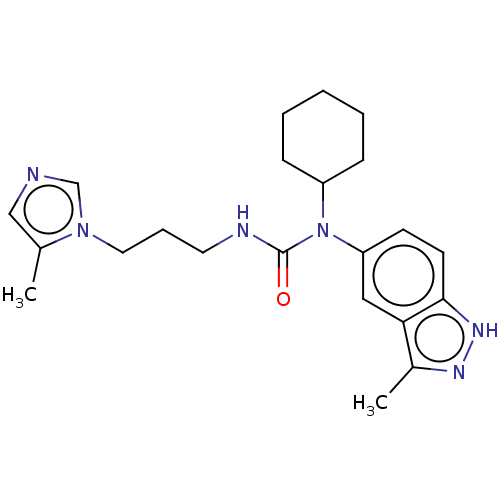
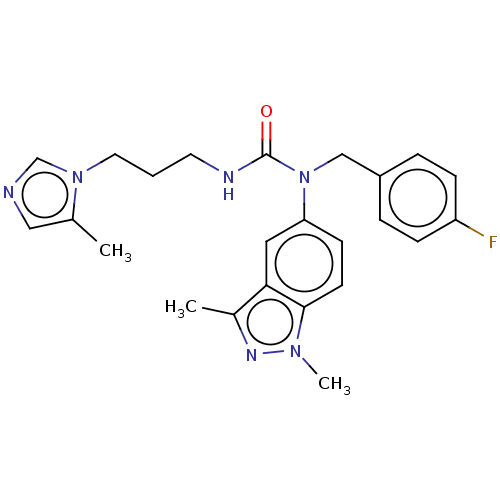
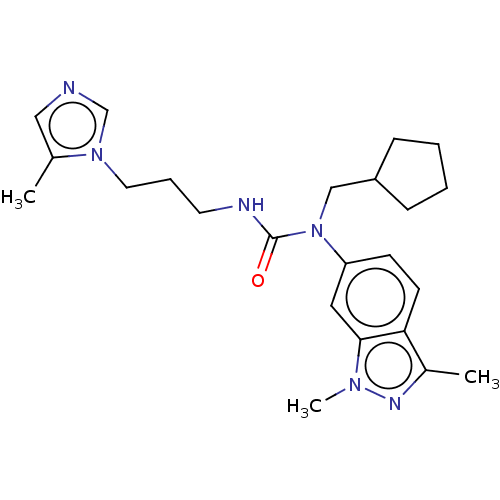
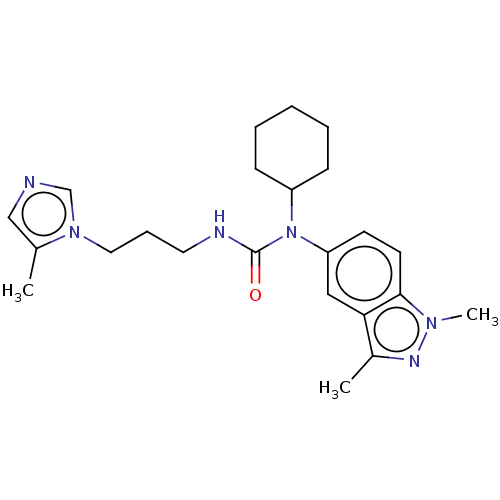
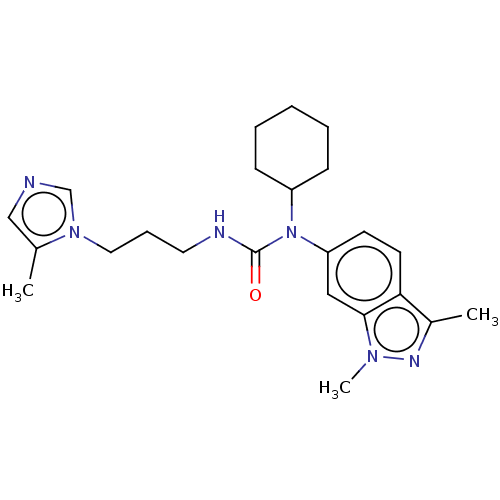
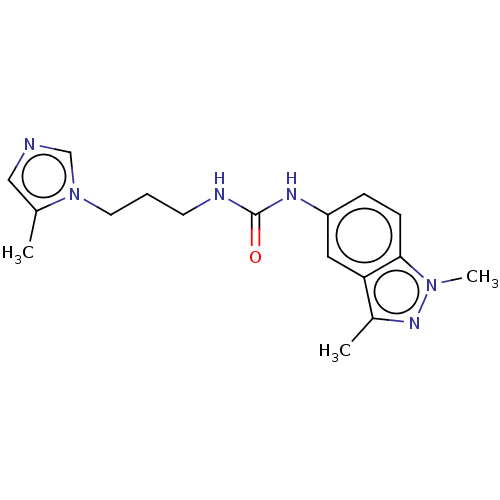
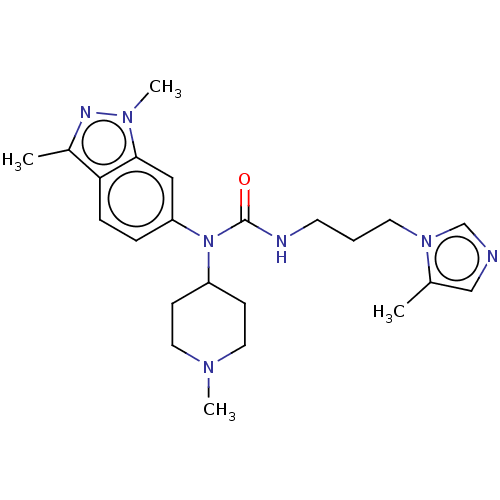
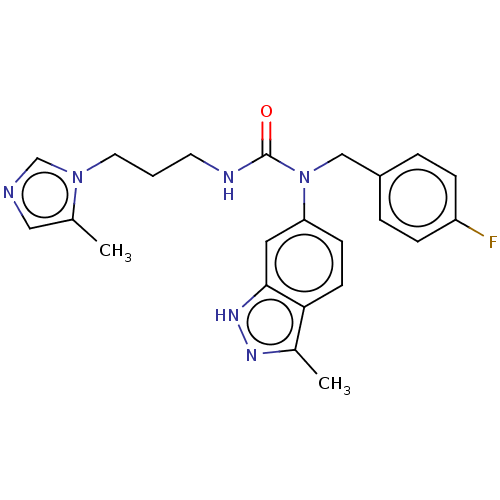
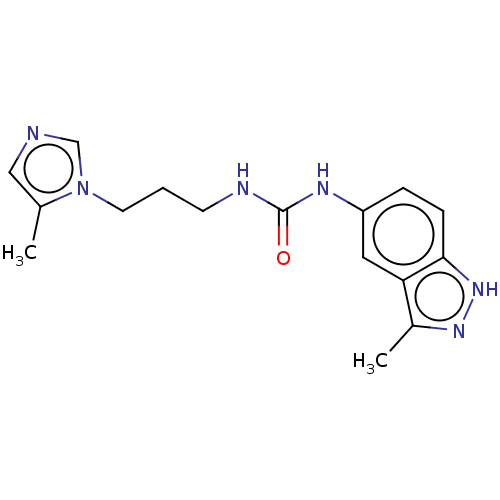
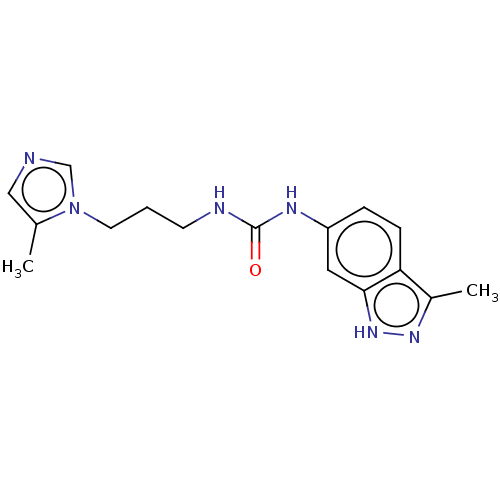
Report error Found 52 Enz. Inhib. hit(s) with all data for entry = 50018794
Affinity DataIC50: 2.30nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 2.30nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 3.20nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 3.40nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 4.40nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 5.30nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 5.60nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 9.80nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 14nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:Inhibition of mouse glutaminyl cyclaseMore data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 23nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 25nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 29nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 29nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 29nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 29nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of mouse glutaminyl cyclaseMore data for this Ligand-Target Pair
Affinity DataIC50: 32nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 34nMAssay Description:Inhibition of mouse glutaminyl cyclaseMore data for this Ligand-Target Pair
Affinity DataIC50: 44nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 54nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 62nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 66nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 66nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 71nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
TargetGlutaminyl-peptide cyclotransferase-like protein(Human)
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 80nMAssay Description:Inhibition of isoQC (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 87nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
TargetGlutaminyl-peptide cyclotransferase-like protein(Human)
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 88nMAssay Description:Inhibition of isoQC (unknown origin)More data for this Ligand-Target Pair
TargetGlutaminyl-peptide cyclotransferase-like protein(Human)
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 106nMAssay Description:Inhibition of isoQC (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 107nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 141nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
Affinity DataIC50: 210nMAssay Description:Inhibition of human glutaminyl cyclase using L-glutamine-7-amido-4-methylcoumarin as substrate incubated for 10 mins in presence of pyroglutamyl pept...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of hERG by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of hERG by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of hERG by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of hERG by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of hERG by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of hERG by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of hERG by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of hERG by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of hERG by fluorescence polarization assayMore data for this Ligand-Target Pair