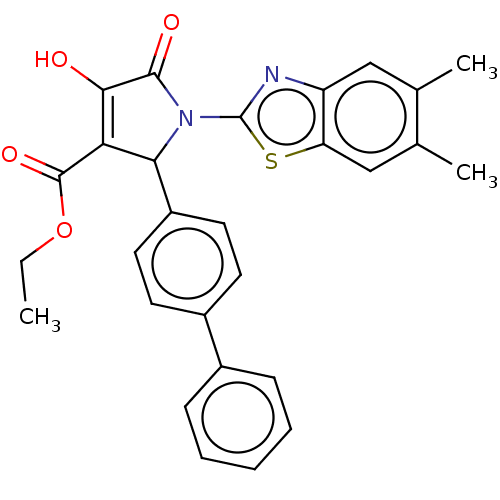
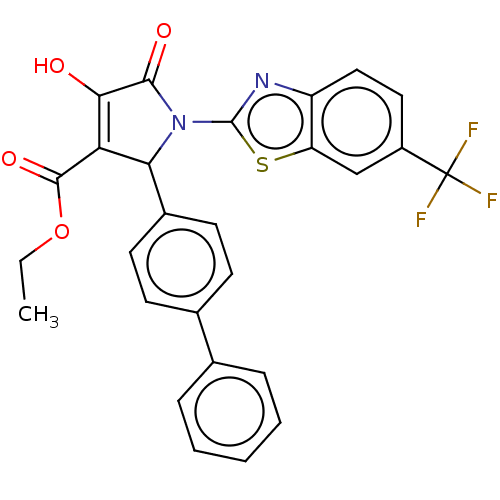
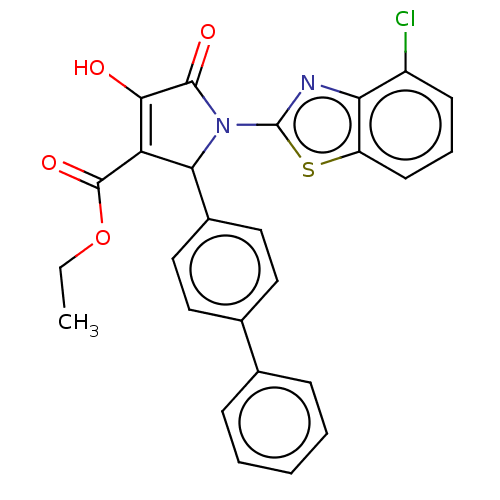
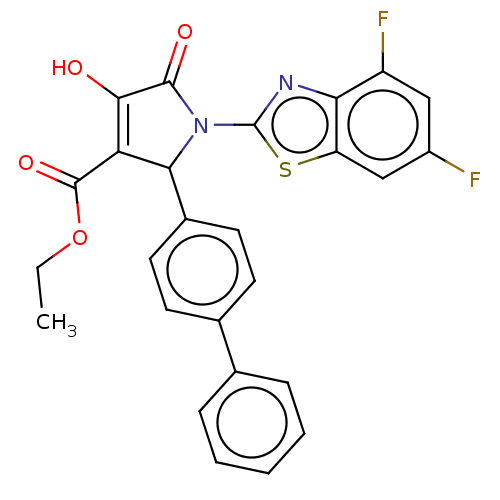
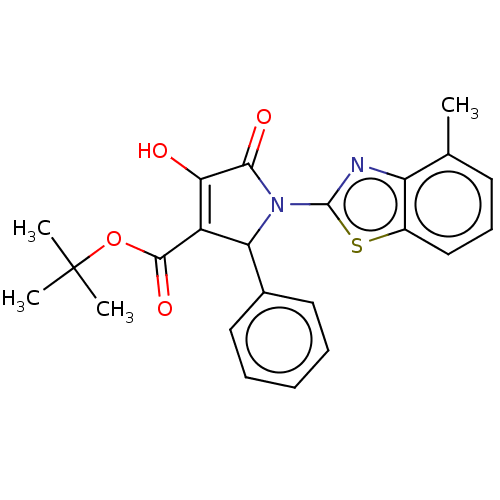
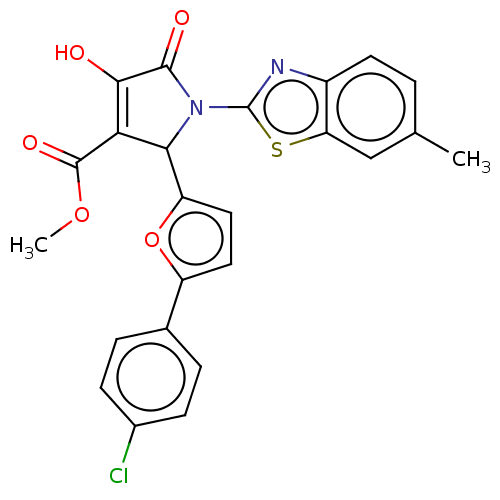
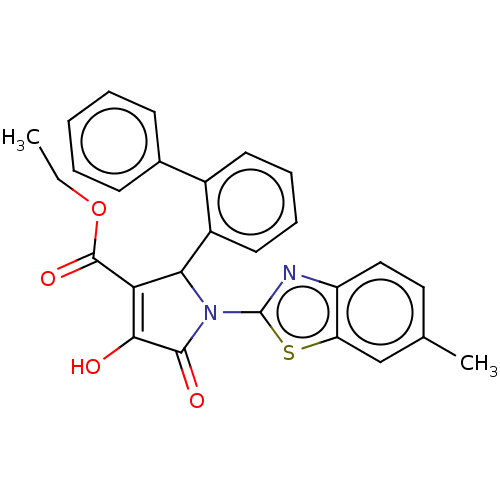
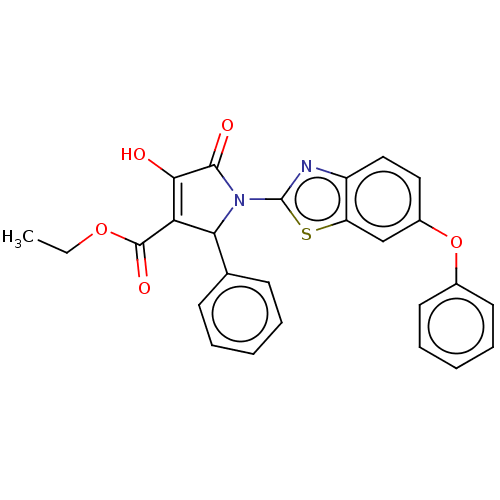
Report error Found 22 Enz. Inhib. hit(s) with all data for entry = 50018064
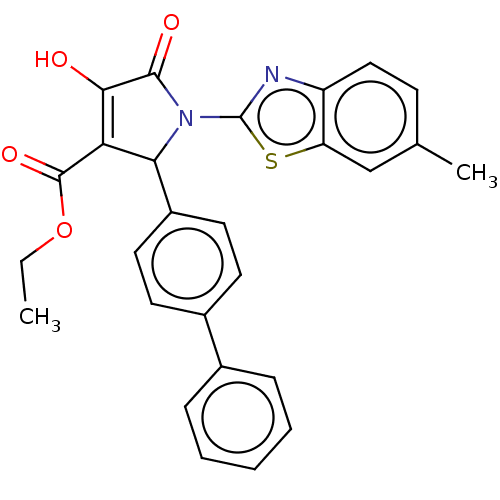
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
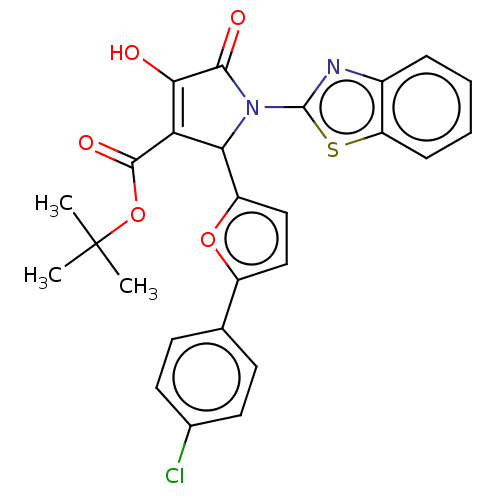
Affinity DataIC50: 4.50E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
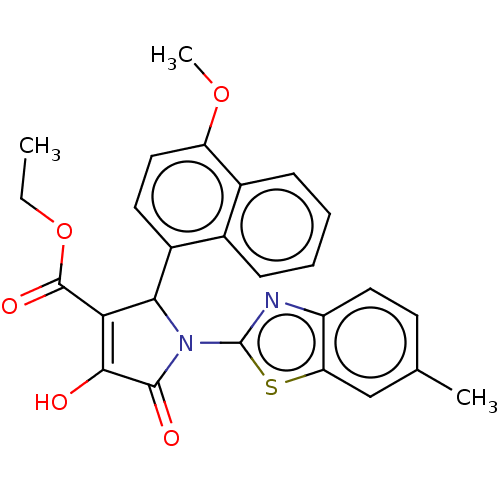
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
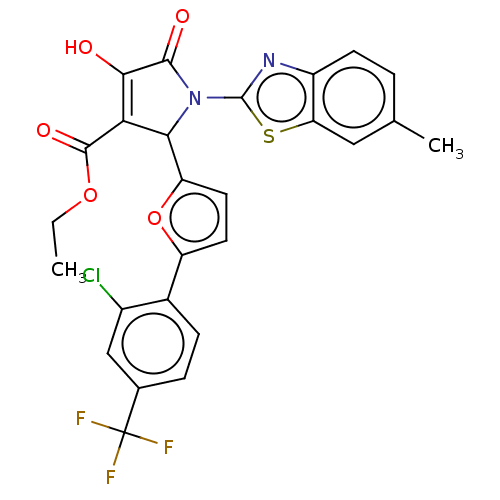
Affinity DataIC50: 4.80E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
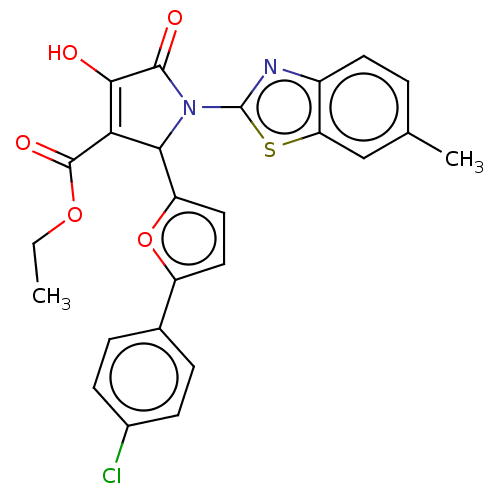
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
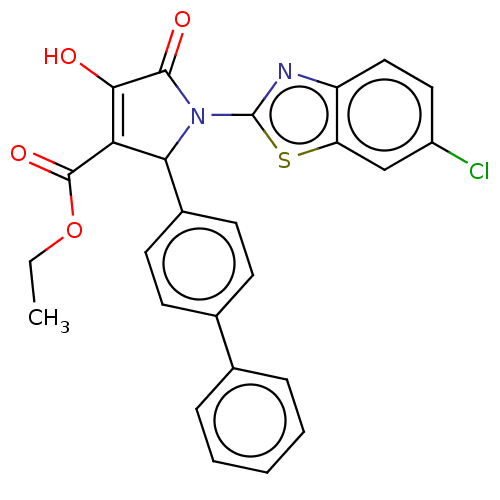
