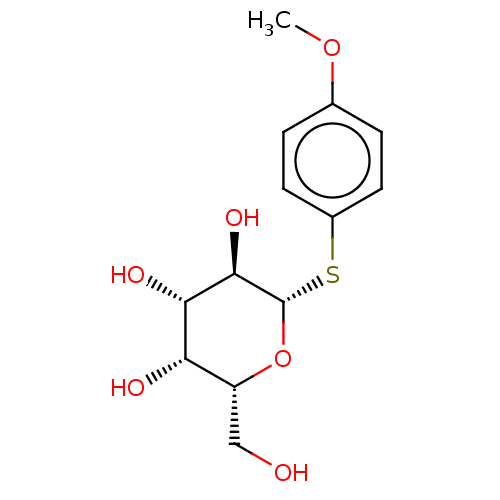
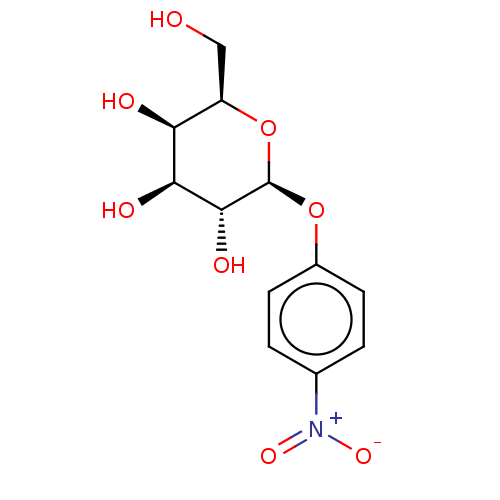
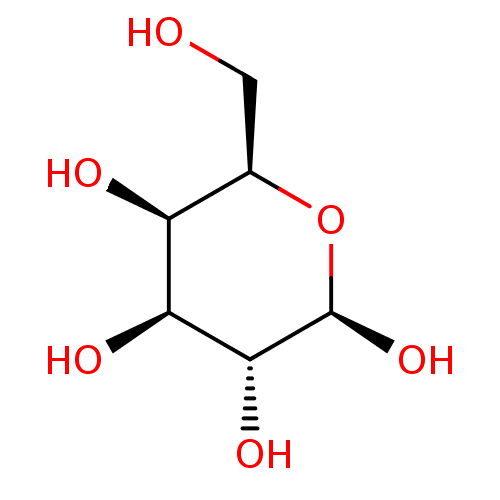
Report error Found 16 Enz. Inhib. hit(s) with all data for entry = 50019976
TargetPA-I galactophilic lectin(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University Paris-Saclay
Curated by ChEMBL
University Paris-Saclay
Curated by ChEMBL
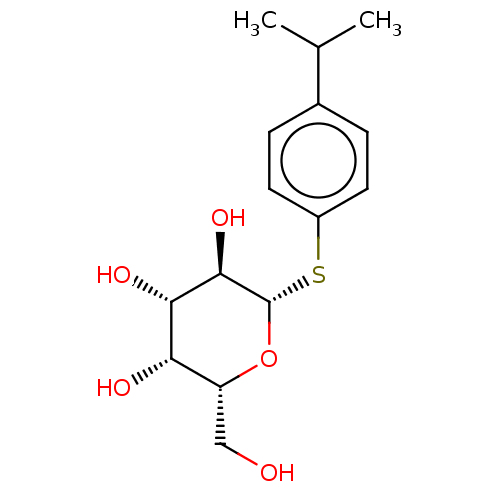
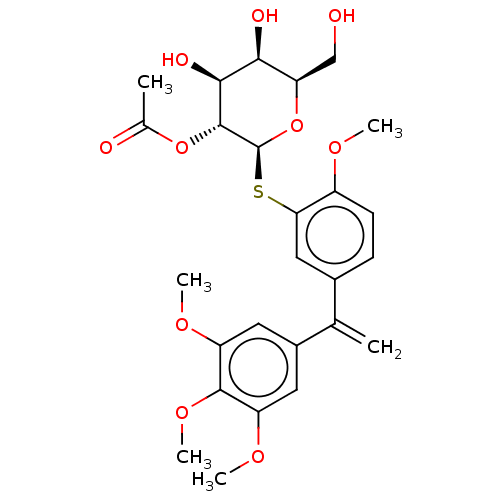
Affinity DataKd: 28nMAssay Description:Binding affinity to Pseudomonas aeruginosa LecA assessed as dissociation constant by isothermal titration calorimetryMore data for this Ligand-Target Pair
Ligand InfoSimilars
TargetPA-I galactophilic lectin(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University Paris-Saclay
Curated by ChEMBL
University Paris-Saclay
Curated by ChEMBL
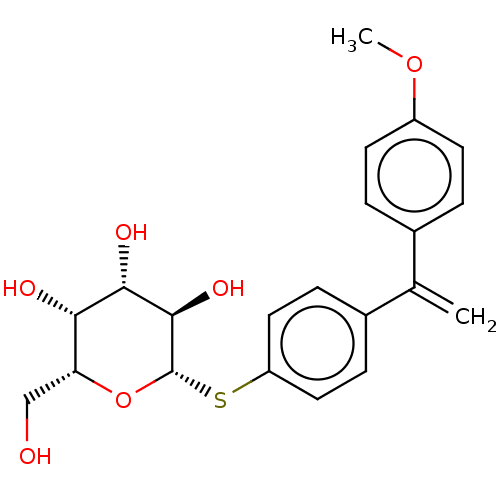
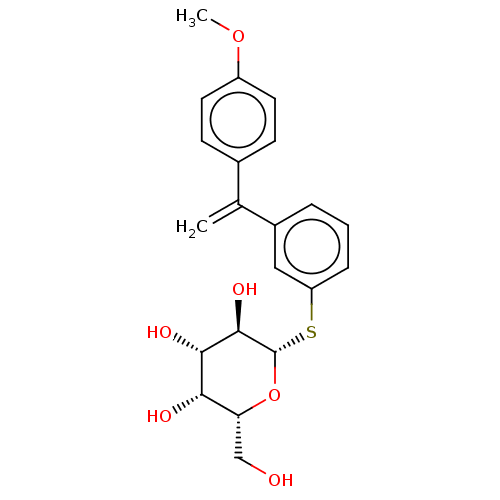
Affinity DataKd: 160nMAssay Description:Binding affinity to Pseudomonas aeruginosa LecA expressed in Escherichia coli BL21 (DE3) cells assessed as dissociation constant by isothermal titrat...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetPA-I galactophilic lectin(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University Paris-Saclay
Curated by ChEMBL
University Paris-Saclay
Curated by ChEMBL
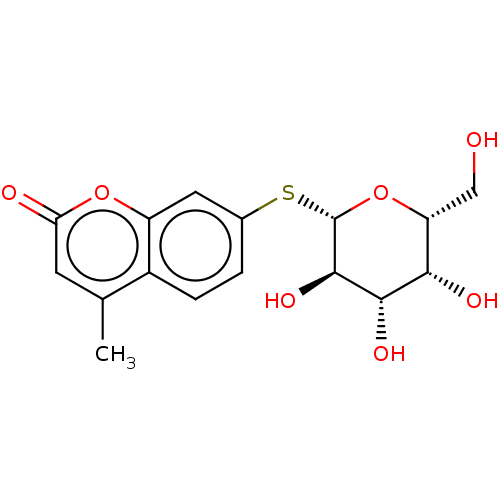
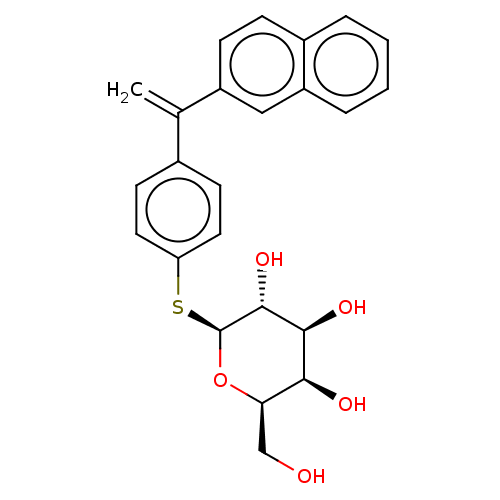
Affinity DataKd: 176nMAssay Description:Binding affinity to Pseudomonas aeruginosa LecAMore data for this Ligand-Target Pair
Ligand InfoSimilars
TargetPA-I galactophilic lectin(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University Paris-Saclay
Curated by ChEMBL
University Paris-Saclay
Curated by ChEMBL
