Report error Found 14 Enz. Inhib. hit(s) with all data for entry = 50015183
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Second Military Medical University
Curated by ChEMBL
Second Military Medical University
Curated by ChEMBL
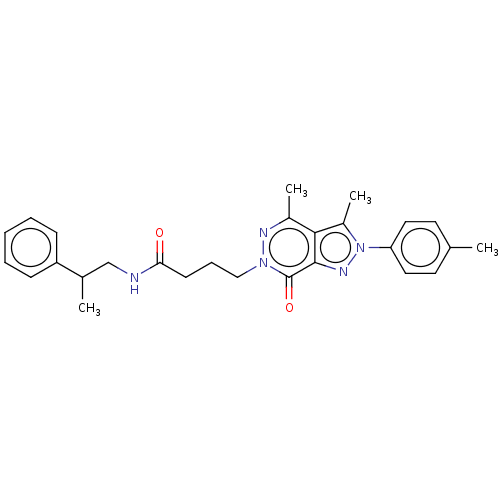
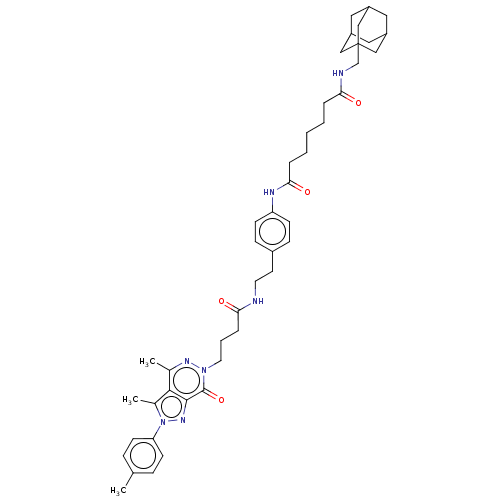
Affinity DataKi: 8nMAssay Description:Binding affinity to PDE delta (unknown origin) assessed as dissociation constant measured after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Second Military Medical University
Curated by ChEMBL
Second Military Medical University
Curated by ChEMBL
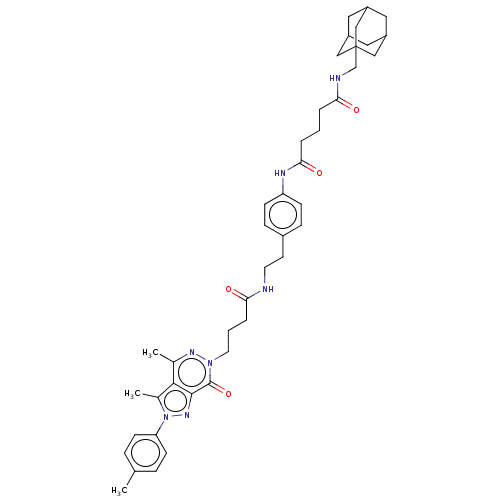
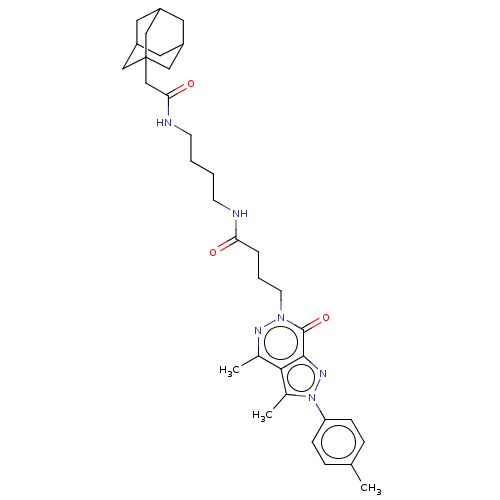
Affinity DataKi: 8.60nMAssay Description:Binding affinity to PDE delta (unknown origin) assessed as dissociation constant measured after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Second Military Medical University
Curated by ChEMBL
Second Military Medical University
Curated by ChEMBL
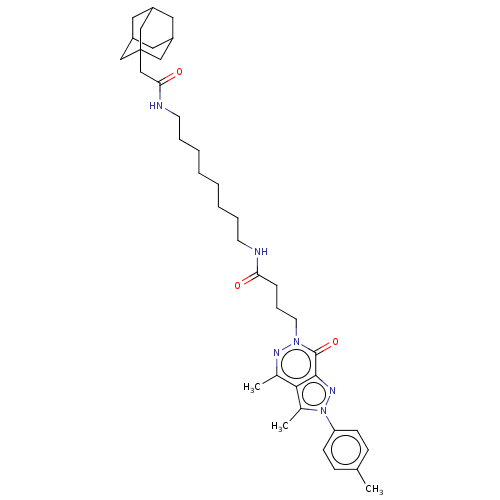
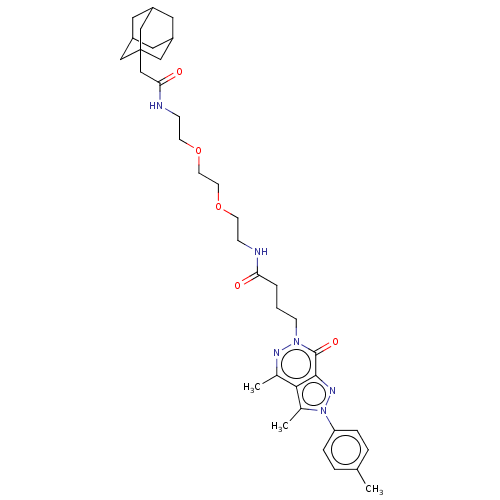
Affinity DataKi: 9.30nMAssay Description:Binding affinity to PDE delta (unknown origin) assessed as dissociation constant measured after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Second Military Medical University
Curated by ChEMBL
Second Military Medical University
Curated by ChEMBL
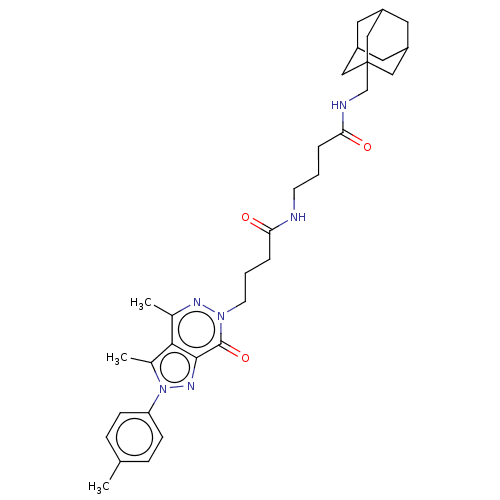
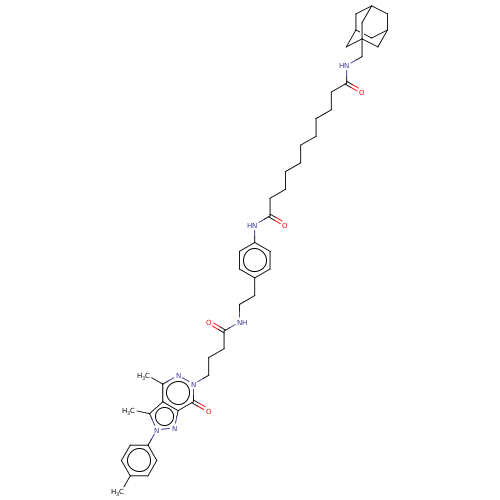
Affinity DataKi: 14nMAssay Description:Binding affinity to PDE delta (unknown origin) assessed as dissociation constant measured after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Second Military Medical University
Curated by ChEMBL
Second Military Medical University
Curated by ChEMBL
Affinity DataKi: 26nMAssay Description:Binding affinity to PDE delta (unknown origin) assessed as dissociation constant measured after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Second Military Medical University
Curated by ChEMBL
Second Military Medical University
Curated by ChEMBL
Affinity DataKi: 29nMAssay Description:Binding affinity to PDE delta (unknown origin) assessed as dissociation constant measured after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Second Military Medical University
Curated by ChEMBL
