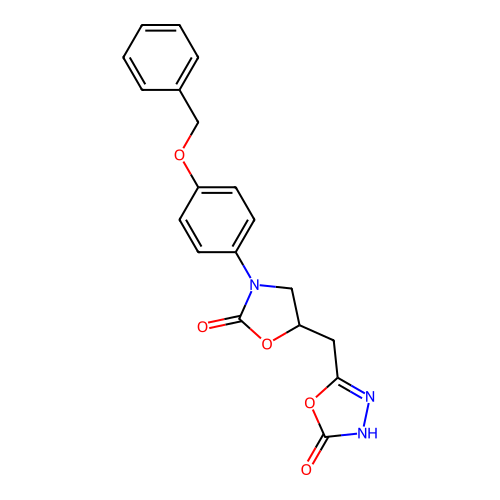
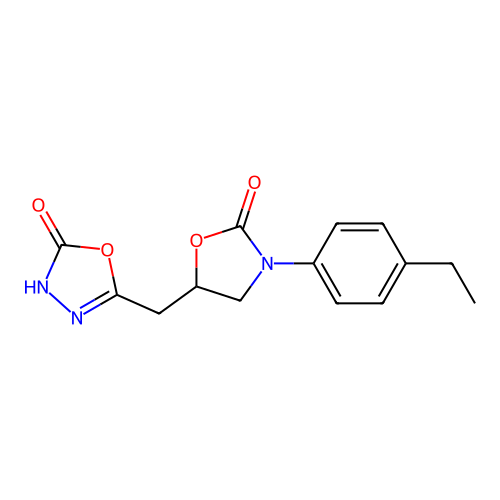
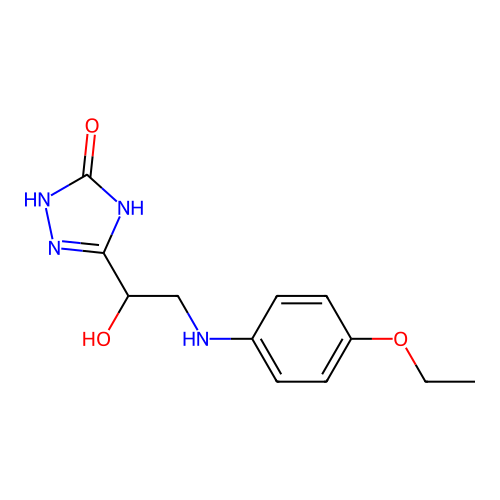
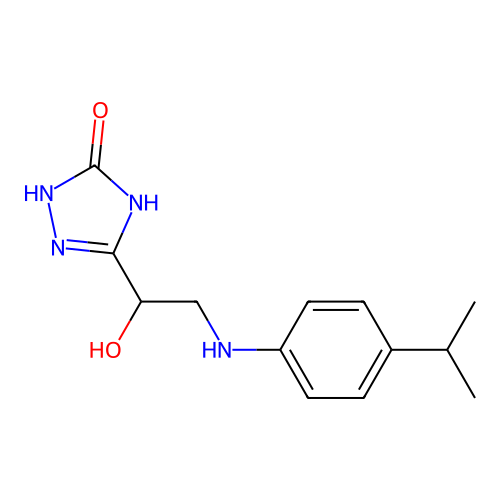
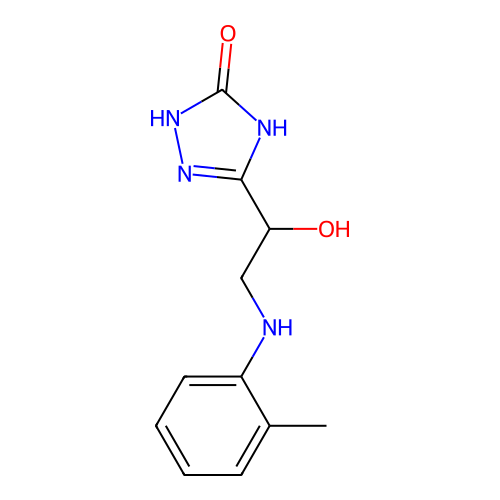
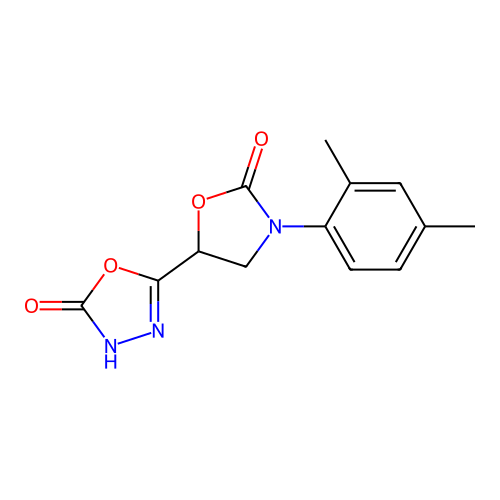
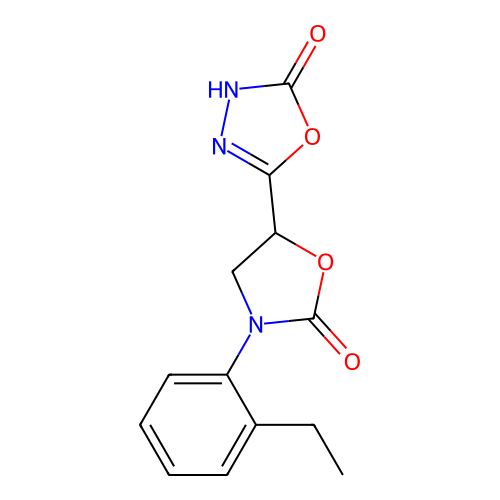
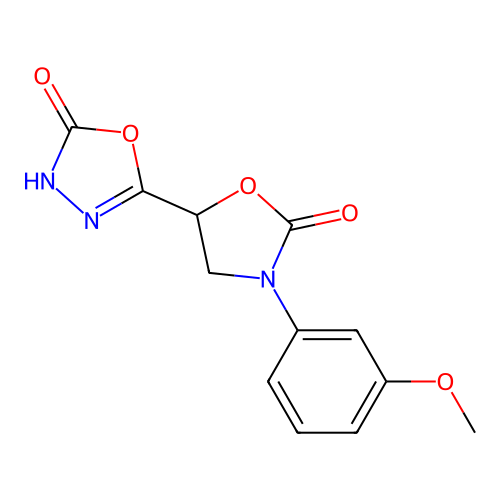
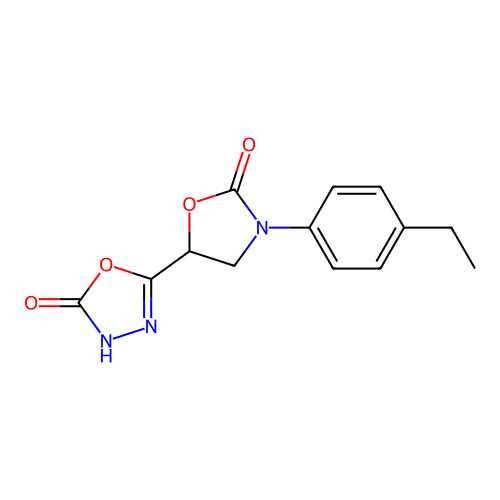
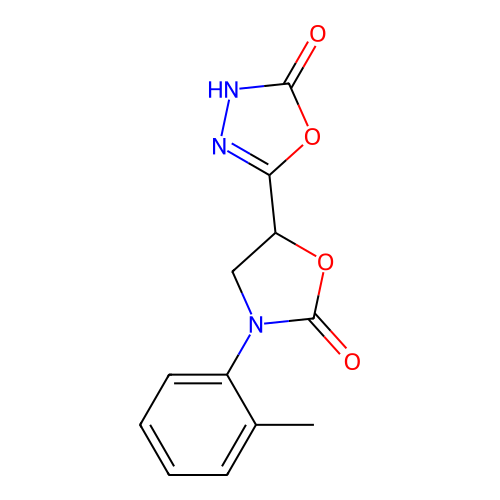
Report error Found 38 Enz. Inhib. hit(s) with all data for entry = 50021762
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
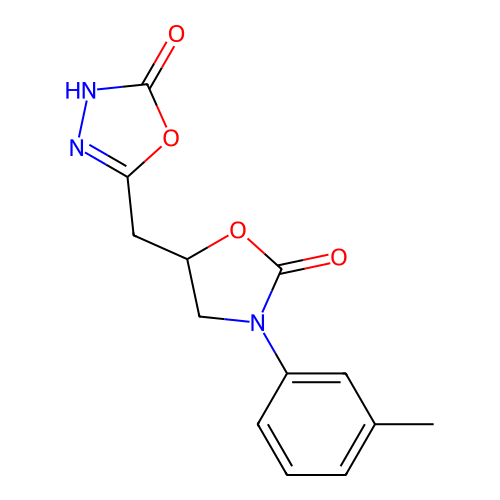
