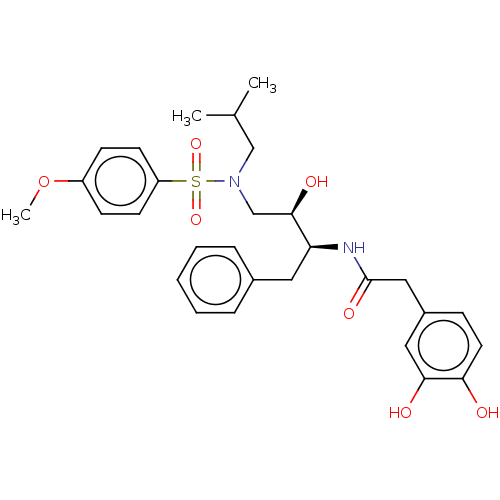
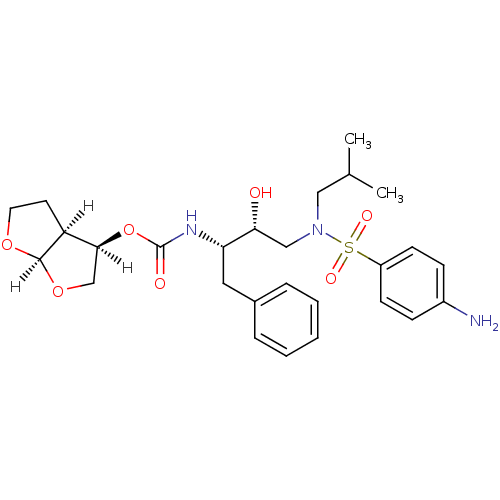
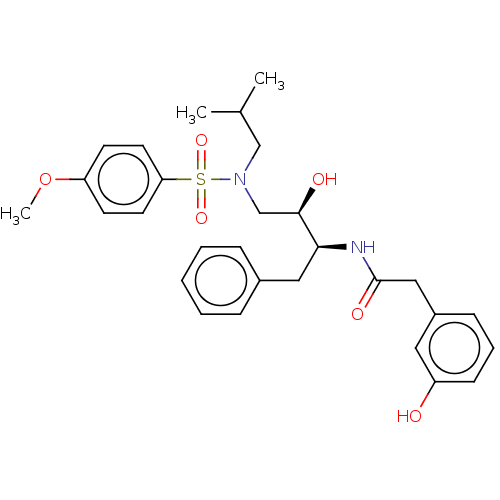
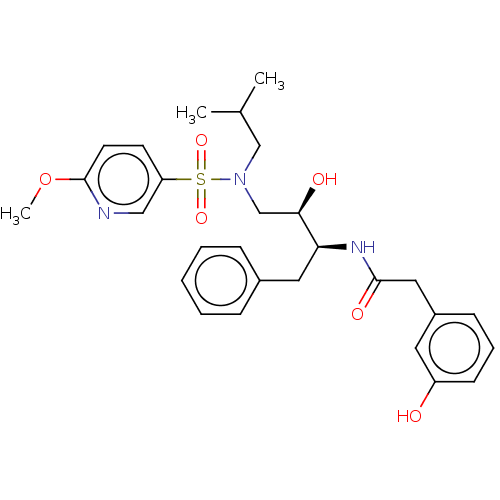
Report error Found 33 Enz. Inhib. hit(s) with all data for entry = 50019606
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
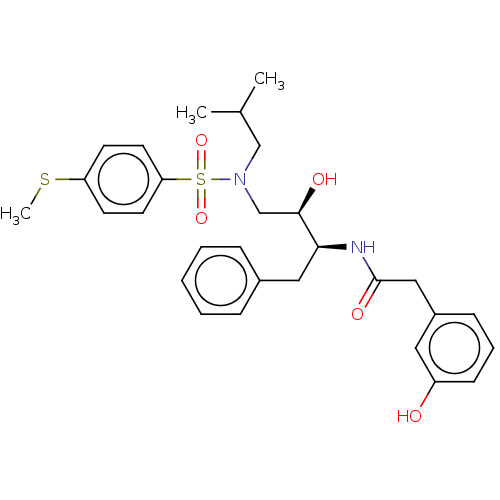
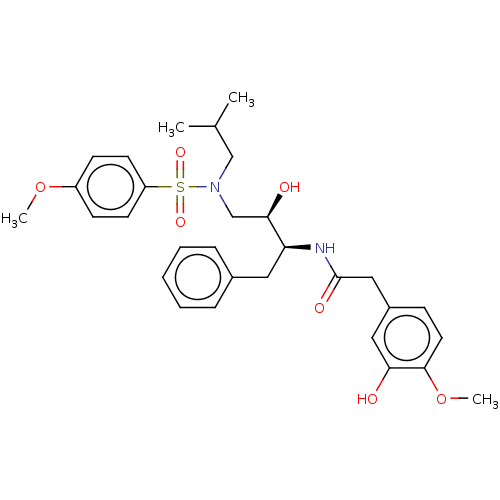
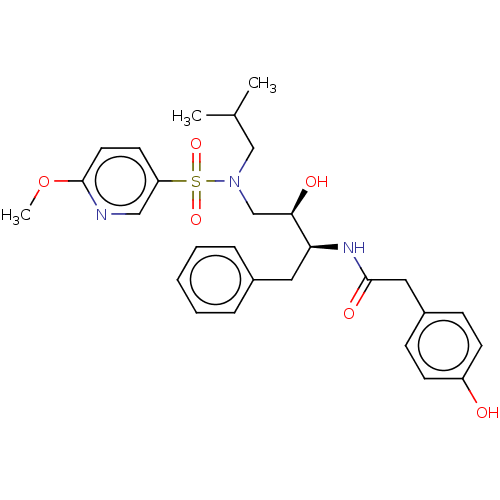
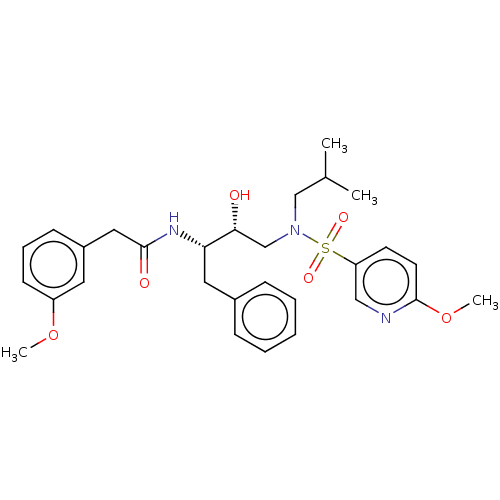
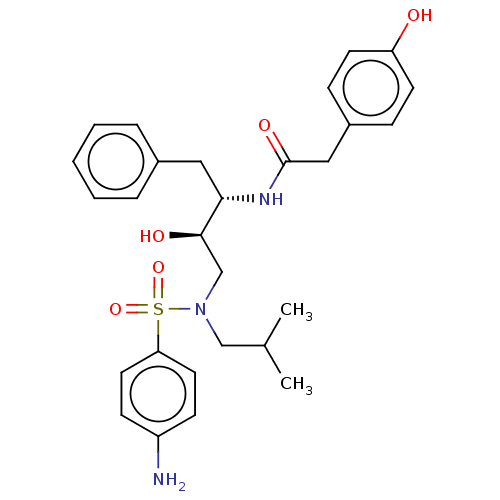
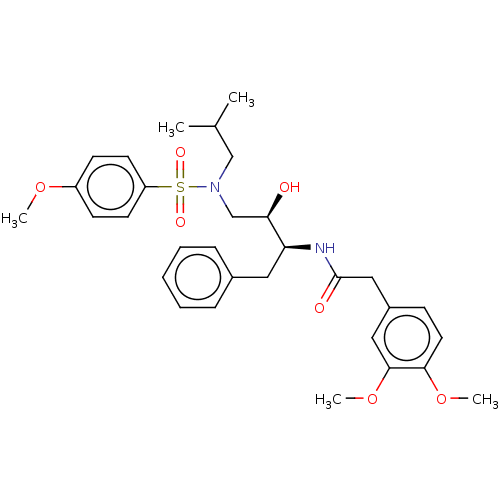
