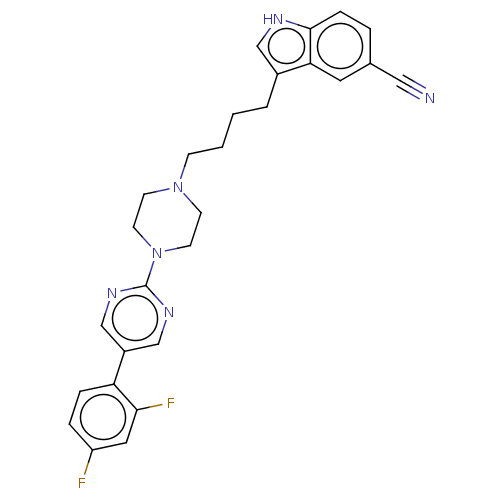
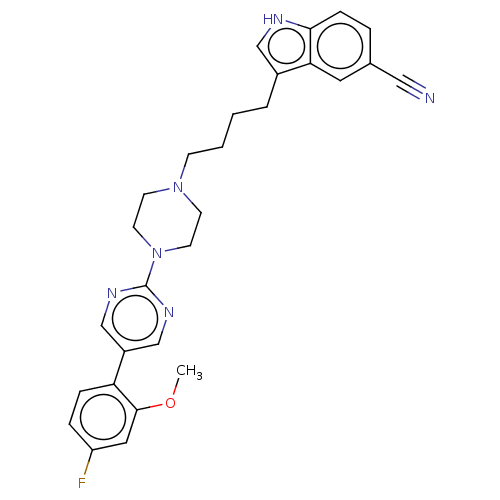
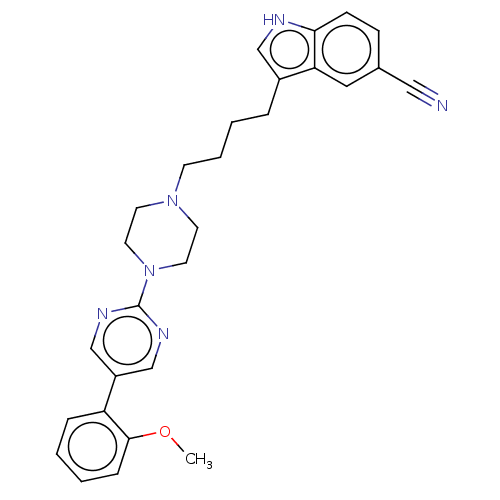
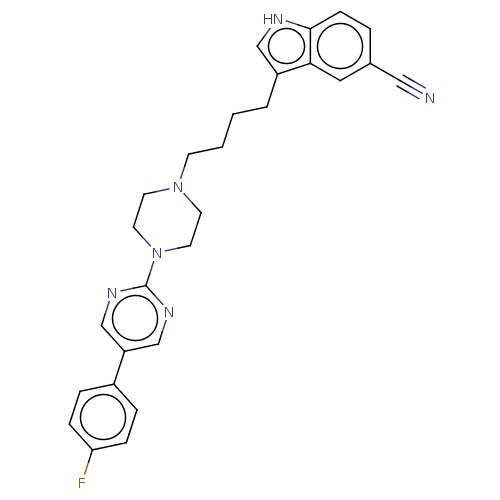
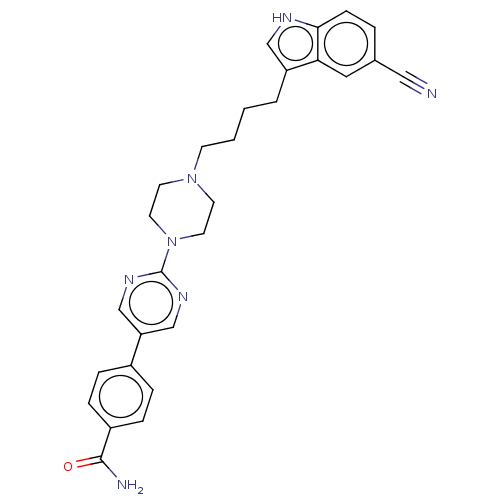
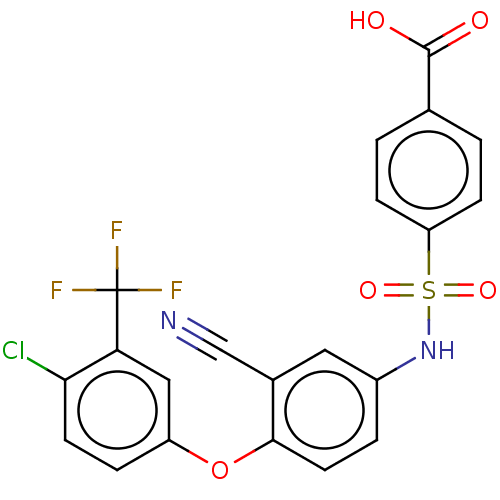
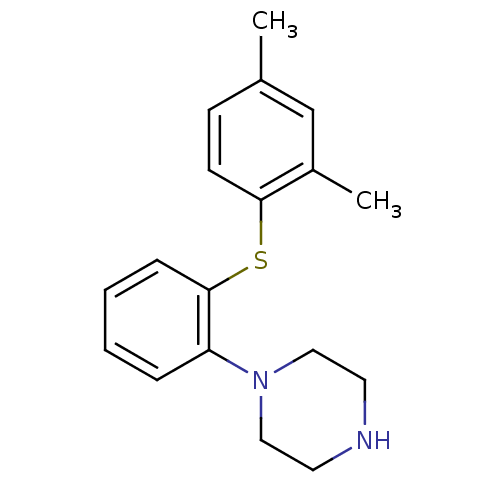
Report error Found 10 Enz. Inhib. hit(s) with all data for entry = 50019192
Affinity DataIC50: 0.320nMAssay Description:Inhibition of SERT receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 0.350nMAssay Description:Inhibition of SERT receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 0.480nMAssay Description:Inhibition of SERT receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 0.580nMAssay Description:Inhibition of SERT receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of