Affinity DataIC50: 4.90E+3nMAssay Description:Inhibition of his tagged Escherichia coli K12 MurA C115D mutant using UDP-N-acetylglucosamine as substrate in presence of DTT and PEP by microplate r...More data for this Ligand-Target Pair
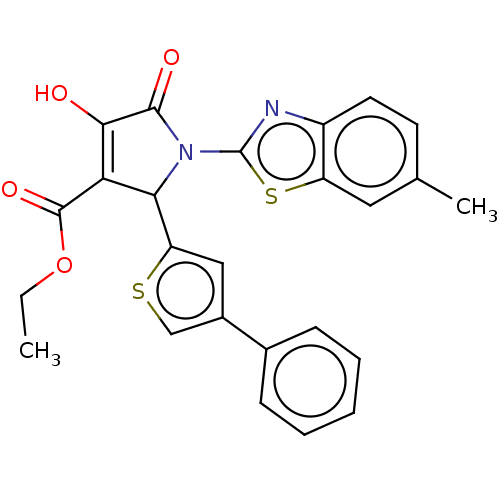
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
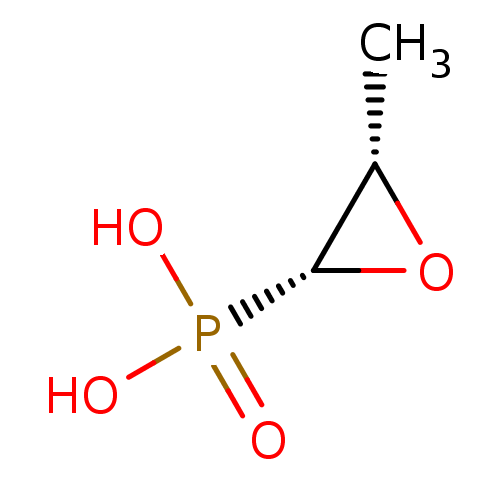
Affinity DataIC50: 5.10E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
Affinity DataIC50: 5.30E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
Affinity DataIC50: 5.80E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
Affinity DataIC50: 6.10E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
Affinity DataIC50: 7.00E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
Affinity DataIC50: 7.50E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
Affinity DataIC50: 8.50E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
Affinity DataIC50: 9.50E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
Affinity DataIC50: 9.90E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
Affinity DataIC50: 9.90E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
Affinity DataIC50: 1.10E+4nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
Affinity DataIC50: 1.50E+4nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
Affinity DataIC50: 1.60E+4nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
Affinity DataIC50: 1.70E+4nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
Affinity DataIC50: 1.70E+4nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
Affinity DataIC50: 2.10E+4nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
Affinity DataIC50: 2.20E+4nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
Affinity DataIC50: 2.20E+4nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Saarland University
Curated by ChEMBL
Saarland University
Curated by ChEMBL
Affinity DataIC50: 2.50E+4nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair