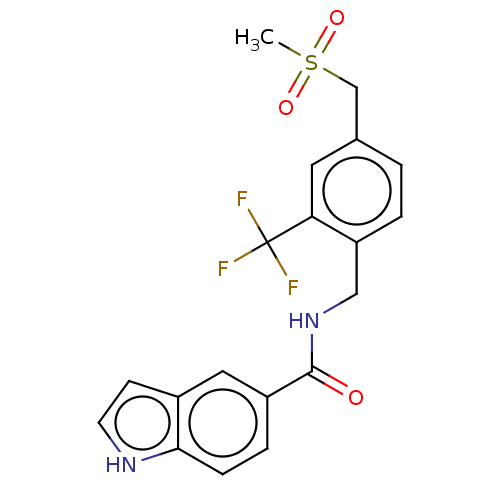
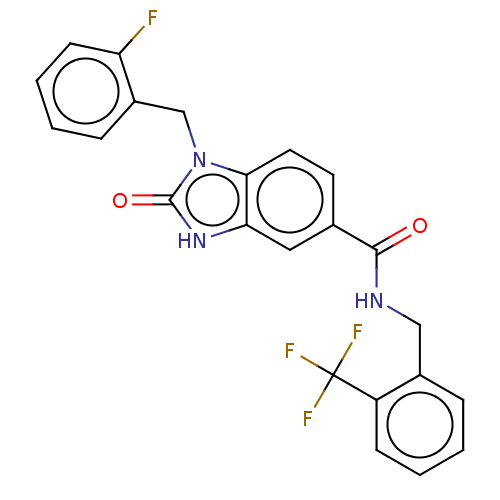
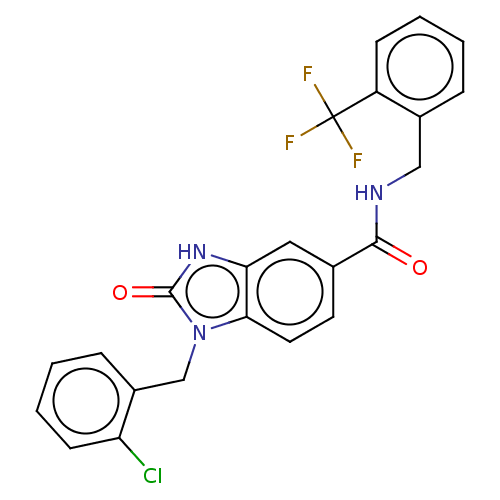
Report error Found 105 Enz. Inhib. hit(s) with all data for entry = 50015470
Affinity DataIC50: 2nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of N-terminal His-tagged mouse sEH (2 to 554 residues) expressed in Escherichia coli Rosetta2 (DE3) assessed as reduction in 6-methoxy-2-n...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 28nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 28nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 29nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 29nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 35nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 36nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 43nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 46nMAssay Description:Inhibition of N-terminal His-tagged mouse