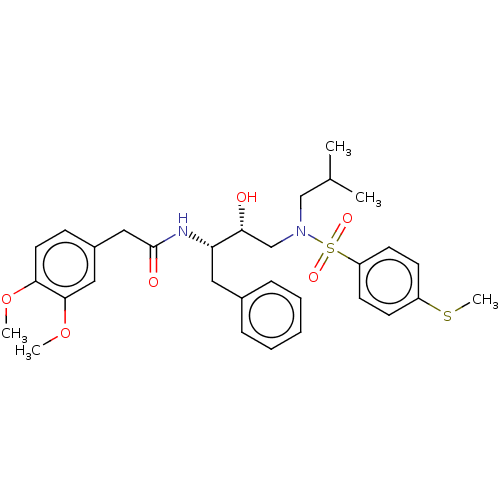
Affinity DataIC50: 0.540nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
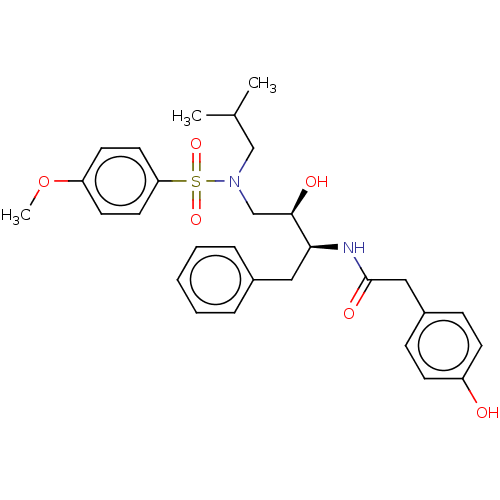
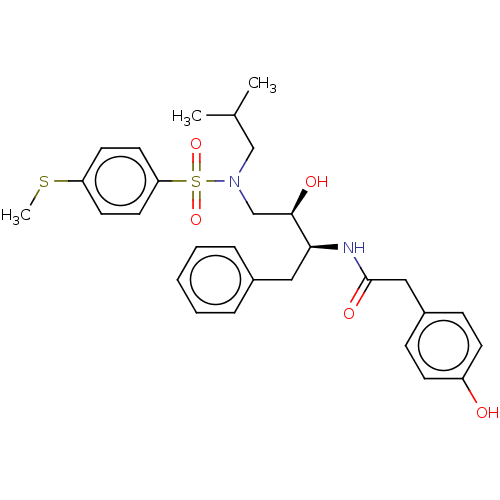
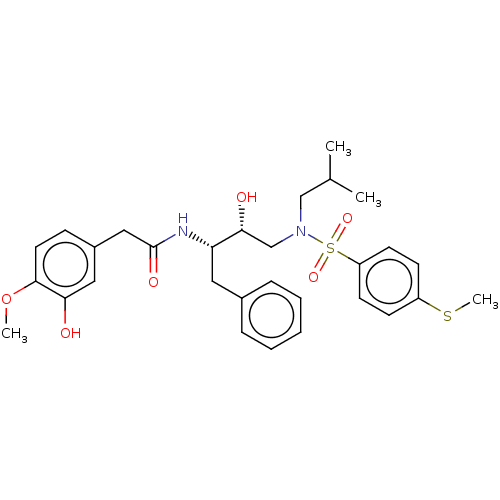
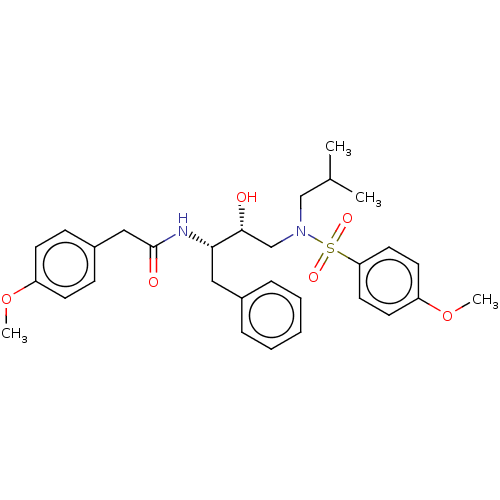
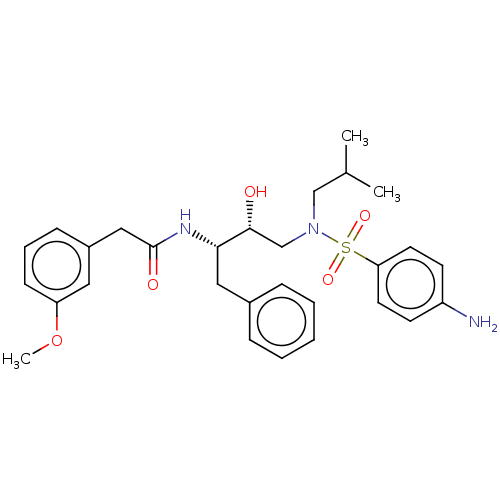
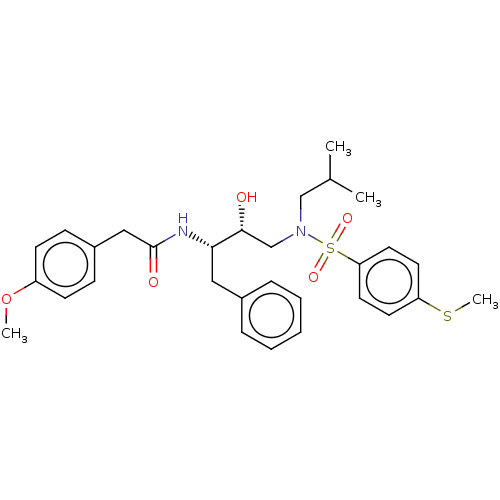
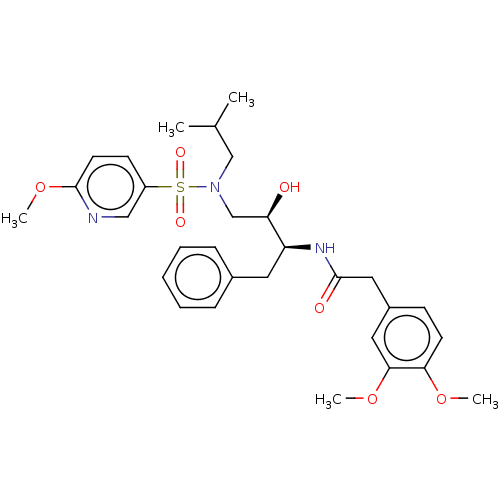
Affinity DataIC50: 0.870nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
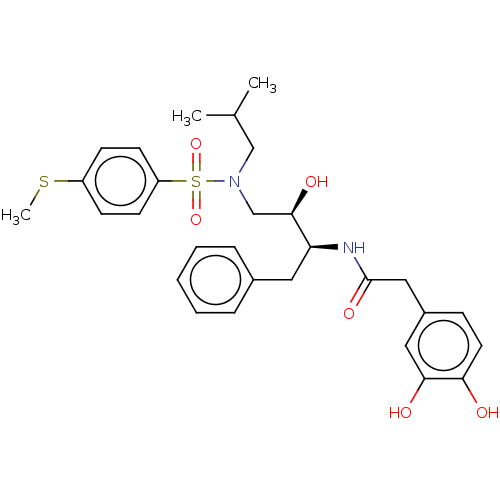