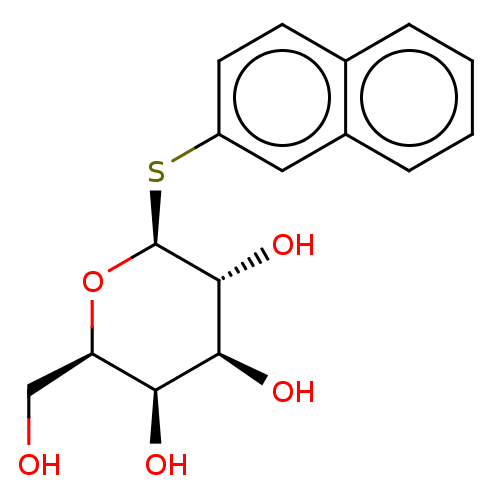
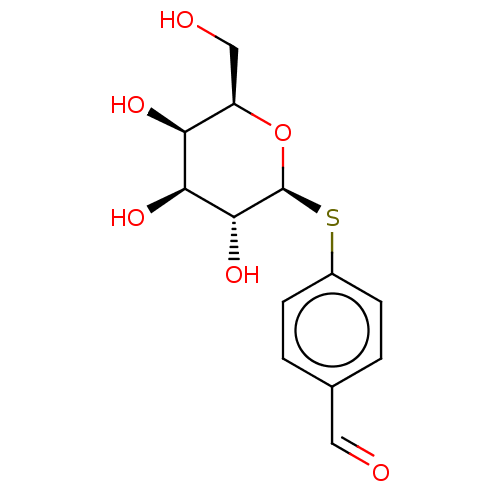
Affinity DataKd: 1.02E+3nMAssay Description:Binding affinity to Pseudomonas aeruginosa LecA expressed in Escherichia coli BL21 (DE3) cells assessed as dissociation constant by isothermal titrat...More data for this Ligand-Target Pair
TargetPA-I galactophilic lectin(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University Paris-Saclay
Curated by ChEMBL
University Paris-Saclay
Curated by ChEMBL
Affinity DataKd: 2.45E+3nMAssay Description:Binding affinity to Pseudomonas aeruginosa LecA expressed in Escherichia coli BL21 (DE3) cells assessed as dissociation constant by isothermal titrat...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetPA-I galactophilic lectin(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University Paris-Saclay
Curated by ChEMBL
University Paris-Saclay
Curated by ChEMBL
Affinity DataKd: 3.00E+3nMAssay Description:Binding affinity to Pseudomonas aeruginosa LecA expressed in Escherichia coli BL21 (DE3) cells assessed as dissociation constant by isothermal titrat...More data for this Ligand-Target Pair
TargetPA-I galactophilic lectin(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University Paris-Saclay
Curated by ChEMBL
University Paris-Saclay
Curated by ChEMBL
Affinity DataKd: 5.40E+3nMAssay Description:Binding affinity to Pseudomonas aeruginosa LecA assessed as dissociation constant by isothermal titration calorimetryMore data for this Ligand-Target Pair
Ligand InfoSimilars
TargetPA-I galactophilic lectin(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University Paris-Saclay
Curated by ChEMBL
University Paris-Saclay
Curated by ChEMBL