Second Military Medical University
Curated by ChEMBL
Affinity DataKi: 39nMAssay Description:Binding affinity to PDE delta (unknown origin) assessed as dissociation constant measured after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Second Military Medical University
Curated by ChEMBL
Second Military Medical University
Curated by ChEMBL
Affinity DataKi: 41nMAssay Description:Binding affinity to PDE delta (unknown origin) assessed as dissociation constant measured after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Second Military Medical University
Curated by ChEMBL
Second Military Medical University
Curated by ChEMBL
Affinity DataKi: 44nMAssay Description:Binding affinity to PDE delta (unknown origin) assessed as dissociation constant measured after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Second Military Medical University
Curated by ChEMBL
Second Military Medical University
Curated by ChEMBL
Affinity DataKi: 45nMAssay Description:Binding affinity to PDE delta (unknown origin) assessed as dissociation constant measured after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Second Military Medical University
Curated by ChEMBL
Second Military Medical University
Curated by ChEMBL
Affinity DataKi: 49nMAssay Description:Binding affinity to PDE delta (unknown origin) assessed as dissociation constant measured after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Second Military Medical University
Curated by ChEMBL
Second Military Medical University
Curated by ChEMBL
Affinity DataKi: 132nMAssay Description:Binding affinity to PDE delta (unknown origin) assessed as dissociation constant measured after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Second Military Medical University
Curated by ChEMBL
Second Military Medical University
Curated by ChEMBL
Affinity DataKi: 147nMAssay Description:Binding affinity to PDE delta (unknown origin) assessed as dissociation constant measured after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Second Military Medical University
Curated by ChEMBL
Second Military Medical University
Curated by ChEMBL
Affinity DataKi: >1.00E+3nMAssay Description:Binding affinity to PDE delta (unknown origin) assessed as dissociation constant measured after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair