Report error Found 26 Enz. Inhib. hit(s) with all data for entry = 50018789
Affinity DataIC50: 4.10nMAssay Description:Inhibition of human recombinant MAO-A assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 724nMAssay Description:Inhibition of human recombinant MAO-A assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 1.21E+3nMAssay Description:Inhibition of human recombinant MAO-A assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 1.68E+3nMAssay Description:Inhibition of human recombinant MAO-A assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 2.03E+3nMAssay Description:Inhibition of human recombinant MAO-A assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 2.79E+3nMAssay Description:Inhibition of human recombinant MAO-B assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 3.89E+3nMAssay Description:Inhibition of human recombinant MAO-A assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 3.90E+3nMAssay Description:Inhibition of human recombinant MAO-B assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 4.72E+3nMAssay Description:Inhibition of human recombinant MAO-A assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 5.17E+3nMAssay Description:Inhibition of human recombinant MAO-A assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 6.66E+3nMAssay Description:Inhibition of human recombinant MAO-A assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 7.15E+3nMAssay Description:Inhibition of human recombinant MAO-A assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 7.82E+3nMAssay Description:Inhibition of human recombinant MAO-A assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 8.43E+3nMAssay Description:Inhibition of human recombinant MAO-A assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 9.79E+3nMAssay Description:Inhibition of human recombinant MAO-A assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 1.05E+4nMAssay Description:Inhibition of human recombinant MAO-A assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 1.34E+4nMAssay Description:Inhibition of human recombinant MAO-B assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+4nMAssay Description:Inhibition of human recombinant MAO-A assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 1.67E+4nMAssay Description:Inhibition of human recombinant MAO-B assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 2.13E+4nMAssay Description:Inhibition of human recombinant MAO-B assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 2.54E+4nMAssay Description:Inhibition of human recombinant MAO-A assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 3.65E+4nMAssay Description:Inhibition of human recombinant MAO-B assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 3.68E+4nMAssay Description:Inhibition of human recombinant MAO-B assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 5.73E+4nMAssay Description:Inhibition of human recombinant MAO-B assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 6.48E+4nMAssay Description:Inhibition of human recombinant MAO-B assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
Affinity DataIC50: 8.42E+4nMAssay Description:Inhibition of human recombinant MAO-B assessed as inhibition of 4-hydroxyquinoline formation using kynuramine as substrate by fluorescence spectropho...More data for this Ligand-Target Pair
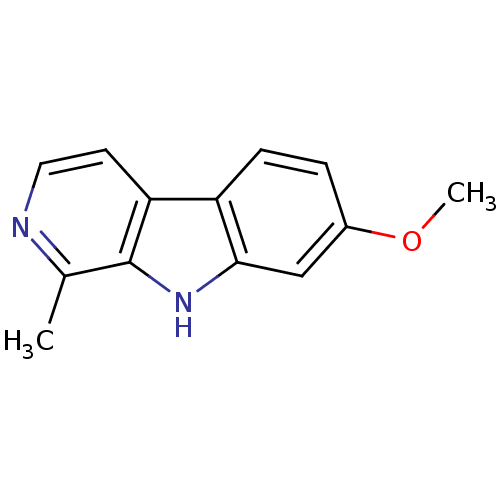
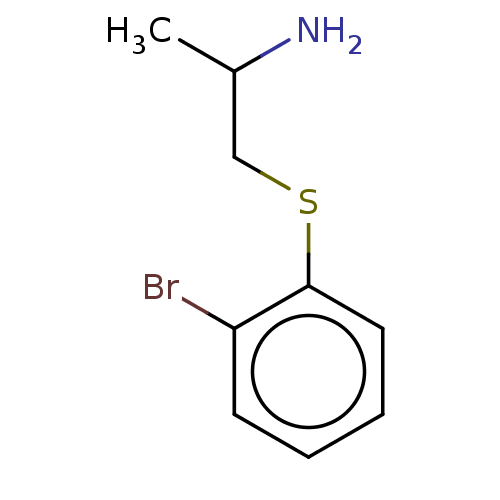
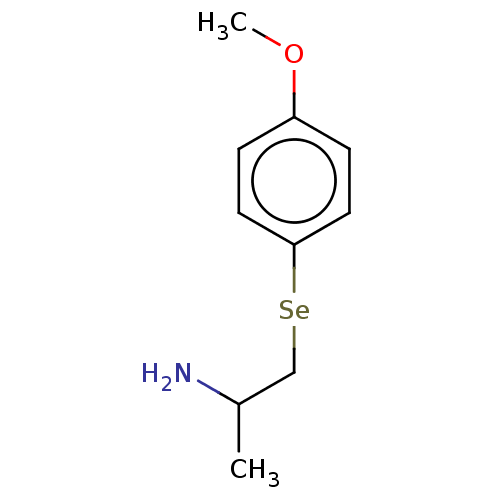
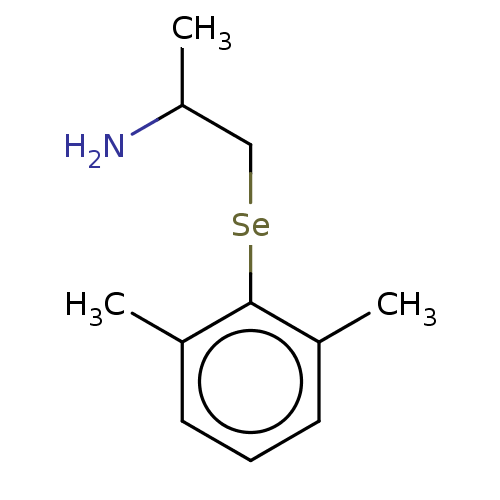
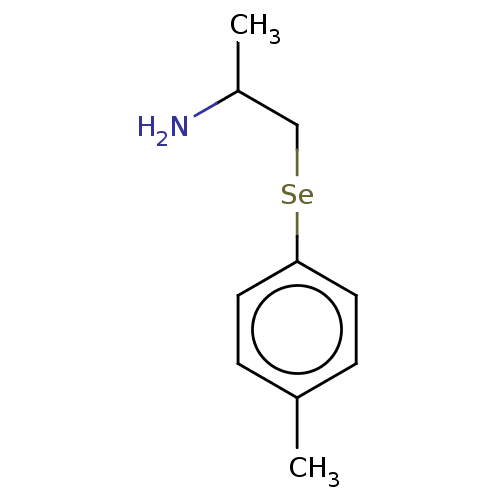

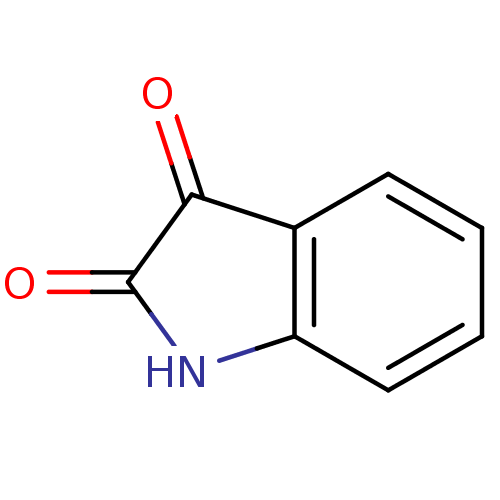
 3D Structure (crystal)
3D Structure (crystal)