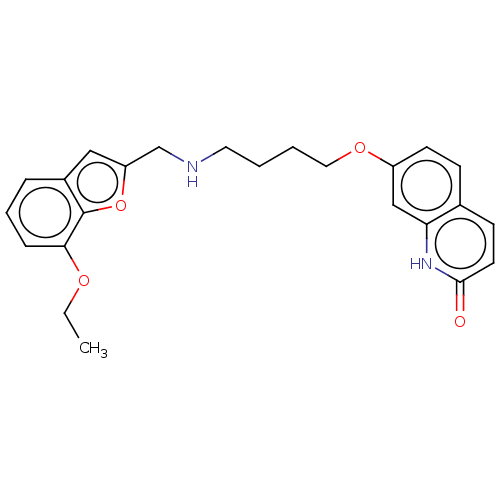
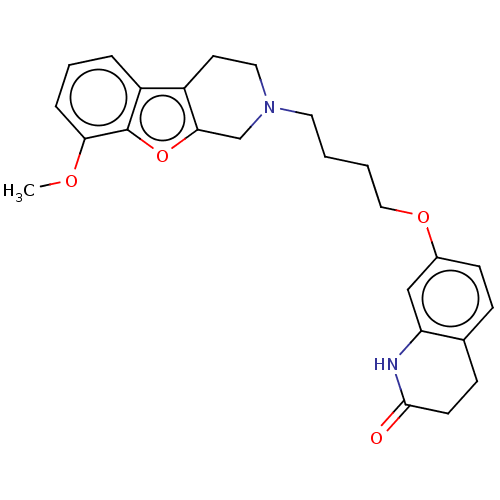
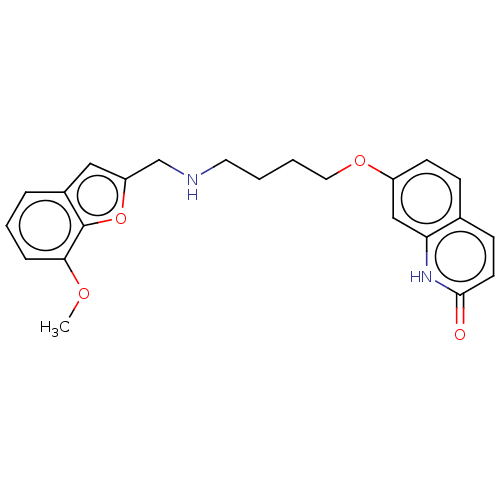
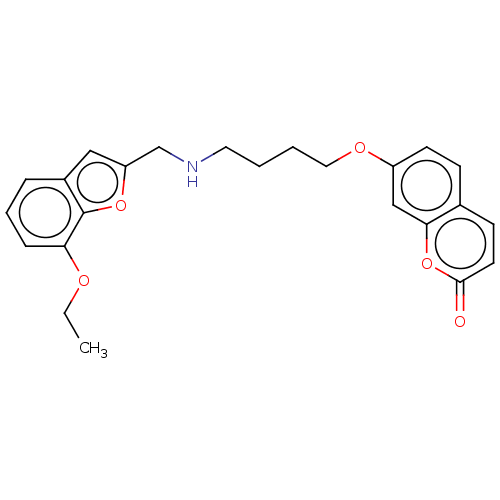
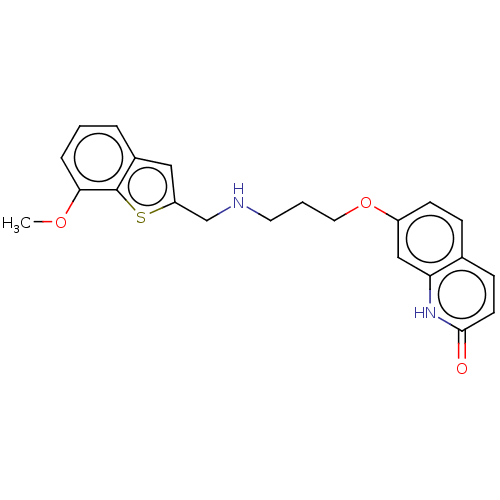
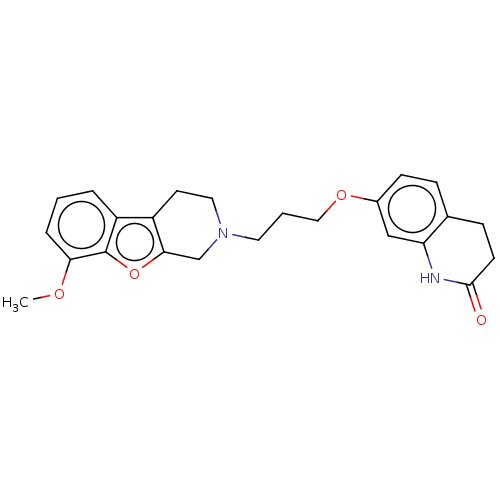
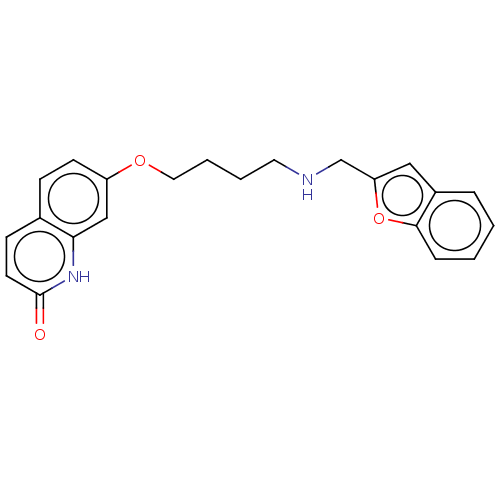
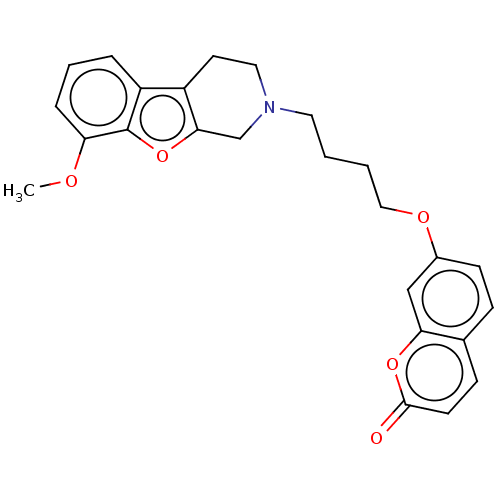
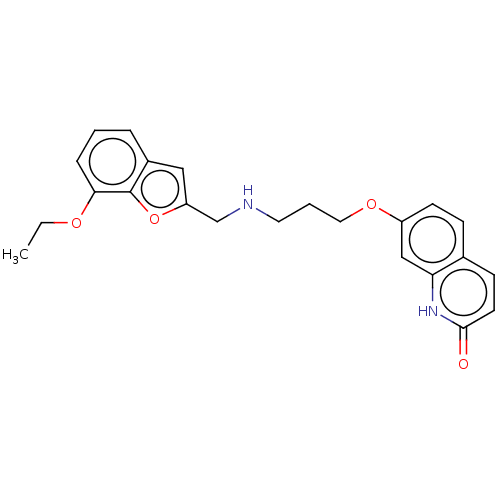
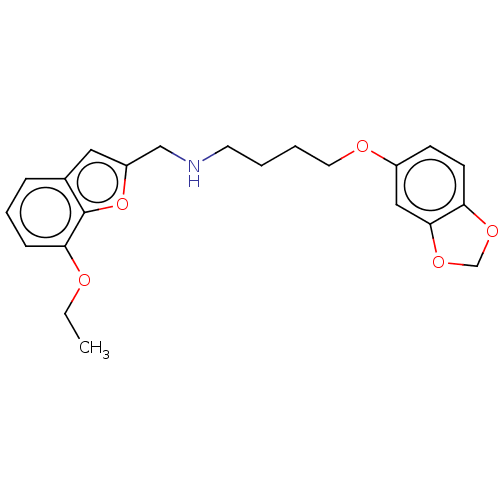
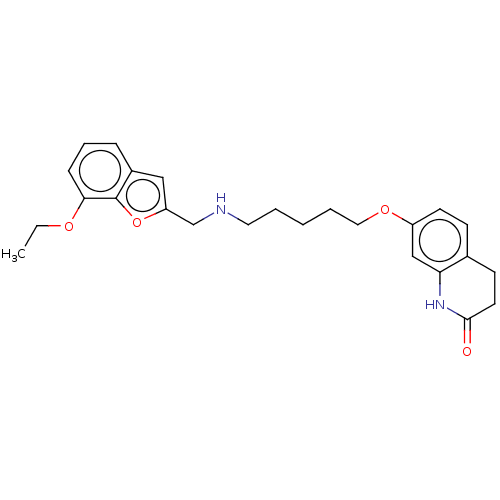
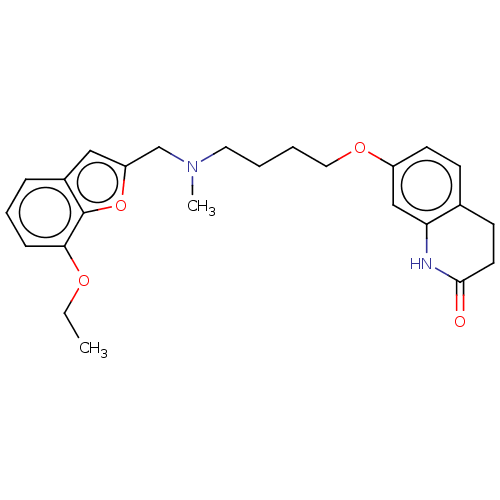
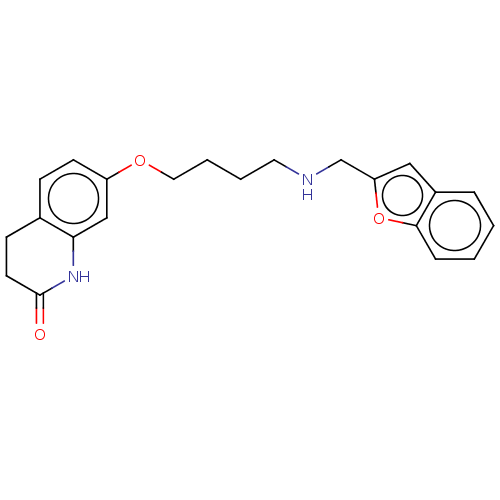
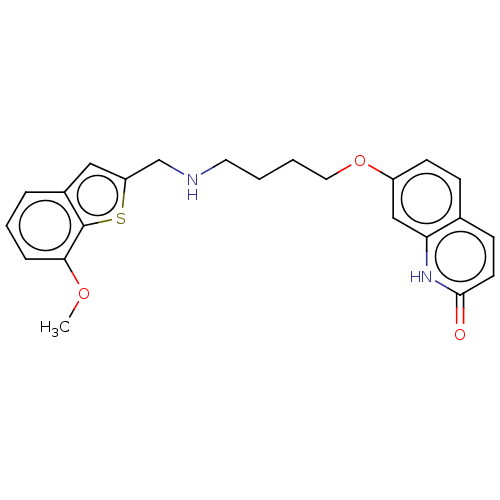
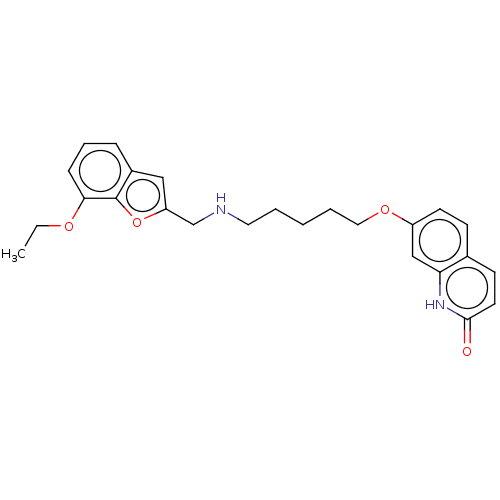
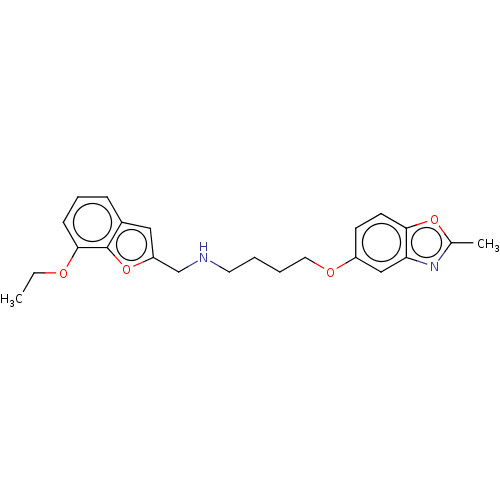
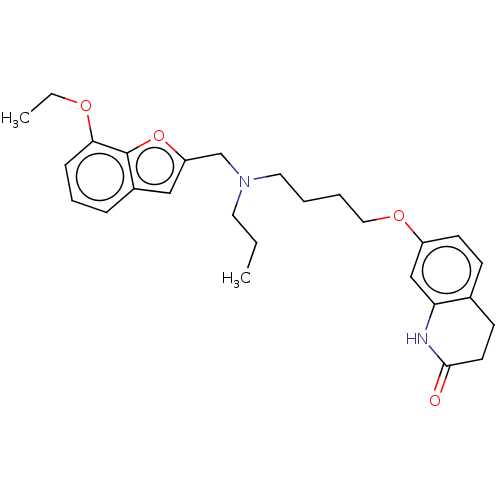
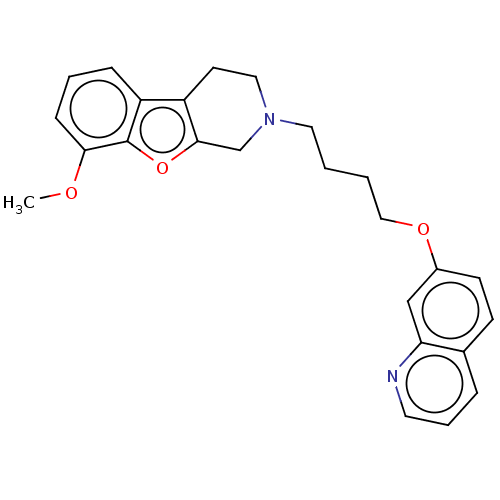
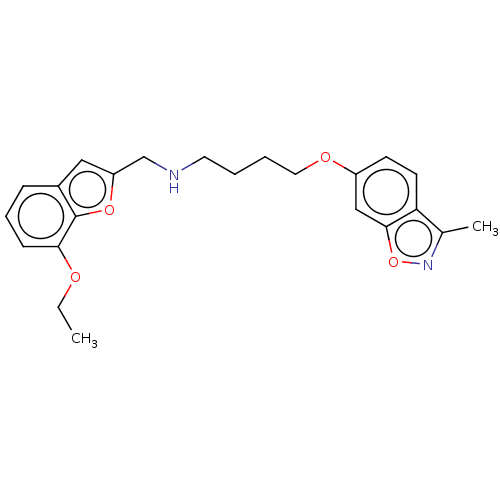
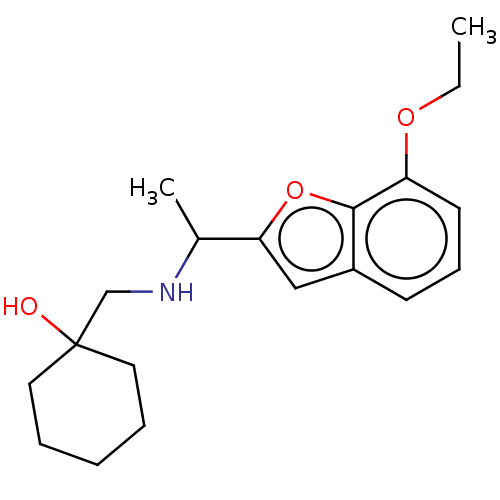
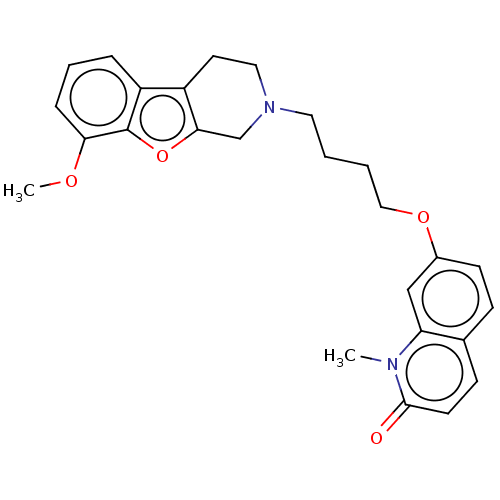
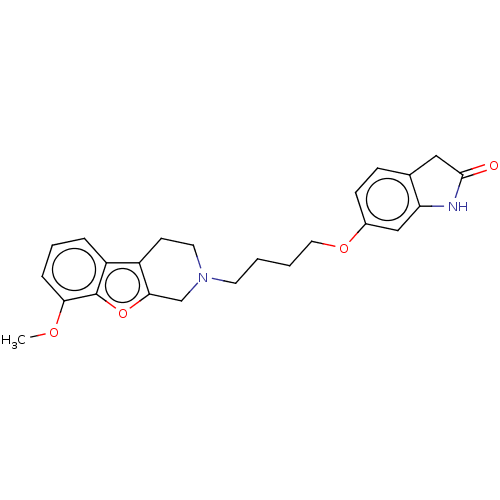
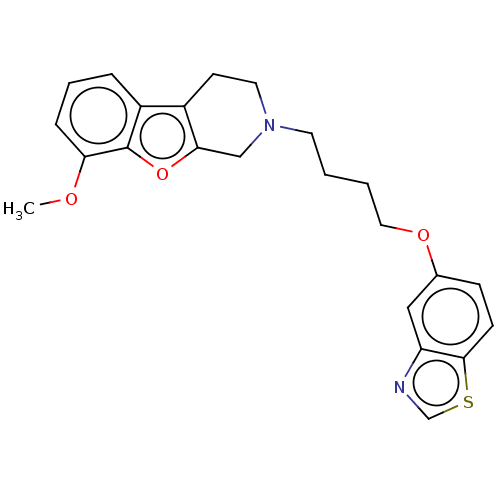
Report error Found 50 Enz. Inhib. hit(s) with all data for entry = 50020714
