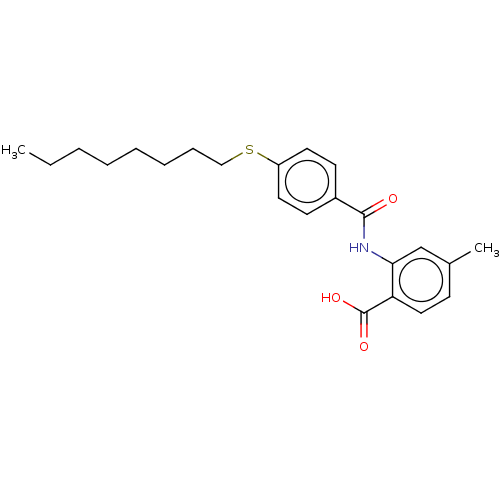
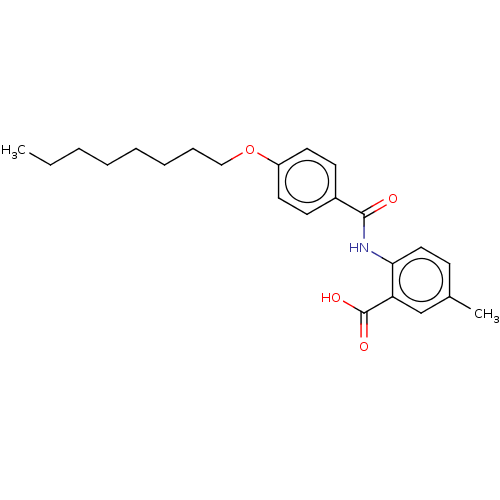
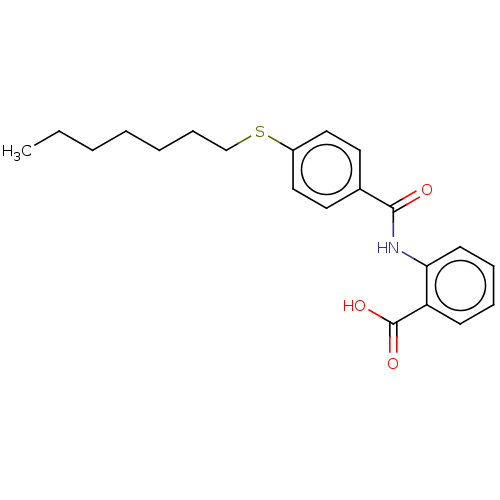
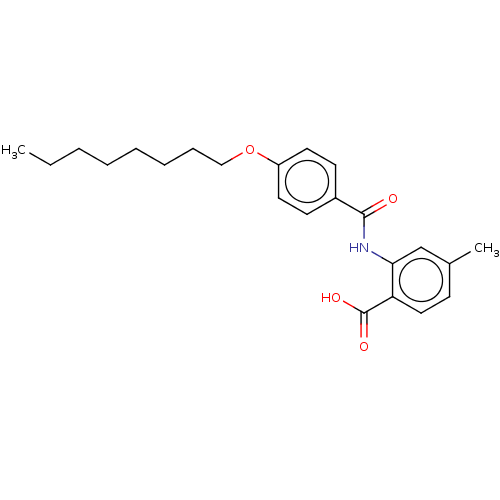
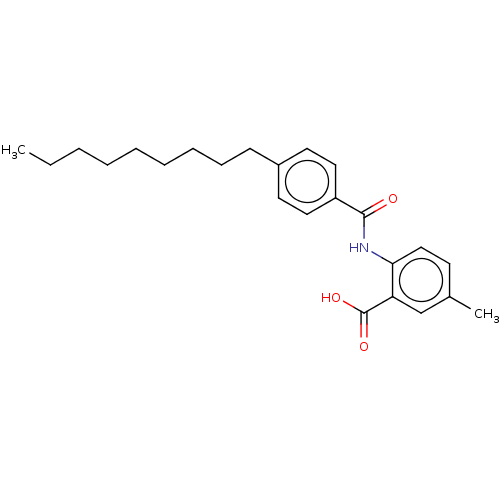
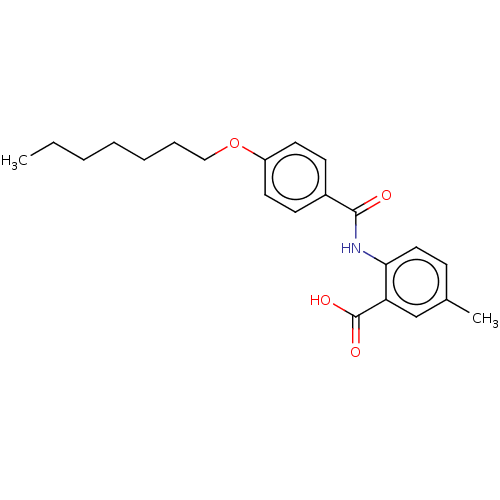
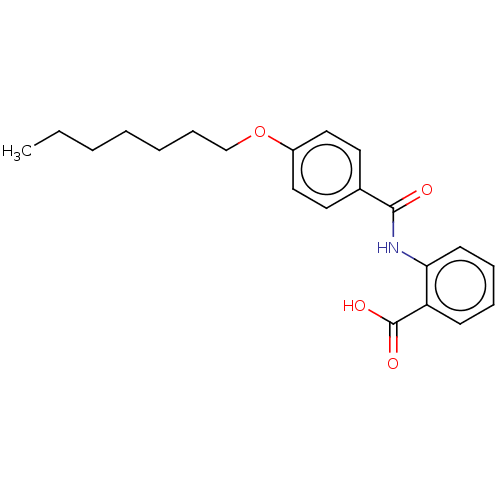
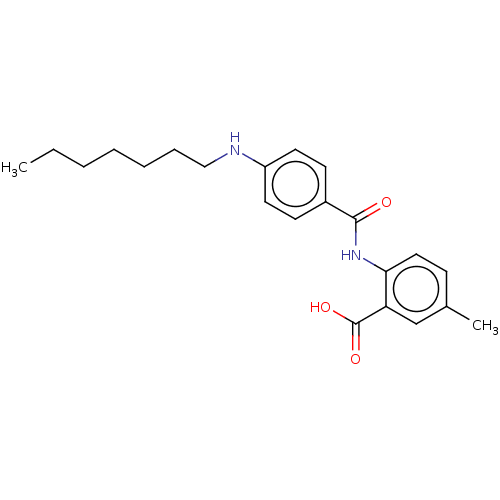
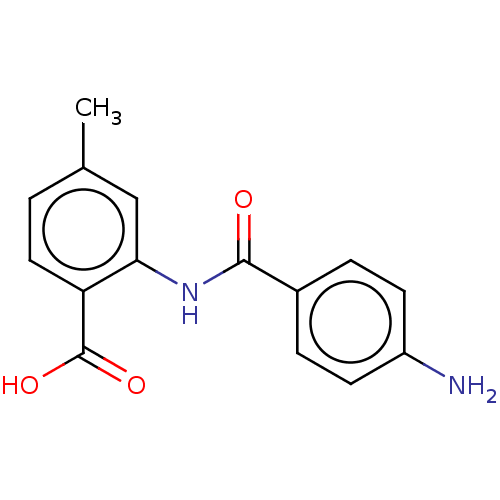
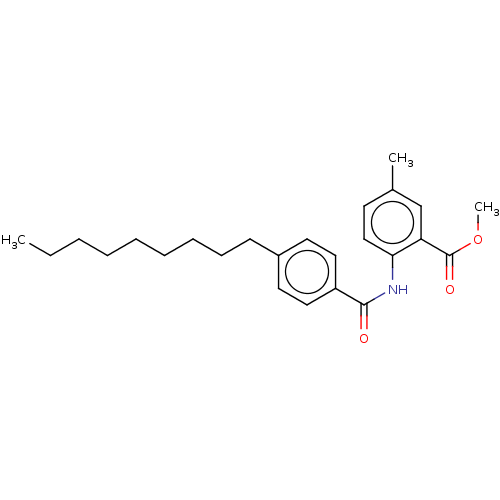
Report error Found 23 Enz. Inhib. hit(s) with all data for entry = 50015014
Affinity DataEC50: 650nMAssay Description:Binding affinity to PLK1 in human PC-3 cells incubated for 2 hrs followed by heat shock for 10 mins by cellular thermal shift assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.34E+4nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 2.35E+4nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 2.38E+4nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 2.55E+4nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 2.61E+4nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 2.68E+4nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 2.71E+4nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+4nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 2.81E+4nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 2.86E+4nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 2.90E+4nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 2.98E+4nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 3.38E+4nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 3.44E+4nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 3.70E+4nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 4.21E+4nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 6.09E+4nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 7.32E+4nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 8.74E+4nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+5nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+5nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+5nMAssay Description:Binding affinity to full length PLK1 (367 to 603 residues) (unknown origin) using MAGPMQS[pT]PLNGAKK tracer peptide as substrate by fluorescence pola...More data for this Ligand-Target Pair