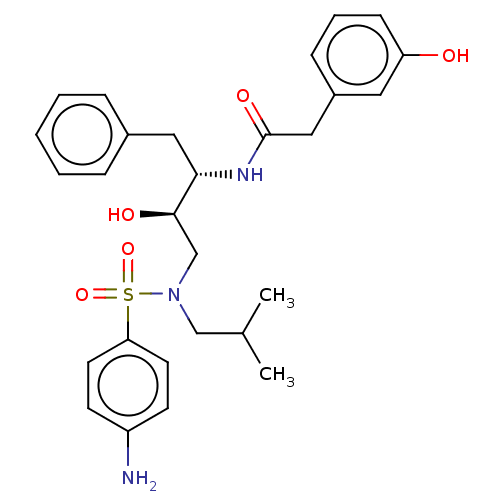
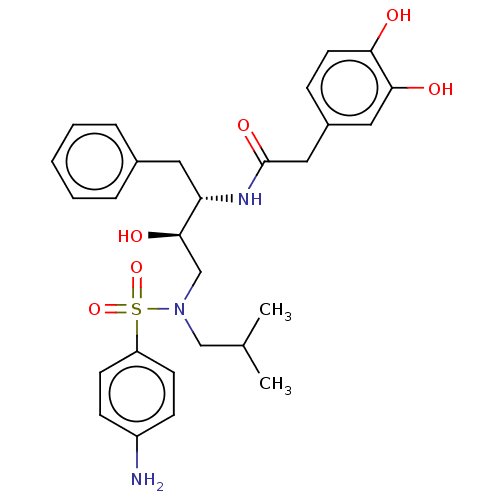
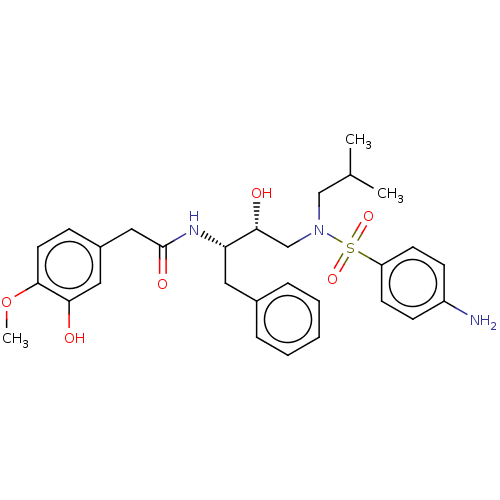
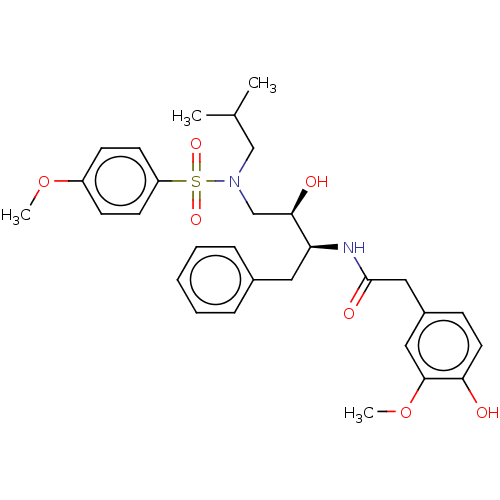
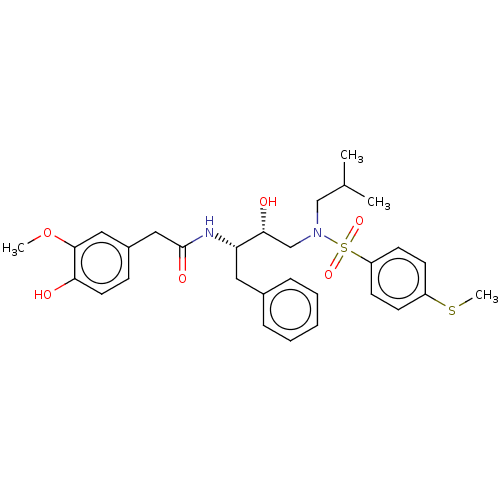
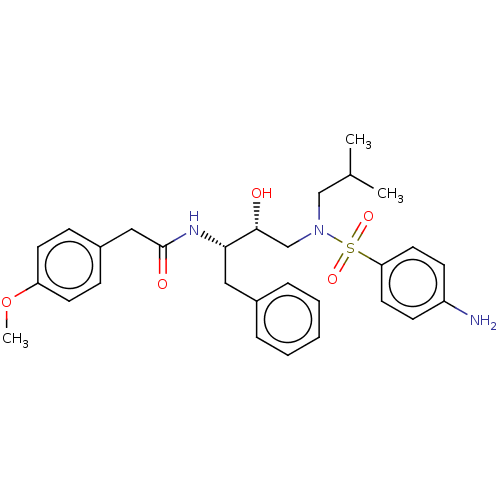
Affinity DataIC50: 0.870nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
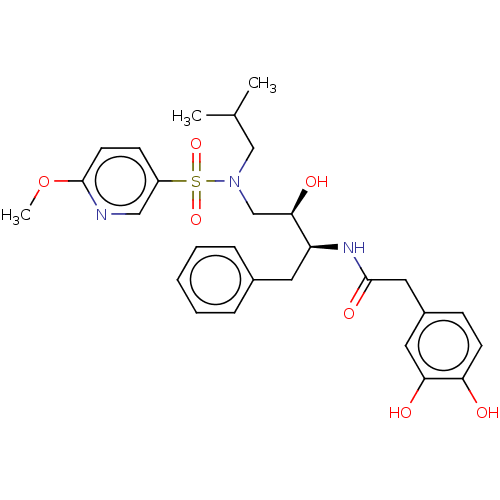
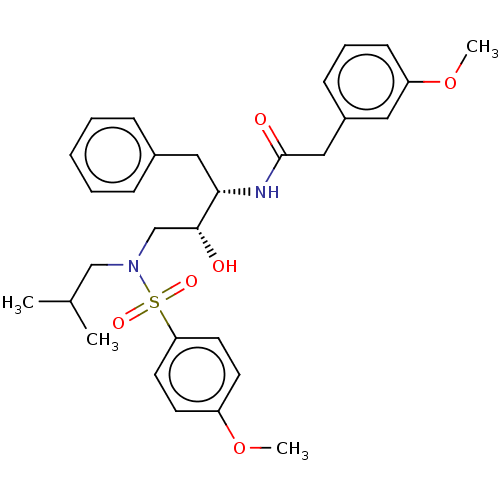
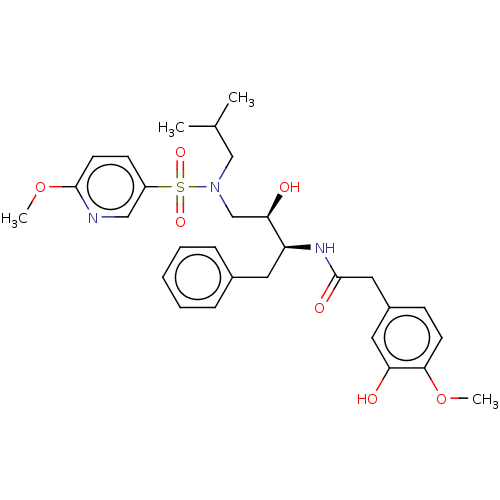
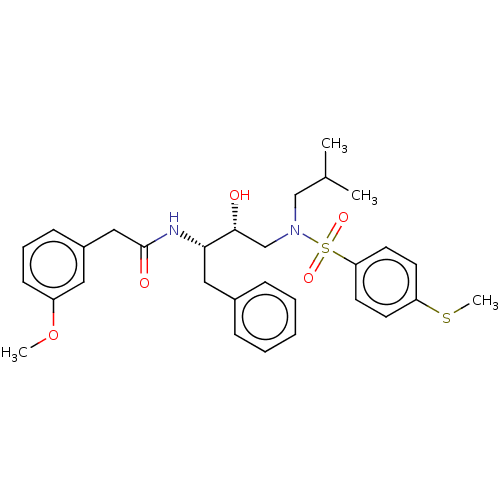
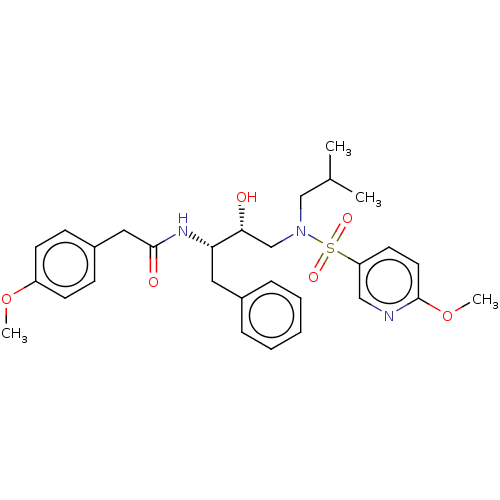