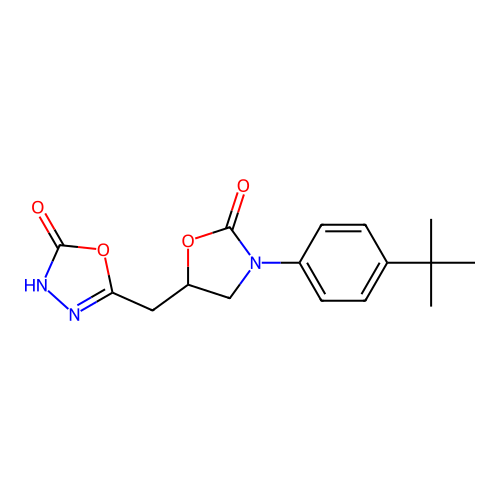
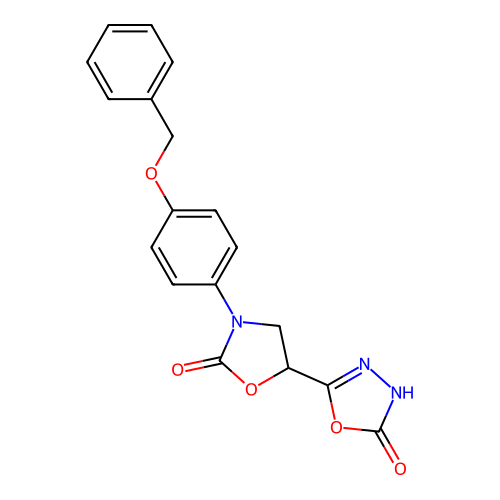
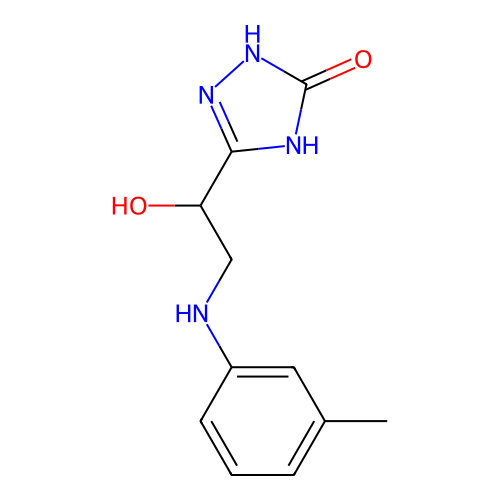
Affinity DataKi: 240nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
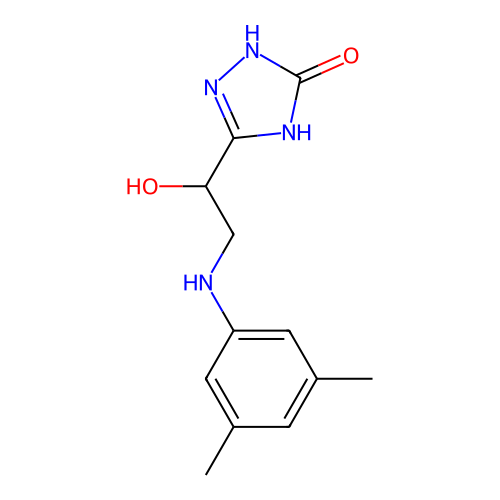
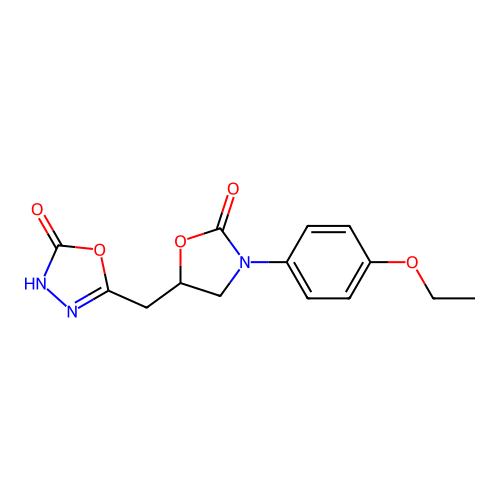
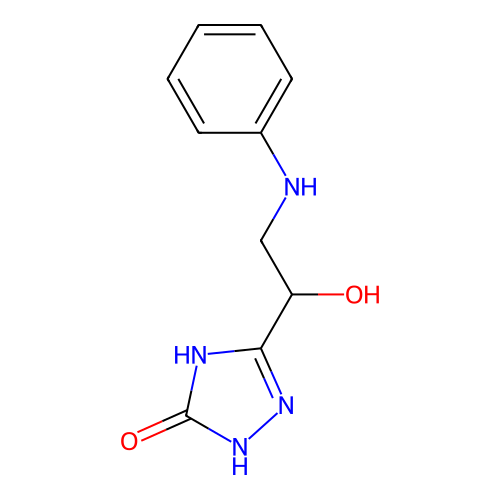
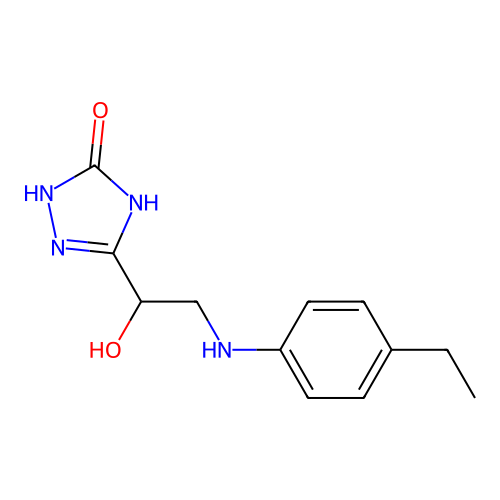
Affinity DataIC50: 1.21E+3nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
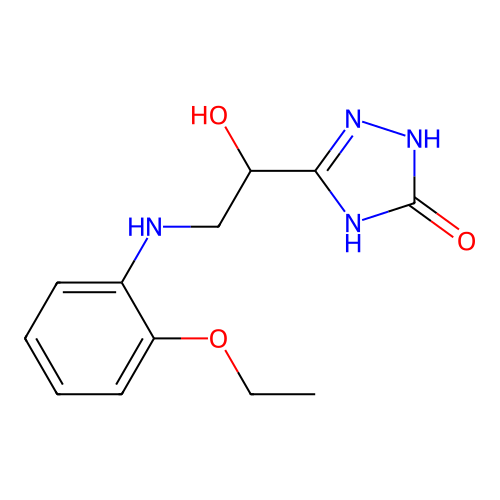
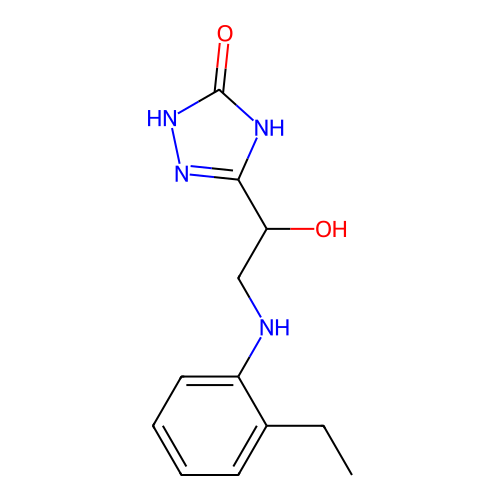
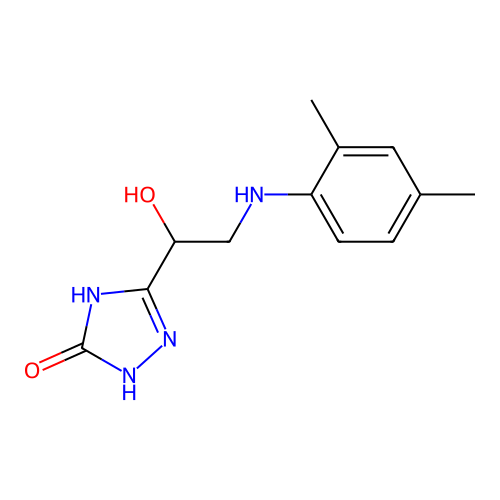
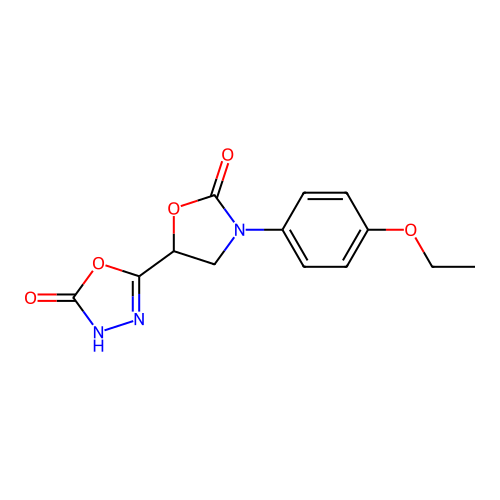
Affinity DataKi: 2.24E+3nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
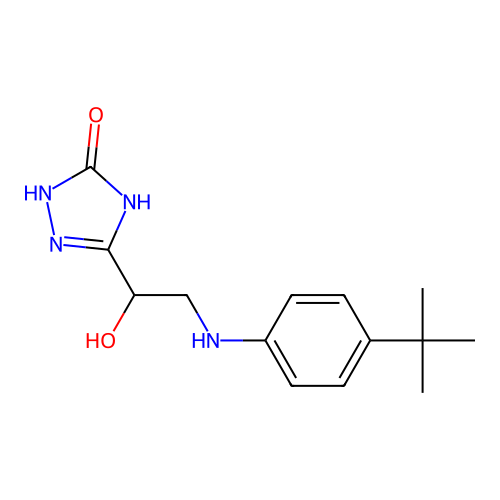
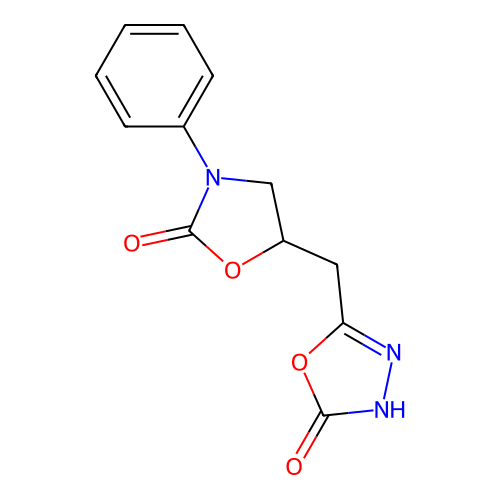
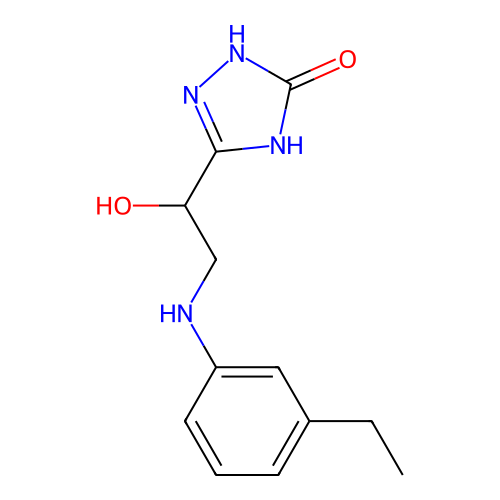
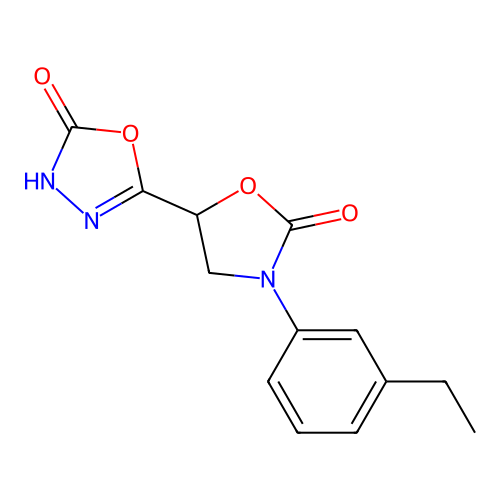
Affinity DataIC50: 6.31E+3nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataKi: 6.75E+3nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 1.01E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 1.91E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 2.07E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 2.12E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 2.21E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 2.46E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 2.66E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 3.06E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 3.07E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 3.88E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 4.02E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 4.57E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 4.61E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 4.68E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 5.14E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 5.68E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 6.13E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 6.26E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 6.52E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 7.31E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 8.52E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 8.75E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 9.05E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 9.35E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 9.87E+4nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 1.02E+5nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 1.08E+5nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 1.08E+5nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 1.10E+5nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 1.16E+5nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 1.51E+5nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 1.62E+5nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 1.86E+5nMAssay Description:Inhibition of Helicobacter pylori urease using urea as substrate assessed as decrease in ammonia production incubated for 0.5 hrs by indole phenol me...More data for this Ligand-Target Pair
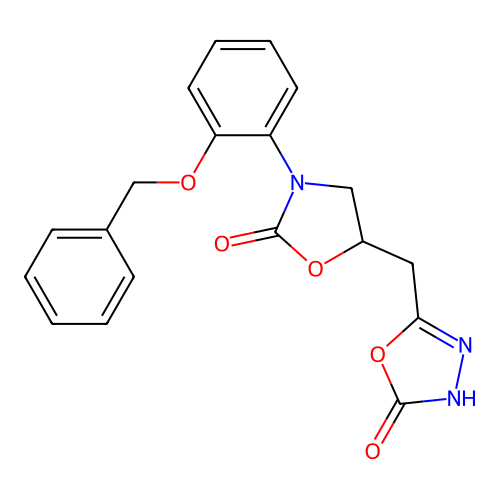
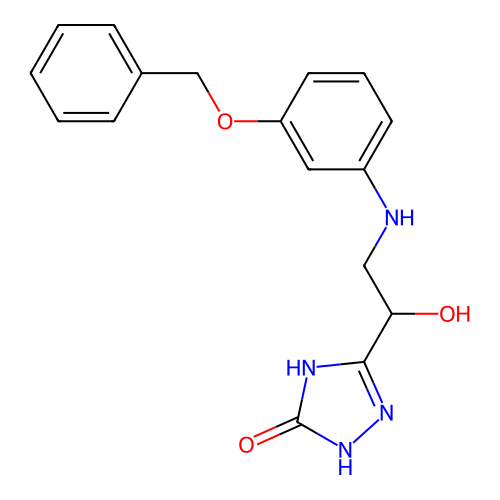
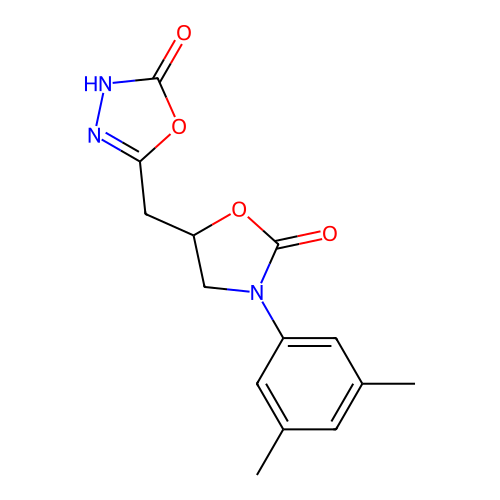
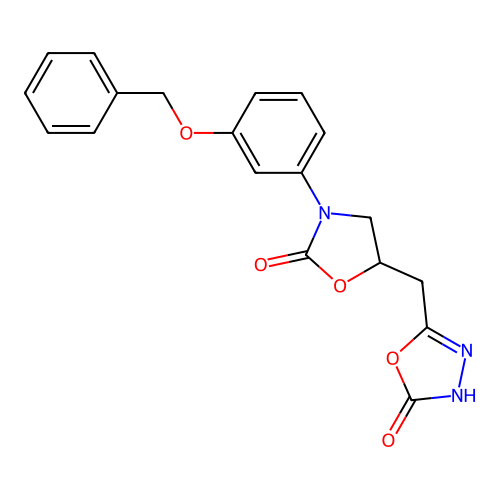
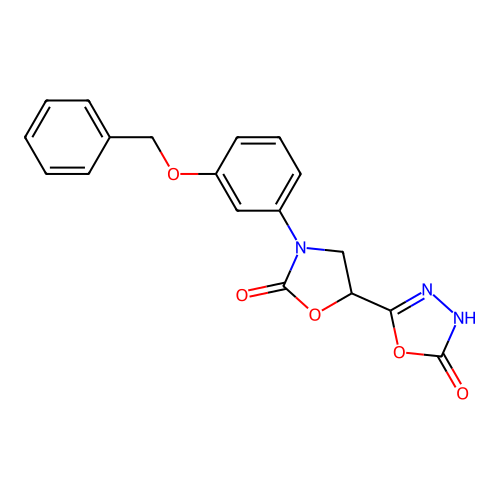
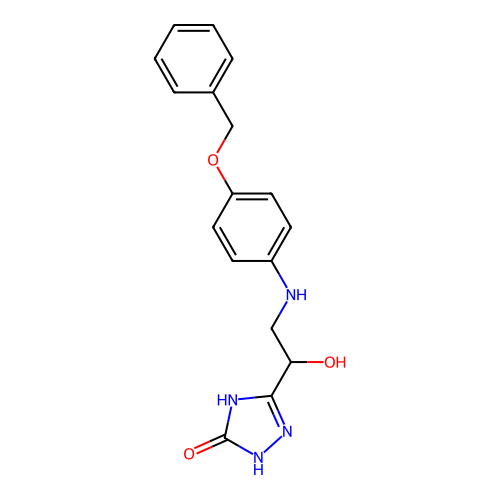
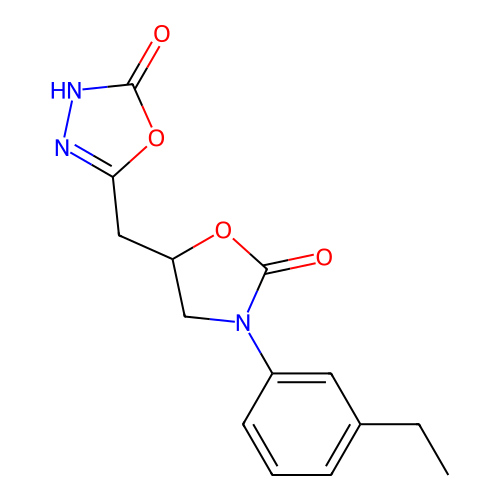
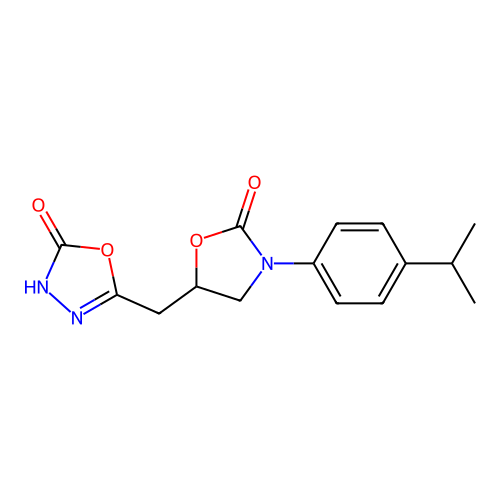
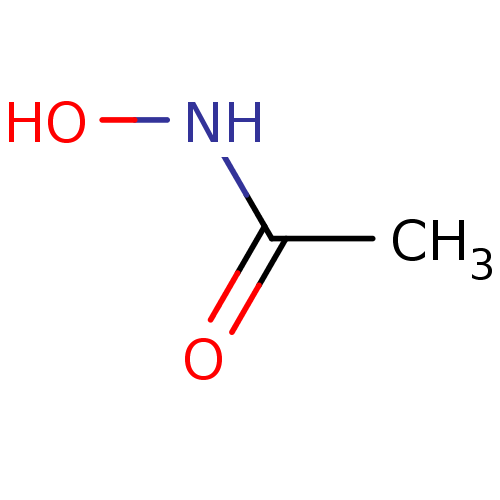
 3D Structure (crystal)
3D Structure (crystal)