sEH (2 to 554 residues) expressed in Escherichia coli Rosetta2 (DE3) assessed as reduction in 6-methoxy-2-n...More data for this Ligand-Target Pair
Affinity DataIC50: 48nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 55nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 82nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 87nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataEC50: 100nMAssay Description:Partial agonist activity at sGSF-PPARgamma (unknown origin) assessed as CBP-1 recruitment by HT-FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 107nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataEC50: 120nMAssay Description:Partial agonist activity at GAL4-tagged human PPARgamma LBD expressed in HEK293T cells incubated for 12 to 14 hrs by dual-Glo luciferase assayMore data for this Ligand-Target Pair
Affinity DataIC50: 142nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataEC50: 150nMAssay Description:Partial agonist activity at GAL4-tagged human PPARgamma LBD expressed in HEK293T cells incubated for 12 to 14 hrs by dual-Glo luciferase assayMore data for this Ligand-Target Pair
Affinity DataIC50: 153nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 157nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 171nMAssay Description:Inhibition of recombinant human full length sEH assessed as reduction in 6-methoxy-2-naphthaldehyde formation using PHOME as substrate preincubated f...More data for this Ligand-Target Pair
Affinity DataIC50: 176nMAssay Description:Inhibition of N-terminal His-tagged mouse sEH (2 to 554 residues) expressed in Escherichia coli Rosetta2 (DE3) assessed as reduction in 6-methoxy-2-n...More data for this Ligand-Target Pair
Affinity DataEC50: 180nMAssay Description:Partial agonist activity at sGSF-PPARgamma (unknown origin) assessed as CBP-1 recruitment by HT-FRET assayMore data for this Ligand-Target Pair