Affinity DataKd: 6.30E+3nMAssay Description:Binding affinity to Pseudomonas aeruginosa LecA assessed as dissociation constant by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetPA-I galactophilic lectin(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University Paris-Saclay
Curated by ChEMBL
University Paris-Saclay
Curated by ChEMBL
Affinity DataKd: 6.90E+3nMAssay Description:Binding affinity to Pseudomonas aeruginosa LecA expressed in Escherichia coli BL21 (DE3) cells assessed as dissociation constant by isothermal titrat...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetPA-I galactophilic lectin(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University Paris-Saclay
Curated by ChEMBL
University Paris-Saclay
Curated by ChEMBL
Affinity DataKd: 8.10E+3nMAssay Description:Binding affinity to Pseudomonas aeruginosa LecA expressed in Escherichia coli BL21 (DE3) cells assessed as dissociation constant by isothermal titrat...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetPA-I galactophilic lectin(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University Paris-Saclay
Curated by ChEMBL
University Paris-Saclay
Curated by ChEMBL
Affinity DataKd: 9.00E+3nMAssay Description:Binding affinity to Pseudomonas aeruginosa LecA expressed in Escherichia coli BL21 (DE3) cells assessed as dissociation constant by isothermal titrat...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetPA-I galactophilic lectin(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University Paris-Saclay
Curated by ChEMBL
University Paris-Saclay
Curated by ChEMBL
Affinity DataKd: 9.20E+3nMAssay Description:Binding affinity to Pseudomonas aeruginosa LecA expressed in Escherichia coli BL21 (DE3) cells assessed as dissociation constant by isothermal titrat...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetPA-I galactophilic lectin(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University Paris-Saclay
