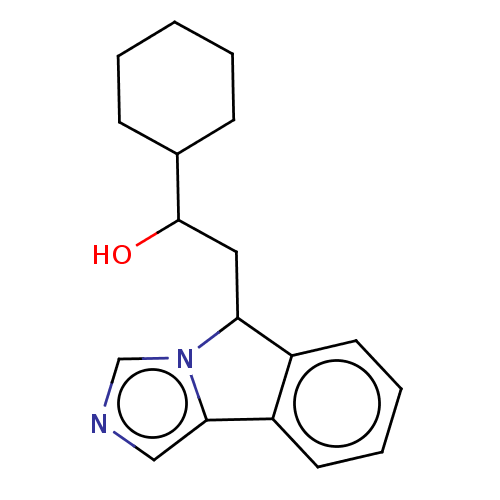
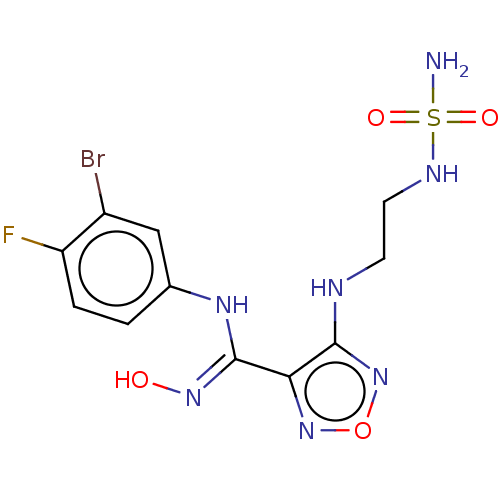
 3D Structure (crystal)
3D Structure (crystal)