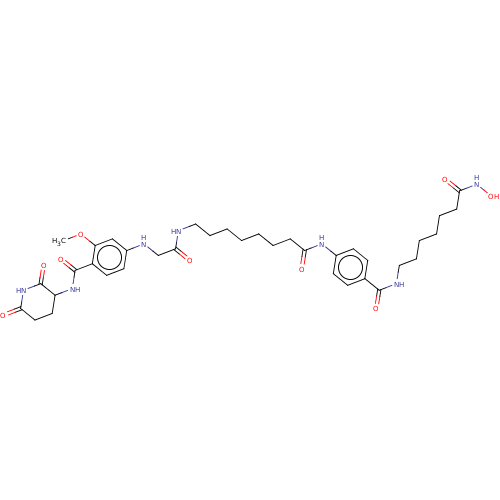
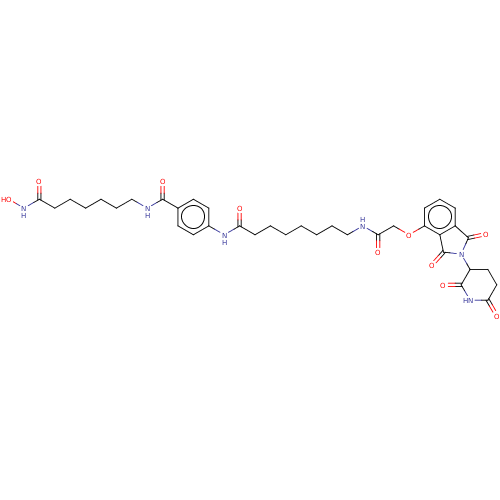
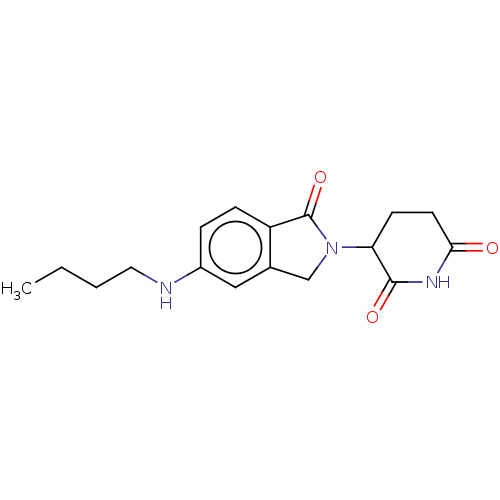
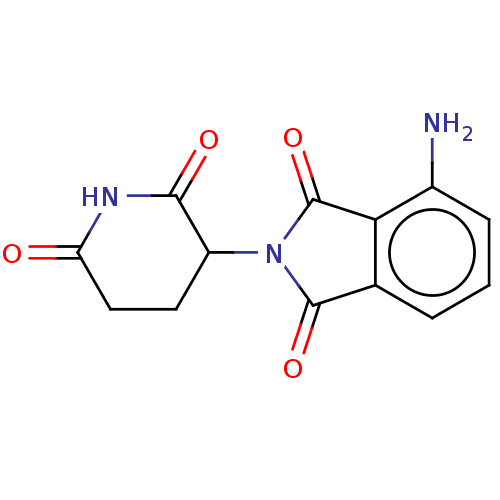
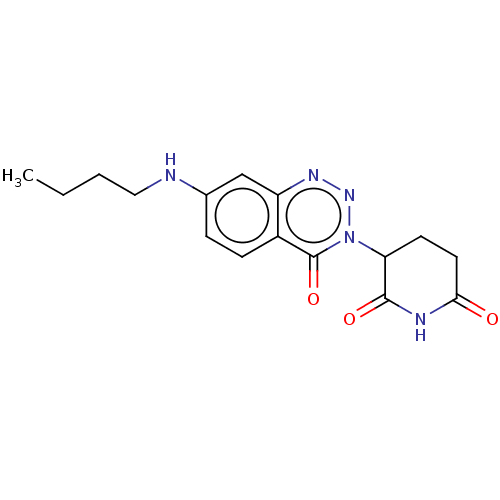
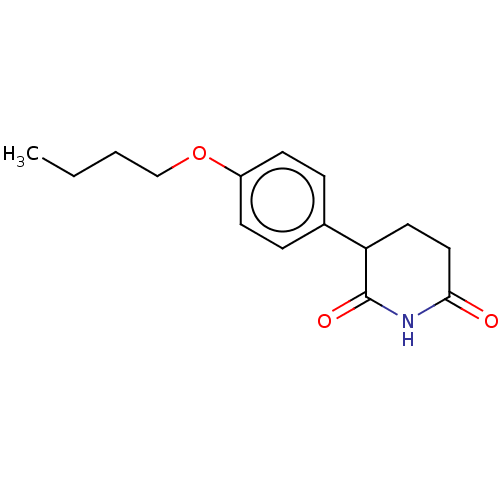
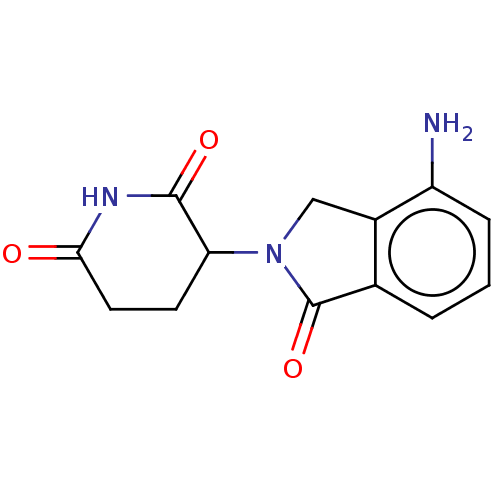
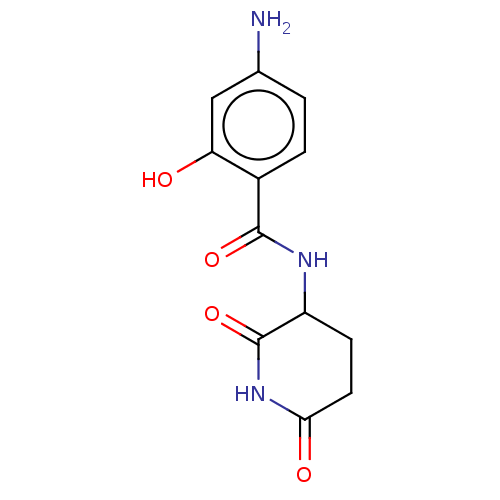
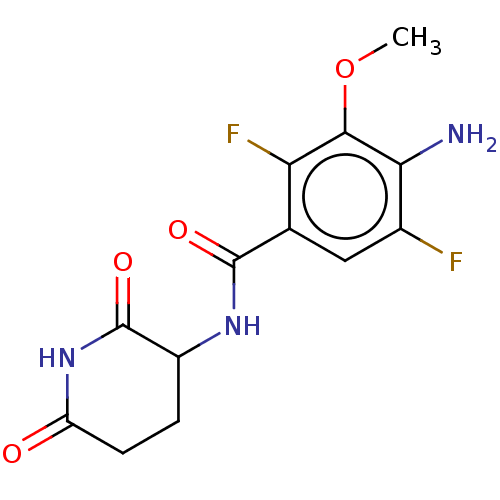
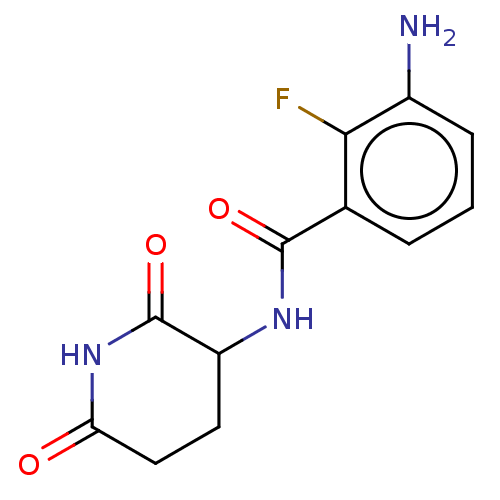
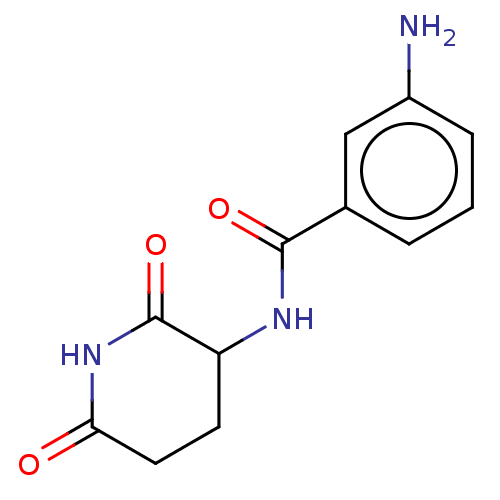
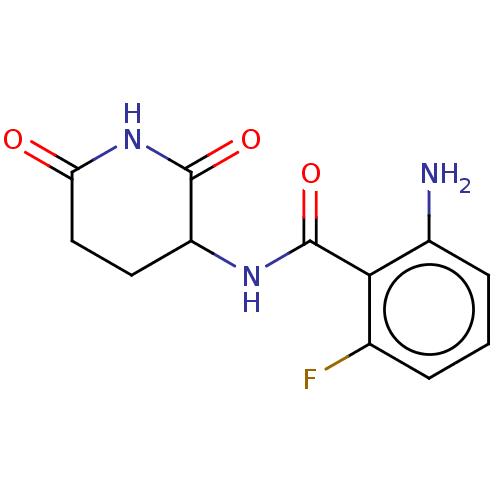
Report error Found 54 Enz. Inhib. hit(s) with all data for entry = 50020502
Affinity DataEC50: 5.80nMAssay Description:Protac activity at HDAC6 in human MM1.S cells assessed as HDAC6 protein degradation incubated for 24 hrs by immunoblotting analysisMore data for this Ligand-Target Pair
Affinity DataEC50: 6.5nMAssay Description:Protac activity at HDAC6 in human MM1.S cells assessed as HDAC6 protein degradation incubated for 24 hrs by immunoblotting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 430nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.30E+3nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.90E+3nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.90E+3nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 6.40E+3nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 6.80E+3nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 7.40E+3nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.10E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+4nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+4nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.50E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+4nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+4nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.50E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.80E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+4nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.90E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.90E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.00E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.10E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.50E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.50E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.60E+4nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 4.10E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 4.20E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 4.40E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 4.50E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 5.30E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.50E+4nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 5.60E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+4nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6.20E+4nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6.30E+4nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 6.30E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6.50E+4nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 6.50E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6.50E+4nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 6.50E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataKi: 6.60E+4nMAssay Description:Binding affinity to human CRBN TBD assessed as inhibition constant using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 7.40E+4nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 7.50E+4nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 8.60E+4nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 8.70E+4nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 9.00E+4nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 9.30E+4nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.07E+5nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.14E+5nMAssay Description:Binding affinity to human CRBN TBD using BODIPY-uracil as substrate by MST assayMore data for this Ligand-Target Pair