Affinity DataEC50: 180nMAssay Description:Partial agonist activity at sGSF-PPARgamma (unknown origin) assessed as CBP-1 recruitment by HT-FRET assayMore data for this Ligand-Target Pair
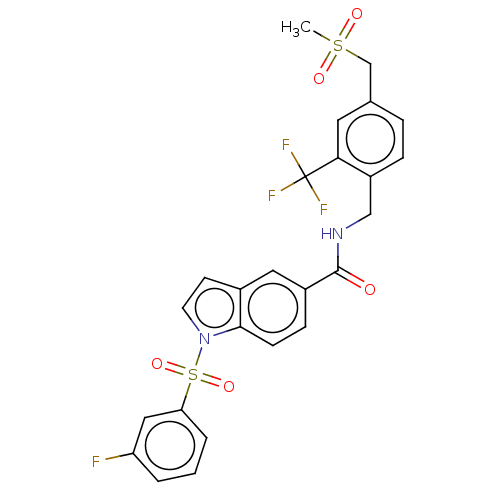
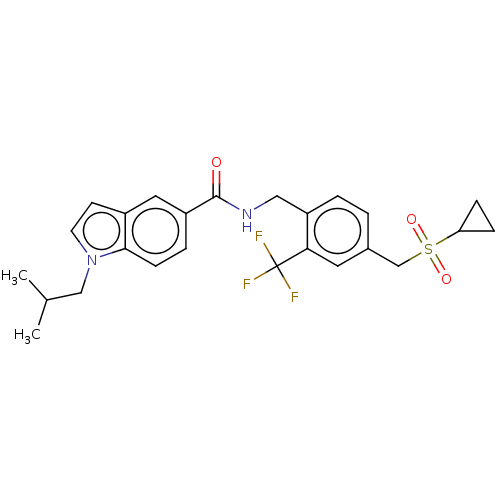

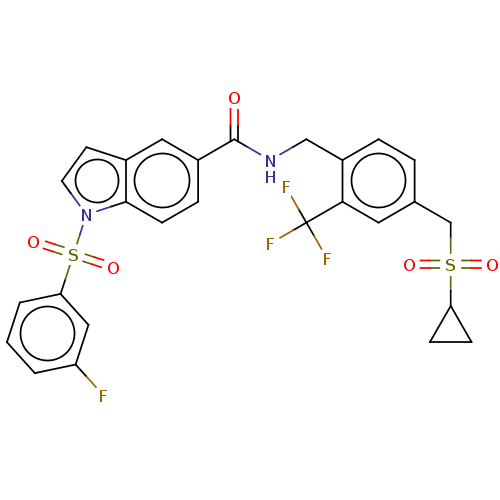
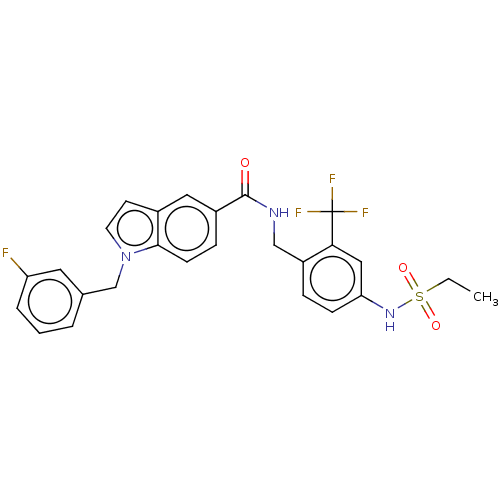
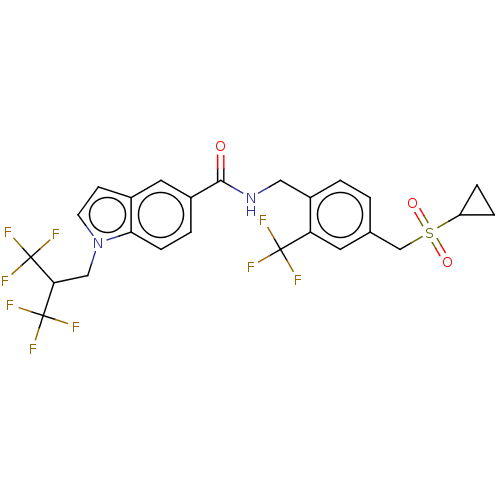

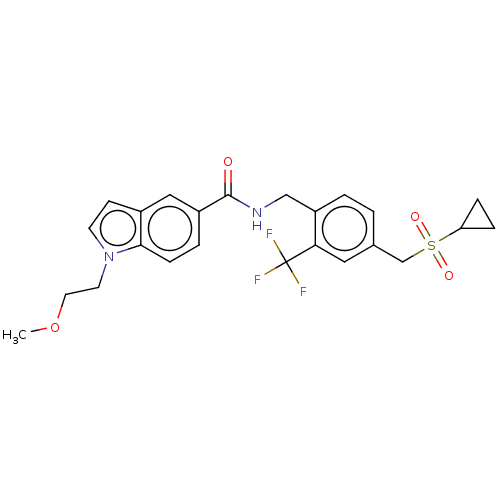
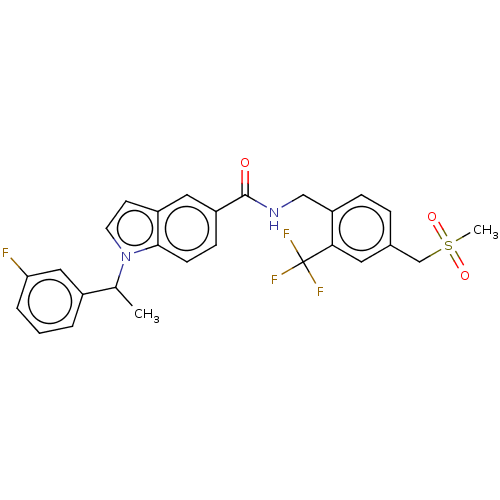

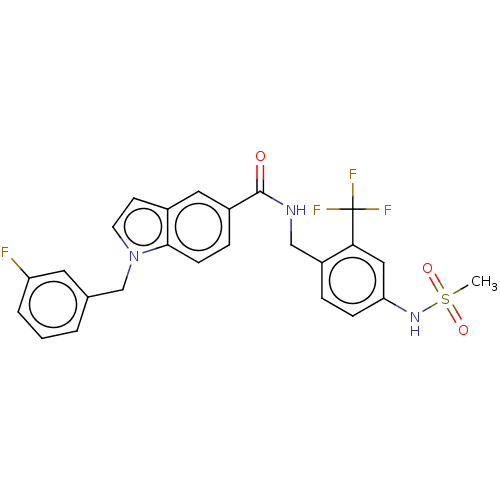
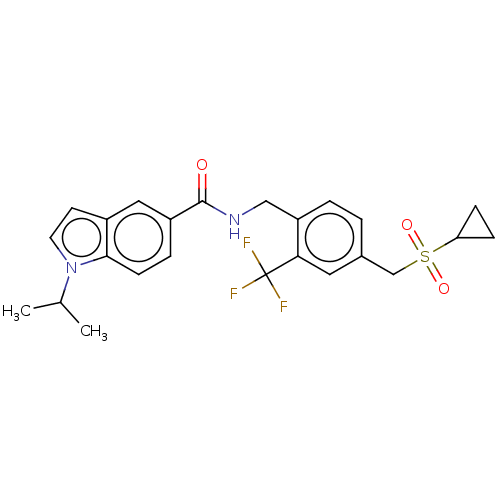
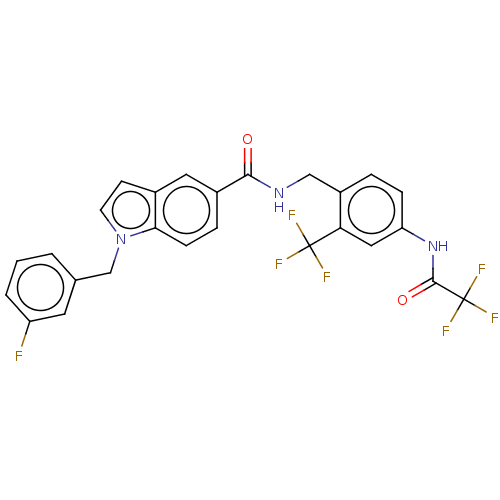
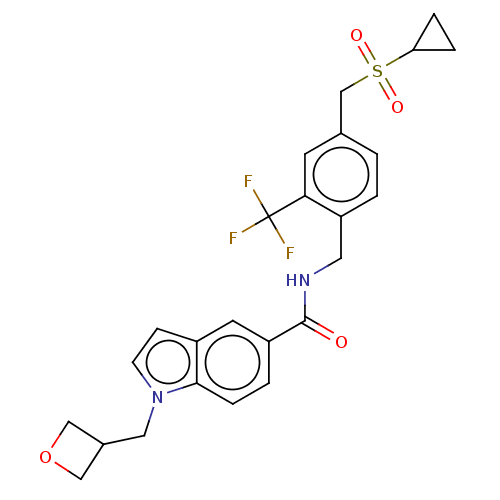
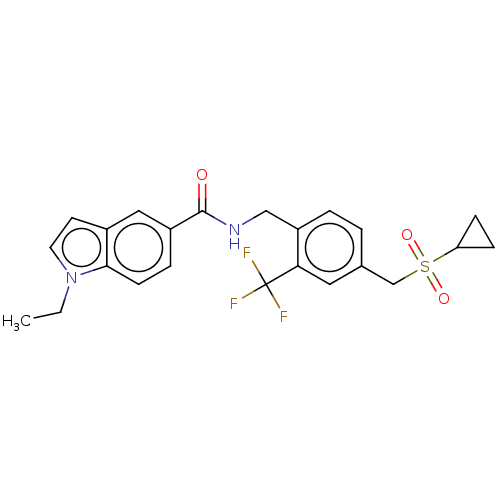
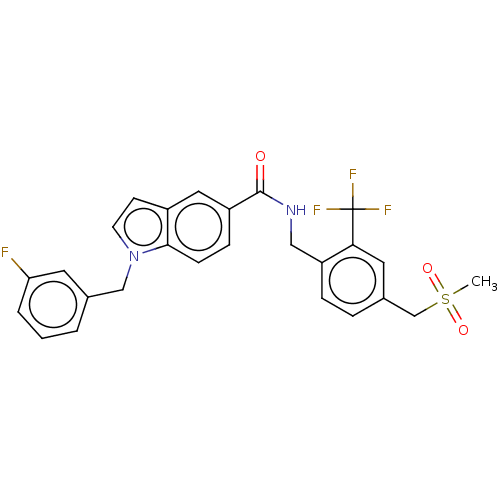
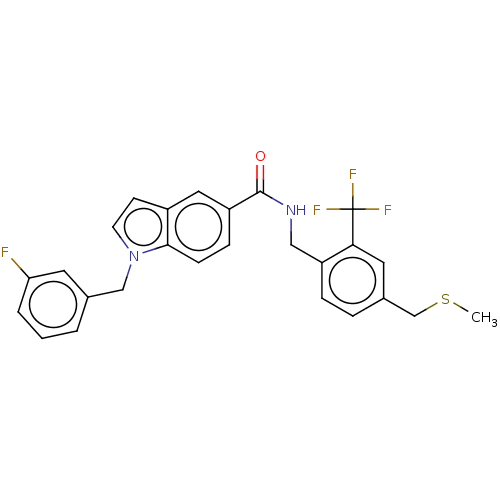

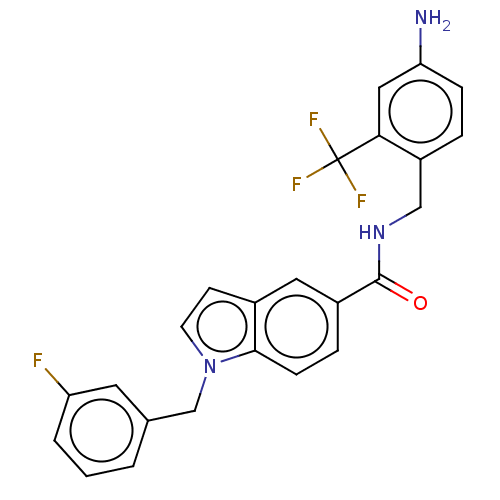
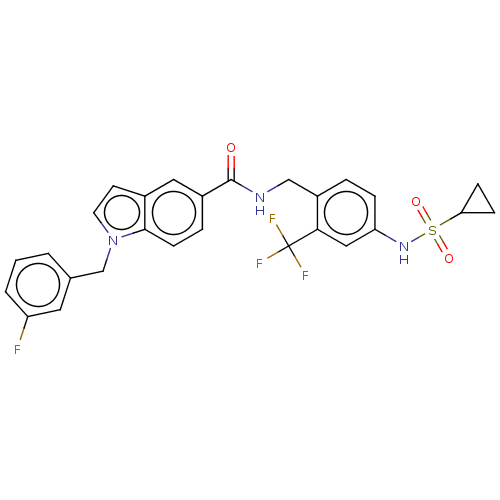
 3D Structure (crystal)
3D Structure (crystal)