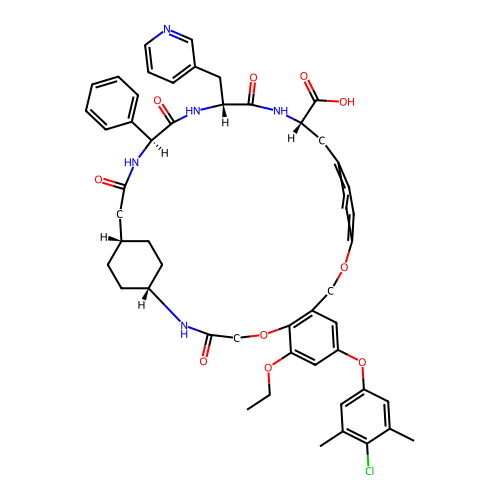
Report error Found 64 Enz. Inhib. hit(s) with all data for entry = 50021451
Affinity DataIC50: 34nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 43nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 59nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 64nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 78nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 80nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 80nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 90nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 98nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 101nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 124nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 137nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 174nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 179nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 212nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 227nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 234nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 253nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 271nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 298nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 382nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 678nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.37E+3nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.43E+3nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.44E+3nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.30E+3nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.30E+3nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.30E+3nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.60E+3nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.70E+3nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 7.10E+3nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 7.30E+3nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 7.35E+3nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 7.70E+3nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+3nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 9.10E+3nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 9.40E+3nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Bcl-xl (1-212) (unknown origin) using BIOTIN-AHXAHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Bcl-xl (1-212) (unknown origin) using BIOTIN-AHXAHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Bcl-xl (1-212) (unknown origin) using BIOTIN-AHXAHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Bcl-xl (1-212) (unknown origin) using BIOTIN-AHXAHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Bcl-xl (1-212) (unknown origin) using BIOTIN-AHXAHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+4nMAssay Description:Inhibition of Mcl-1 (172-327) (unknown origin) using BIOTIN-AHXGQVGRQLAIIGDDINR-amide peptide by TR-FRET based BAK peptide displacement assayMore data for this Ligand-Target Pair