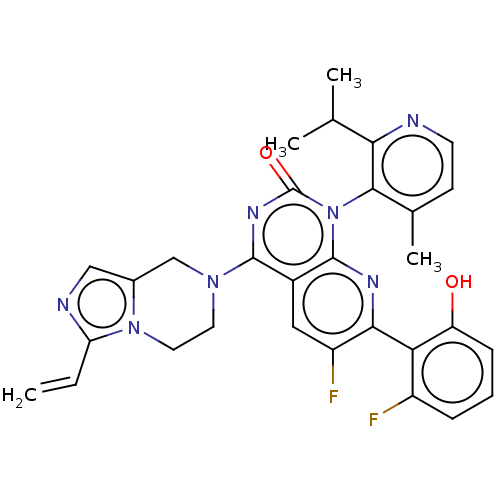
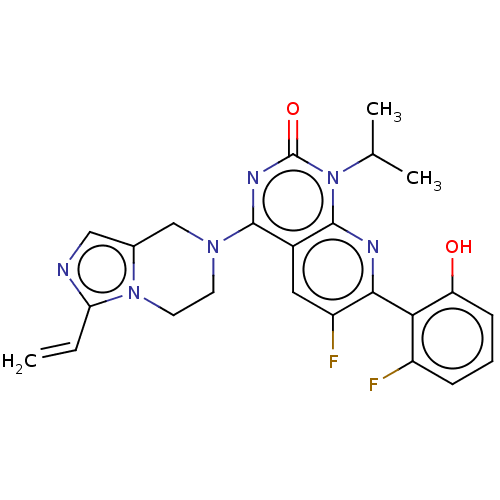
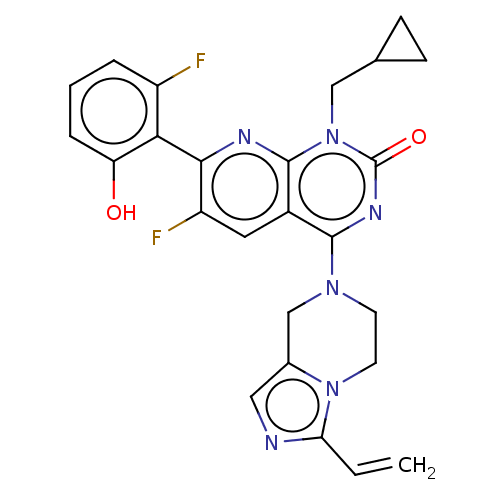
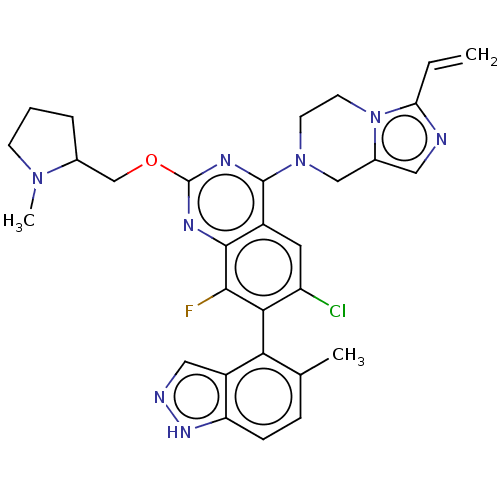
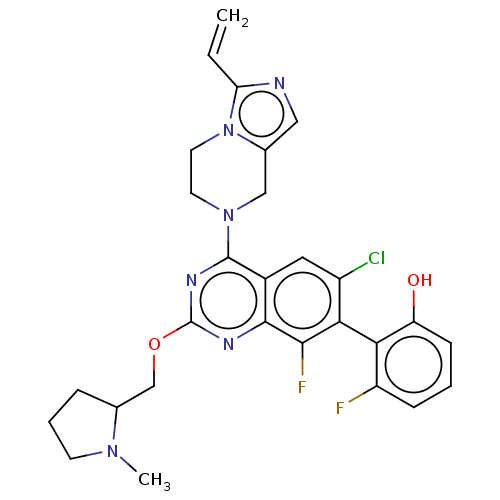
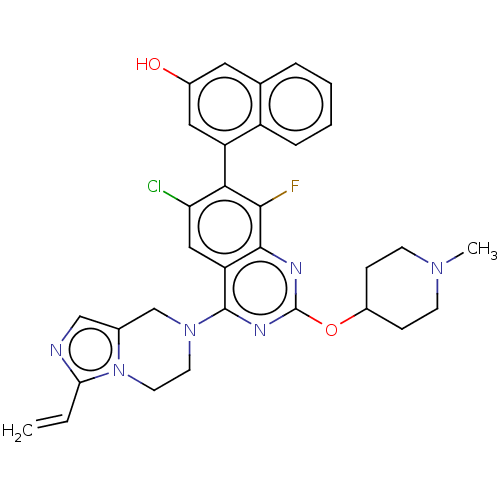
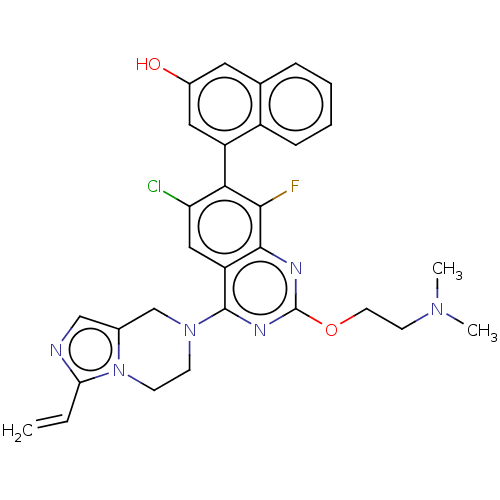
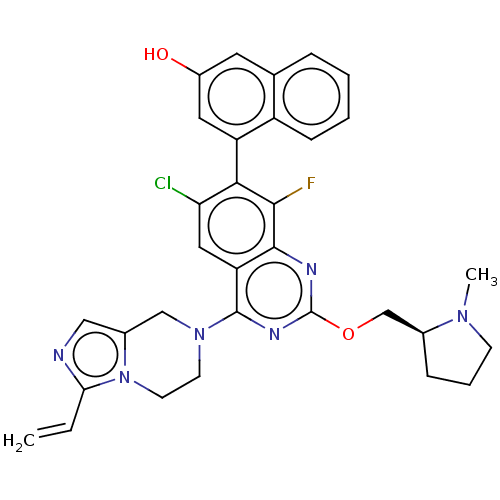
Report error Found 23 Enz. Inhib. hit(s) with all data for entry = 50016757
Affinity DataIC50: 100nMAssay Description:Inhibition of BODIPY loaded KRAS G12C mutant (unknown origin) assessed as inhibition of GDP-GTP exchange rate in presence of SOS1/GppNHp incubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of BODIPY loaded KRAS G12C mutant (unknown origin) assessed as inhibition of GDP-GTP exchange rate in presence of SOS1/GppNHp incubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of BODIPY loaded KRAS G12C mutant (unknown origin) assessed as inhibition of GDP-GTP exchange rate in presence of SOS1/GppNHp incubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of BODIPY loaded KRAS G12C mutant (unknown origin) assessed as inhibition of GDP-GTP exchange rate in presence of SOS1/GppNHp incubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of BODIPY loaded KRAS G12C mutant (unknown origin) assessed as inhibition of GDP-GTP exchange rate in presence of SOS1/GppNHp incubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of BODIPY loaded KRAS G12C mutant (unknown origin) assessed as inhibition of GDP-GTP exchange rate in presence of SOS1/GppNHp incubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of BODIPY loaded KRAS G12C mutant (unknown origin) assessed as inhibition of GDP-GTP exchange rate in presence of SOS1/GppNHp incubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of BODIPY loaded KRAS G12C mutant (unknown origin) assessed as inhibition of GDP-GTP exchange rate in presence of SOS1/GppNHp incubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of BODIPY loaded KRAS G12C mutant (unknown origin) assessed as inhibition of GDP-GTP exchange rate in presence of SOS1/GppNHp incubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of BODIPY loaded KRAS G12C mutant (unknown origin) assessed as inhibition of GDP-GTP exchange rate in presence of SOS1/GppNHp incubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of BODIPY loaded KRAS G12C mutant (unknown origin) assessed as inhibition of GDP-GTP exchange rate in presence of SOS1/GppNHp incubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of KRAS G12C mutant in human MIA PaCa-2 cells assessed as reduction in ERK phosphorylation incubated for 2 to 4 hrs by HTRF assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of KRAS G12C mutant in human MIA PaCa-2 cells assessed as reduction in ERK phosphorylation incubated for 2 to 4 hrs by HTRF assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of KRAS G12C mutant in human MIA PaCa-2 cells assessed as reduction in ERK phosphorylation incubated for 2 to 4 hrs by HTRF assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of KRAS G12C mutant in human MIA PaCa-2 cells assessed as reduction in ERK phosphorylation incubated for 2 to 4 hrs by HTRF assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of KRAS G12C mutant in human MIA PaCa-2 cells assessed as reduction in ERK phosphorylation incubated for 2 to 4 hrs by HTRF assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of KRAS G12C mutant in human MIA PaCa-2 cells assessed as reduction in ERK phosphorylation incubated for 2 to 4 hrs by HTRF assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of KRAS G12C mutant in human MIA PaCa-2 cells assessed as reduction in ERK phosphorylation incubated for 2 to 4 hrs by HTRF assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of KRAS G12C mutant in human MIA PaCa-2 cells assessed as reduction in ERK phosphorylation incubated for 2 to 4 hrs by HTRF assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of KRAS G12C mutant in human MIA PaCa-2 cells assessed as reduction in ERK phosphorylation incubated for 2 to 4 hrs by HTRF assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of KRAS G12C mutant in human MIA PaCa-2 cells assessed as reduction in ERK phosphorylation incubated for 2 to 4 hrs by HTRF assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of KRAS G12C mutant in human MIA PaCa-2 cells assessed as reduction in ERK phosphorylation incubated for 2 to 4 hrs by HTRF assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of KRAS G12C mutant in human MIA PaCa-2 cells assessed as reduction in ERK phosphorylation incubated for 2 to 4 hrs by HTRF assayMore data for this Ligand-Target Pair