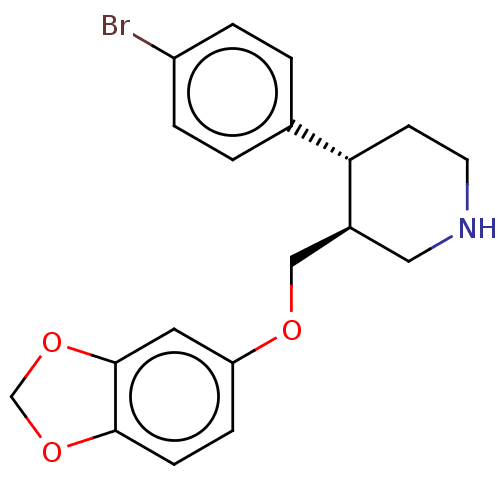
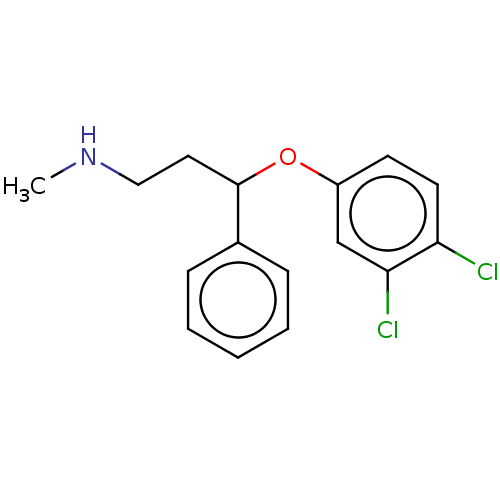
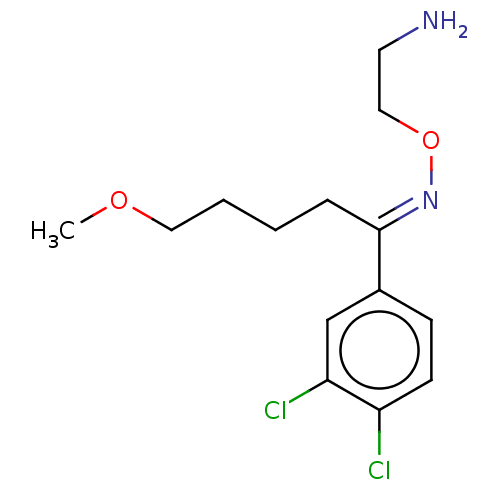
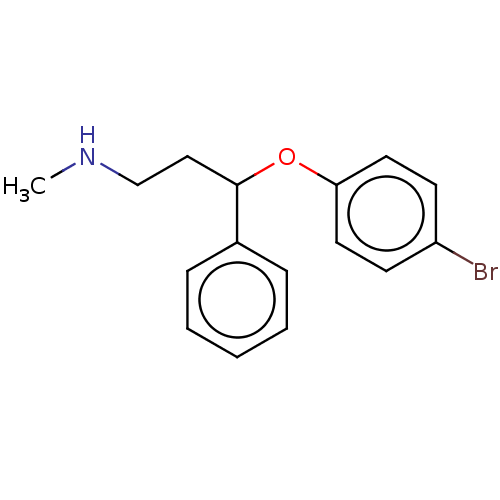
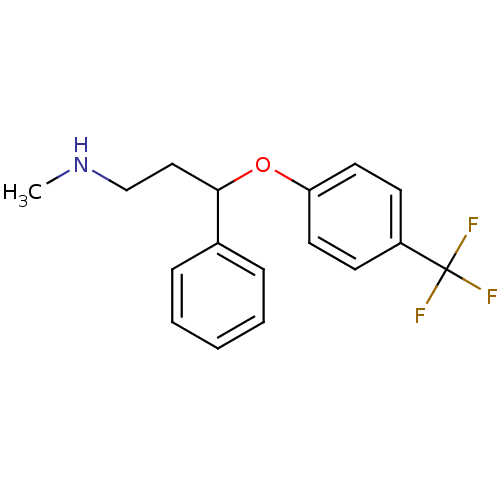
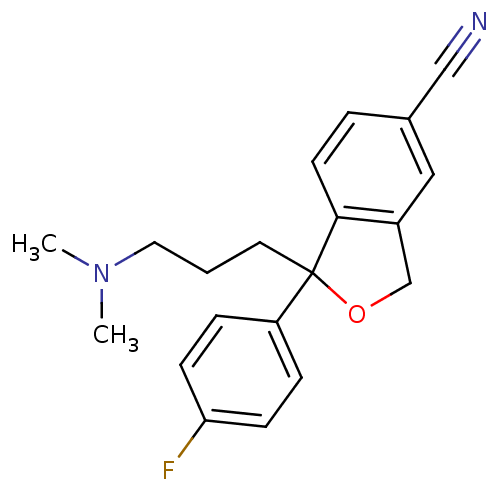
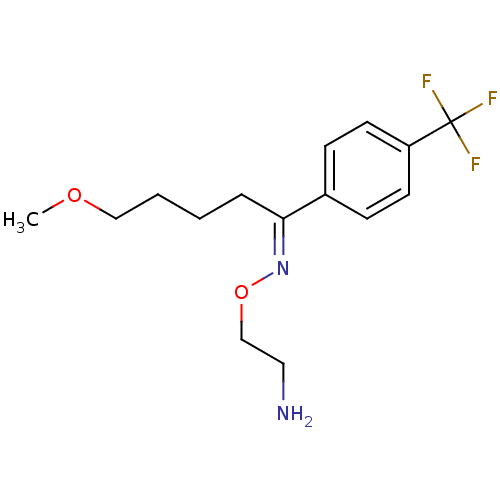
Report error Found 33 Enz. Inhib. hit(s) with all data for entry = 50014665
Affinity DataKi: 0.0800nMAssay Description:Displacement of [125I]RTI55 binding from human wild type SERTMore data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Displacement of [125I]RTI55 binding from human wild type SERTMore data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Inhibition of human wild type SERT expressed in COS7 cells assessed as inhibition of [3H]5HT uptake measured for 40 hrs by scintillation counting ana...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataKi: 19nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataKi: 21nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataKi: 24nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataKi: 31nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataKi: 31nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataKi: 41nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 43nMAssay Description:Inhibition of human wild type SERT expressed in COS7 cells assessed as inhibition of [3H]5HT uptake measured for 40 hrs by scintillation counting ana...More data for this Ligand-Target Pair
Affinity DataKi: 52nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataKi: 55nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 59nMAssay Description:Inhibition of human wild type SERT expressed in COS7 cells assessed as inhibition of [3H]5HT uptake measured for 40 hrs by scintillation counting ana...More data for this Ligand-Target Pair
Affinity DataKi: 61nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataKi: 62nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataKi: 66nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 135nMAssay Description:Inhibition of human wild type SERT expressed in COS7 cells assessed as inhibition of [3H]5HT uptake measured for 40 hrs by scintillation counting ana...More data for this Ligand-Target Pair
Affinity DataKi: 137nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataKi: 202nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataKi: 349nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataKi: 458nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataKi: 543nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 881nMAssay Description:Inhibition of human wild type SERT expressed in COS7 cells assessed as inhibition of [3H]5HT uptake measured for 40 hrs by scintillation counting ana...More data for this Ligand-Target Pair
Affinity DataKi: 1.46E+3nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataKi: 1.62E+3nMAssay Description:Displacement of [3H]citalopram from human SERT in HEK293 cells by Topcount scintillation analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 5.66E+3nMAssay Description:Inhibition of human wild type SERT expressed in COS7 cells assessed as inhibition of [3H]5HT uptake measured for 40 hrs by scintillation counting ana...More data for this Ligand-Target Pair
Affinity DataIC50: 5.66E+3nMAssay Description:Inhibition of human wild type SERT expressed in COS7 cells assessed as inhibition of [3H]5HT uptake measured for 40 hrs by scintillation counting ana...More data for this Ligand-Target Pair
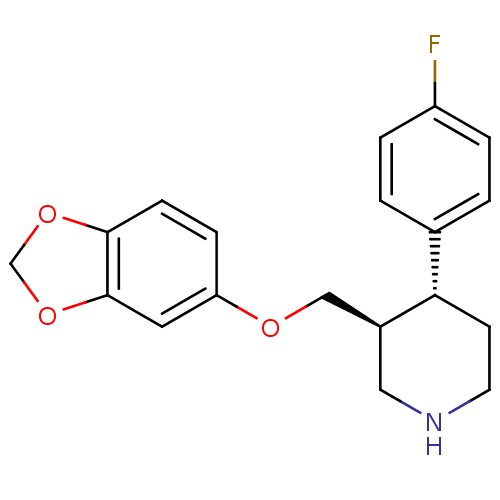
 3D Structure (crystal)
3D Structure (crystal)