Affinity DataKi: 0.120nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 3.10nMAssay Description:Displacement of [3H]-LSD from 5-HT1A receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligand competitive bind...More data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 6.5nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Displacement of [3H]-LSD from 5-HT1A receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligand competitive bind...More data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D3 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 19nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D4 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 24nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D3 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 26nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D4 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 33nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 36nMAssay Description:Displacement of [3H]-SCH23390 from Dopamine D1 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligand compet...More data for this Ligand-Target Pair
Affinity DataKi: 39nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 63nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 68nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 84nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 89nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 99nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 109nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 139nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 149nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 159nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D5 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 181nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 189nMAssay Description:Displacement of [3H]-SCH23390 from Dopamine D1 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligand compet...More data for this Ligand-Target Pair
Affinity DataKi: 204nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 239nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 292nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 423nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 438nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 439nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 539nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 555nMAssay Description:Displacement of [3H]-LSD from 5-HT2C receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligand competitive bind...More data for this Ligand-Target Pair
Affinity DataKi: 582nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 611nMAssay Description:Displacement of [3H]-LSD from 5-HT2C receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligand competitive bind...More data for this Ligand-Target Pair
Affinity DataKi: 707nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 745nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 935nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 1.04E+3nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D5 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 1.11E+3nMAssay Description:Displacement of [3H]-LSD from 5-HT2A receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligand competitive bind...More data for this Ligand-Target Pair
Affinity DataKi: 1.27E+3nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 1.34E+3nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 1.79E+3nMAssay Description:Displacement of [3H]-LSD from 5-HT2A receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligand competitive bind...More data for this Ligand-Target Pair
Affinity DataKi: 2.08E+3nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 2.13E+3nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 2.18E+3nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 2.28E+3nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 3.19E+3nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair
Affinity DataKi: 2.59E+4nMAssay Description:Displacement of [3H]N-methylspiperone from Dopamine D2 receptor (unknown origin) expressed in HEK293T cell membrane incubated for 2 hrs by radioligan...More data for this Ligand-Target Pair