Curated by ChEMBL
University Paris-Saclay
Curated by ChEMBL
Affinity DataKd: 1.19E+4nMAssay Description:Binding affinity to Pseudomonas aeruginosa LecA expressed in Escherichia coli BL21 (DE3) cells assessed as dissociation constant by isothermal titrat...More data for this Ligand-Target Pair
Ligand InfoSimilars
TargetPA-I galactophilic lectin(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University Paris-Saclay
Curated by ChEMBL
University Paris-Saclay
Curated by ChEMBL
Affinity DataKd: 1.41E+4nMAssay Description:Binding affinity to Pseudomonas aeruginosa LecA assessed as dissociation constant by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetPA-I galactophilic lectin(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University Paris-Saclay
Curated by ChEMBL
University Paris-Saclay
Curated by ChEMBL
Affinity DataKd: 3.24E+4nMAssay Description:Binding affinity to Pseudomonas aeruginosa LecA assessed as dissociation constant by isothermal titration calorimetryMore data for this Ligand-Target Pair
TargetPA-I galactophilic lectin(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University Paris-Saclay
Curated by ChEMBL
University Paris-Saclay
Curated by ChEMBL
Affinity DataKd: 8.75E+4nMAssay Description:Binding affinity to Pseudomonas aeruginosa LecA assessed as dissociation constant by isothermal titration calorimetryMore data for this Ligand-Target Pair
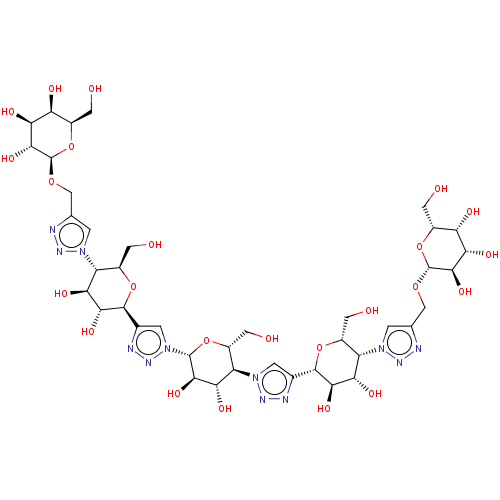
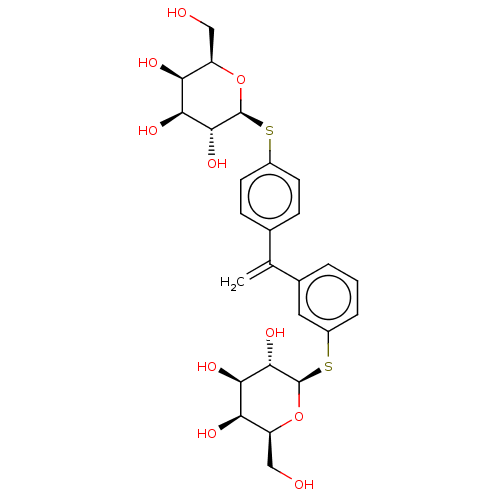
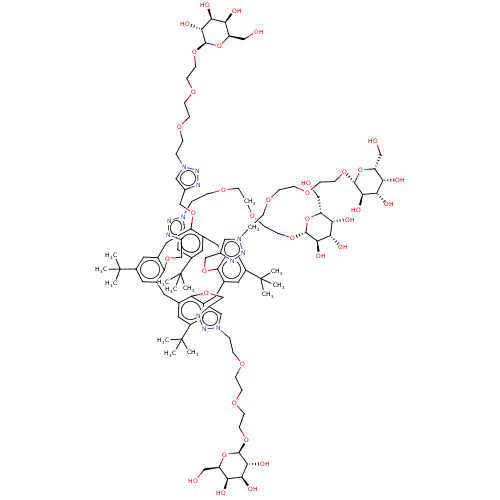
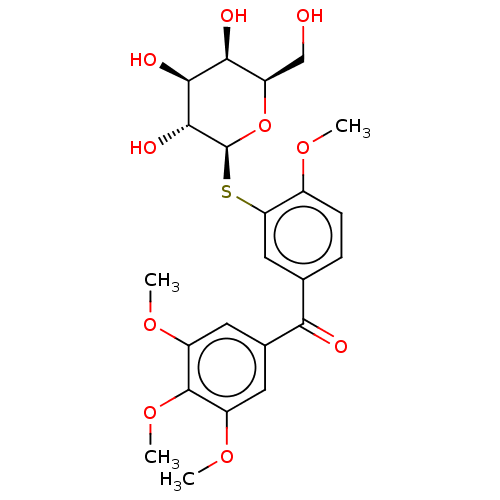
 3D Structure (crystal)
3D Structure (crystal)