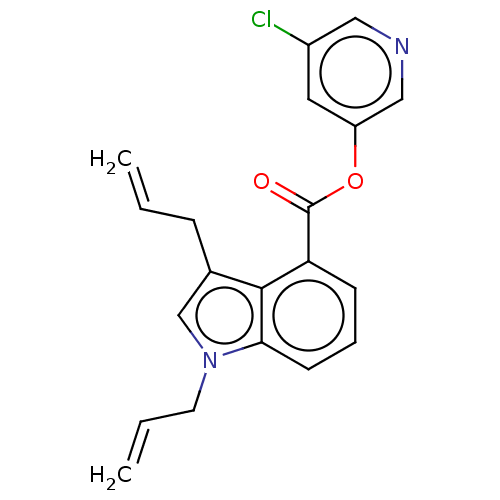
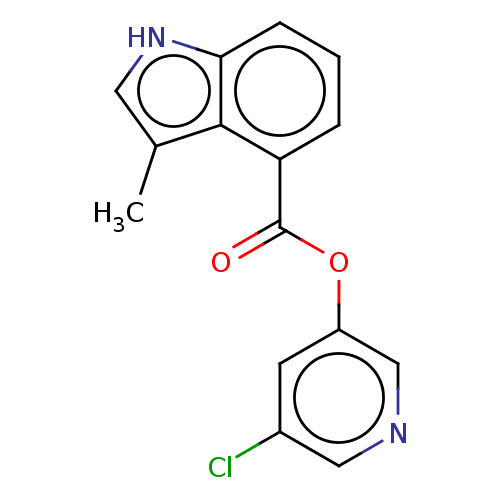
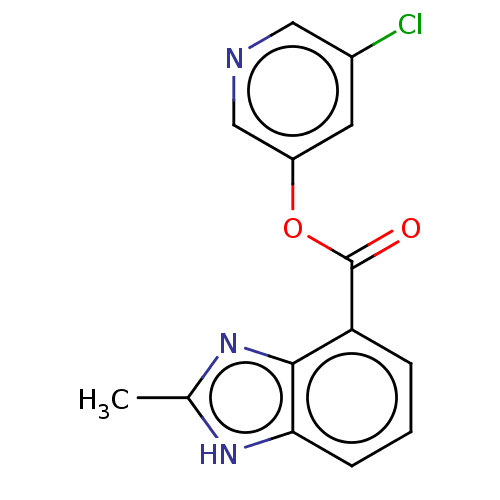
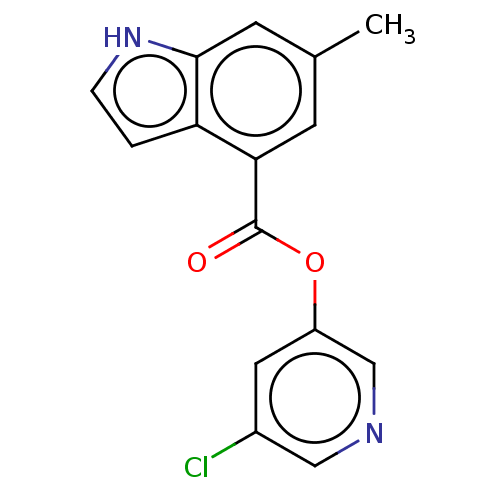
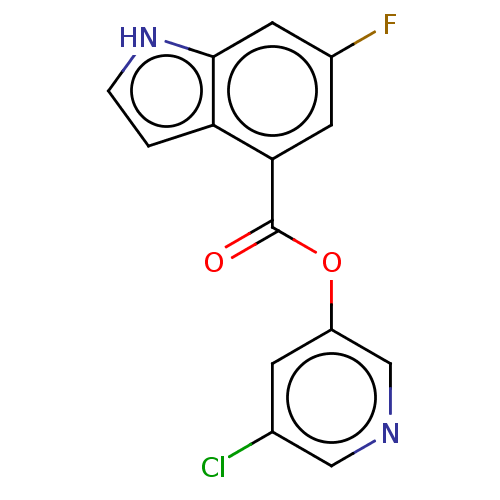
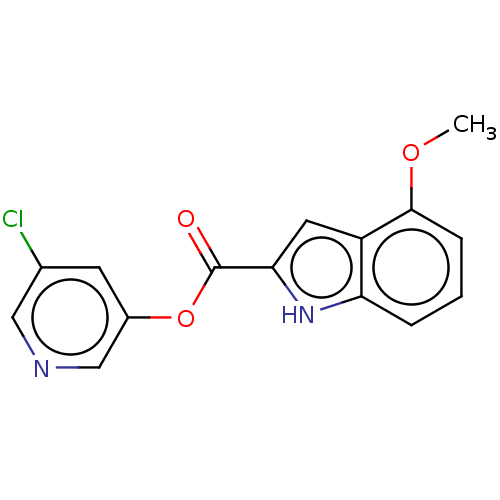
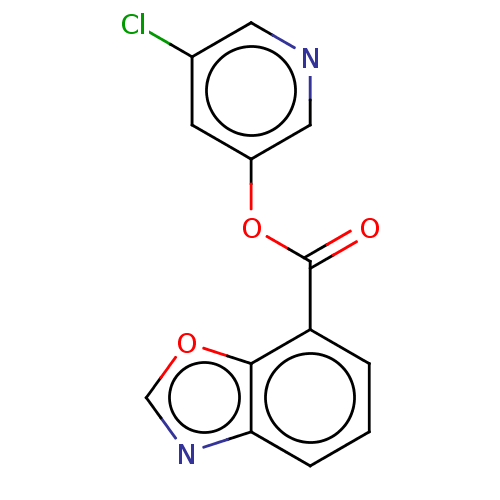
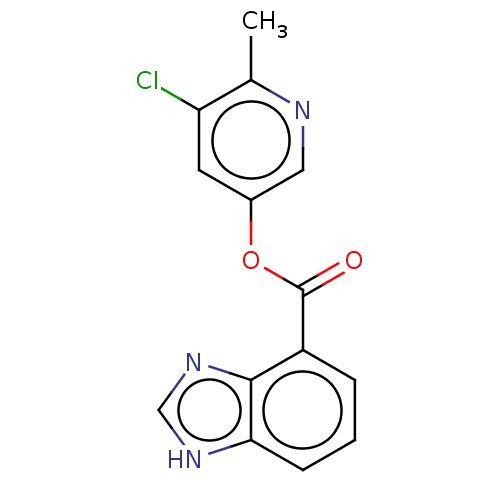
Report error Found 44 Enz. Inhib. hit(s) with all data for entry = 9879
Affinity DataKi: 18nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataIC50: 73nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataIC50: 250nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataIC50: 310nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataIC50: 320nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataIC50: 340nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataIC50: 380nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataIC50: 419nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataIC50: 470nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataIC50: 489nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataIC50: 590nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataIC50: 870nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataIC50: 900nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.19E+3nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.22E+3nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataEC50: 2.80E+3nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataEC50: 3.10E+3nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataEC50: 4.20E+3nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.43E+3nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataEC50: 8.10E+3nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.03E+4nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.15E+4nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.40E+4nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.50E+4nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.50E+4nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.53E+4nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.93E+4nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataEC50: 3.00E+4nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataEC50: 4.37E+4nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataEC50: 5.67E+4nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataEC50: 5.70E+4nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataEC50: 5.71E+4nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataEC50: 6.98E+4nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+5nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+5nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.00E+5nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+5nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.00E+5nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:IC50 values were determined for compounds that covalently inhibit SARS-CoV-2 3CLpro using our recently described assay and data fitting methods that ...More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+5nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+5nMAssay Description:VeroE6 cells and TMPRSS2-overexpressing VeroE6 (VeroE6TMPRSS2) cells were obtained from the Japanese Collection of Research Bioresources (JCRB) Cell ...More data for this Ligand-Target Pair