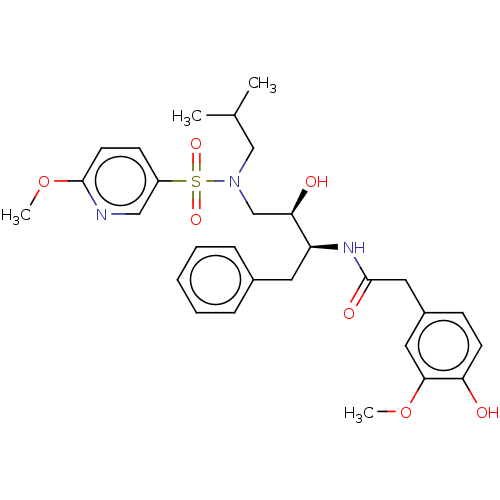
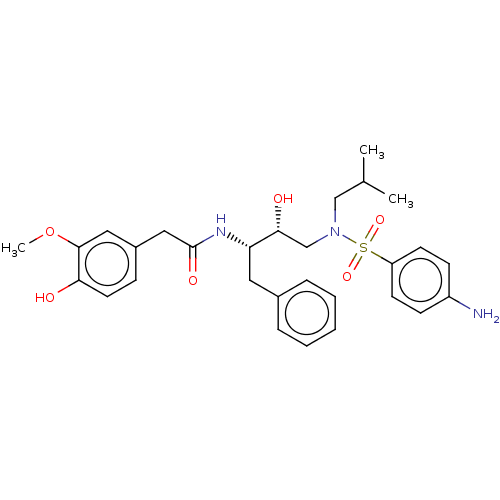
Affinity DataIC50: 1.20nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 2.20nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 2.20nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 2.30nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 2.90nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 4.20nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 5.60nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
Ligand InfoSimilars
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 7nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 7.30nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 9.30nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 9.5nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 13nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 13nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 21nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 28nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 30nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 32nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 35nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
Ligand InfoSimilars
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 50nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 52nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 85nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 86nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 116nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 170nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 171nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 176nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 377nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 379nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Xuzhou Medical University
Curated by ChEMBL
Xuzhou Medical University
Curated by ChEMBL
Affinity DataIC50: 1.02E+3nMAssay Description:Inhibition of HIV-1 protease by fluorescence resonance energy transfer analysisMore data for this Ligand-Target Pair