SERT receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of SERT receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of SERT receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of SERT receptor (unknown origin)More data for this Ligand-Target Pair
TargetPlatelet-activating factor acetylhydrolase(Human)
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 5nMAssay Description:Inhibition of Lp-PLA2 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 5.40nMAssay Description:Inhibition of SERT receptor (unknown origin)More data for this Ligand-Target Pair
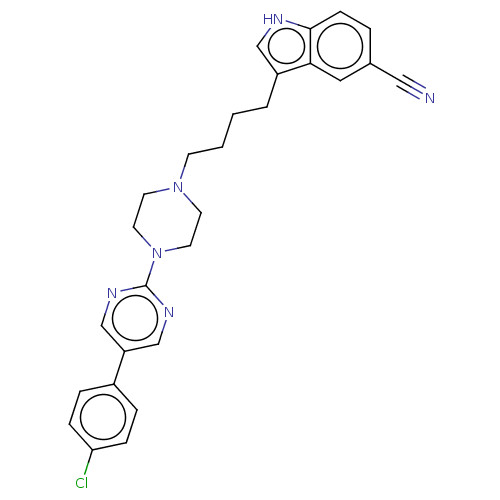
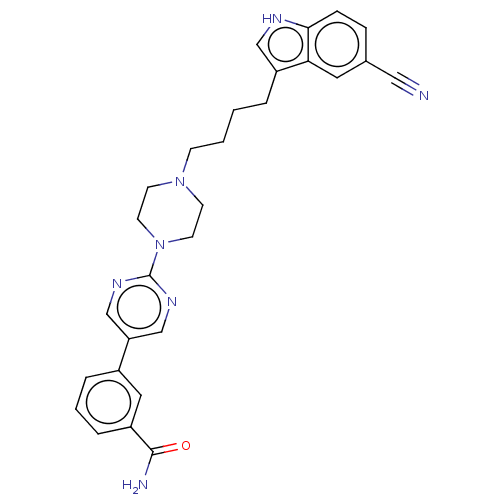
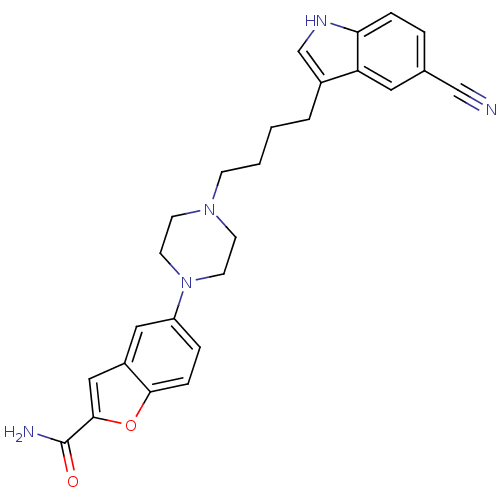
 3D Structure (crystal)
3D Structure (crystal)