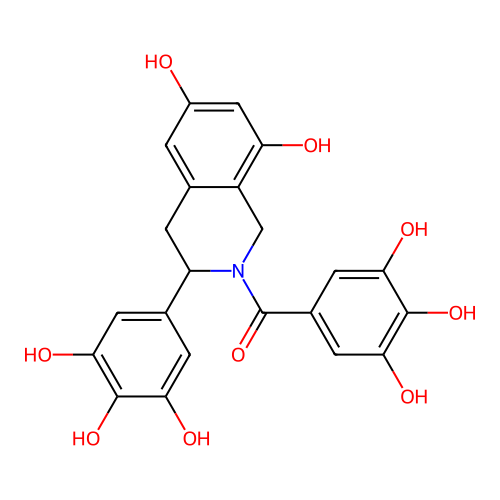
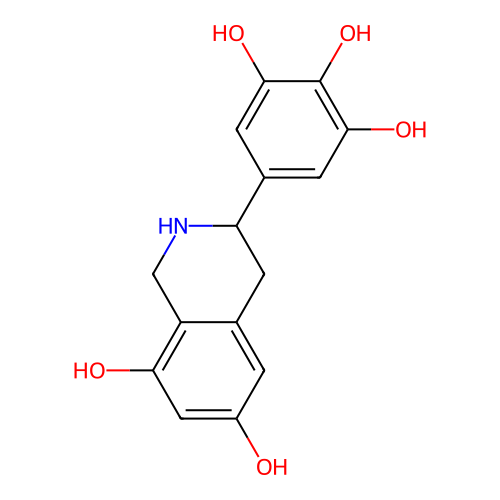
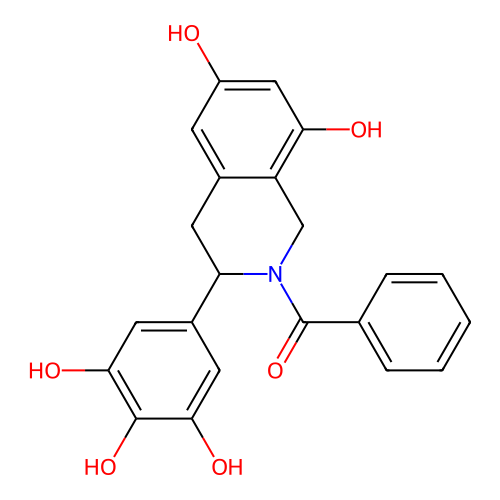
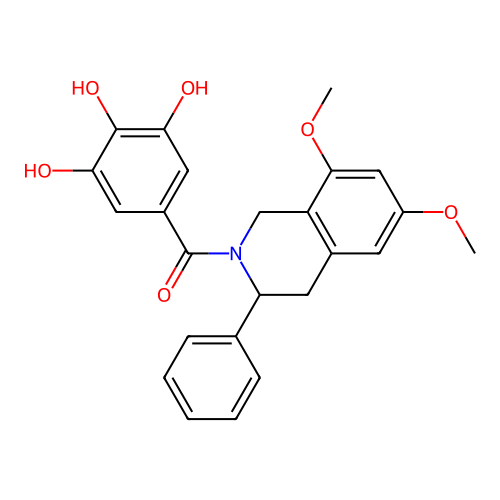
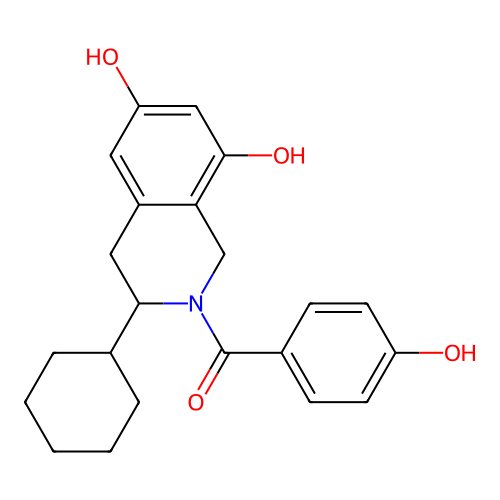
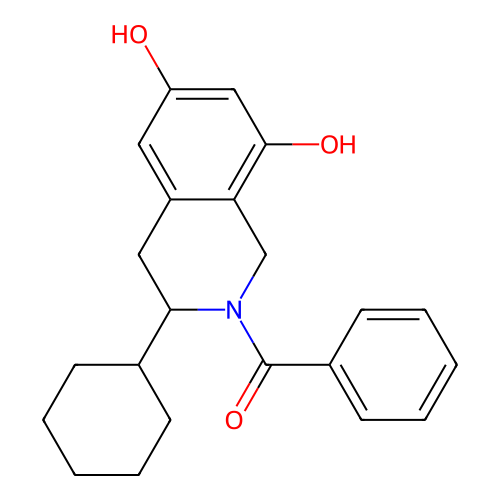
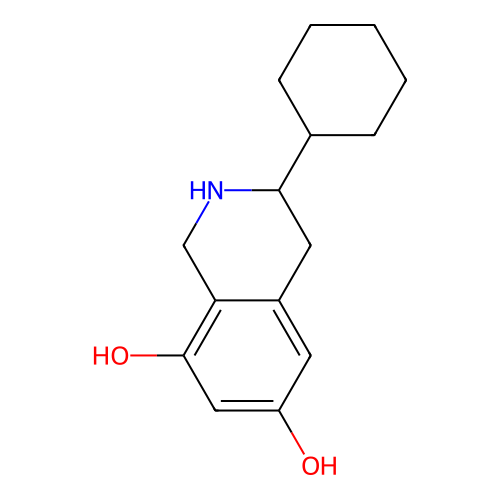
Report error Found 21 Enz. Inhib. hit(s) with all data for entry = 50021253
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Rat)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 20nMAssay Description:Inhibition of recombinant 6His-tagged rat DYRK1A (1 to 502 residues) expressed in Escherichia coli BL21 (DE3) cells using KISGRLSPIMTEQ as substrate ...More data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 80nMAssay Description:Competitive inhibition of DYRK1A (unknown origin)More data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 200nMAssay Description:Inhibition of DYRK1A (unknown origin) using KISGRLSPIMTEQ as substrate incubated for 30 mins in presence of ATP by UFLC-based fluorimetric analysisMore data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 240nMAssay Description:Inhibition of C-terminal FLAG-tagged recombinant human DYRK1A (126 to 490 residues) expressed in Escherichia coli incubated for 10 mins in presence o...More data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 300nMAssay Description:Inhibition of human DYRK1A using KKISGRLSPIMTEQ as substrate incubated for 40 mins in presence of [gamma-33P]-ATP by radioactivity based assayMore data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 330nMAssay Description:Inhibition of human DYRK1A using KKISGRLSPIMTEQ as substrate incubated for 40 mins in presence of [gamma-33P]-ATP by radioactivity based assayMore data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 500nMAssay Description:Inhibition of DYRK1A (unknown origin) using KISGRLSPIMTEQ as substrate incubated for 30 mins in presence of ATP by UFLC-based fluorimetric analysisMore data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 8.20E+3nMAssay Description:Inhibition of DYRK1A (unknown origin) using KISGRLSPIMTEQ as substrate incubated for 30 mins in presence of ATP by UFLC-based fluorimetric analysisMore data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 8.40E+3nMAssay Description:Inhibition of DYRK1A (unknown origin) using KISGRLSPIMTEQ as substrate incubated for 30 mins in presence of ATP by UFLC-based fluorimetric analysisMore data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 1.35E+4nMAssay Description:Inhibition of DYRK1A (unknown origin) using KISGRLSPIMTEQ as substrate incubated for 30 mins in presence of ATP by UFLC-based fluorimetric analysisMore data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 1.57E+4nMAssay Description:Inhibition of DYRK1A (unknown origin) using KISGRLSPIMTEQ as substrate incubated for 30 mins in presence of ATP by UFLC-based fluorimetric analysisMore data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of DYRK1A (unknown origin) using KISGRLSPIMTEQ as substrate incubated for 30 mins in presence of ATP by UFLC-based fluorimetric analysisMore data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of DYRK1A (unknown origin) using KISGRLSPIMTEQ as substrate incubated for 30 mins in presence of ATP by UFLC-based fluorimetric analysisMore data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of DYRK1A (unknown origin) using KISGRLSPIMTEQ as substrate incubated for 30 mins in presence of ATP by UFLC-based fluorimetric analysisMore data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of DYRK1A (unknown origin) using KISGRLSPIMTEQ as substrate incubated for 30 mins in presence of ATP by UFLC-based fluorimetric analysisMore data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of DYRK1A (unknown origin) using KISGRLSPIMTEQ as substrate incubated for 30 mins in presence of ATP by UFLC-based fluorimetric analysisMore data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of DYRK1A (unknown origin) using KISGRLSPIMTEQ as substrate incubated for 30 mins in presence of ATP by UFLC-based fluorimetric analysisMore data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of DYRK1A (unknown origin) using KISGRLSPIMTEQ as substrate incubated for 30 mins in presence of ATP by UFLC-based fluorimetric analysisMore data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of DYRK1A (unknown origin) using KISGRLSPIMTEQ as substrate incubated for 30 mins in presence of ATP by UFLC-based fluorimetric analysisMore data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of DYRK1A (unknown origin) using KISGRLSPIMTEQ as substrate incubated for 30 mins in presence of ATP by UFLC-based fluorimetric analysisMore data for this Ligand-Target Pair
TargetDual specificity tyrosine-phosphorylation-regulated kinase 1A(Human)
Sorbonne Universit£
Curated by ChEMBL
Sorbonne Universit£
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of DYRK1A (unknown origin) using KISGRLSPIMTEQ as substrate incubated for 30 mins in presence of ATP by UFLC-based fluorimetric analysisMore data for this Ligand-Target Pair