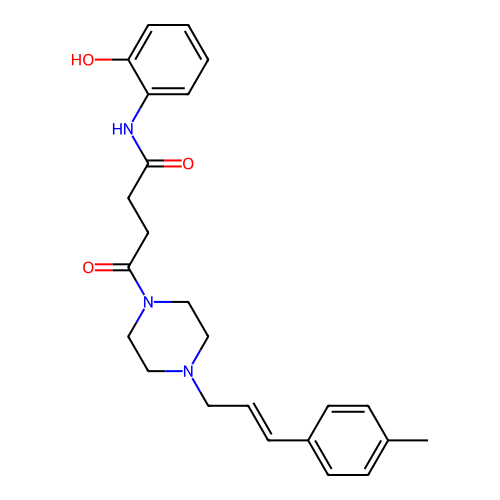
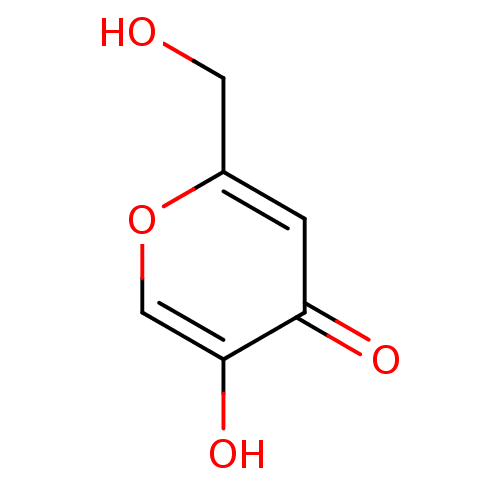
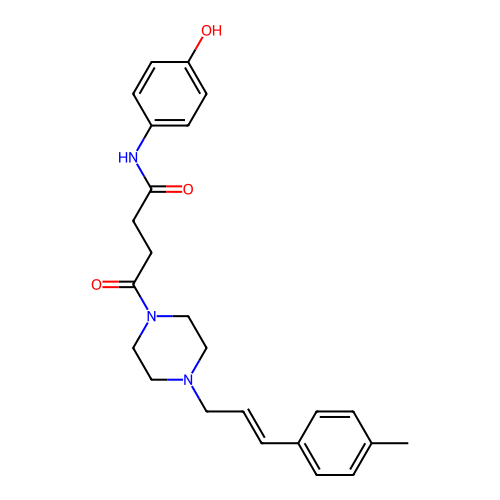
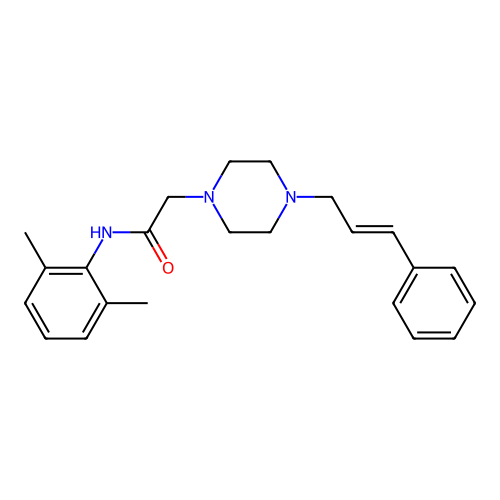
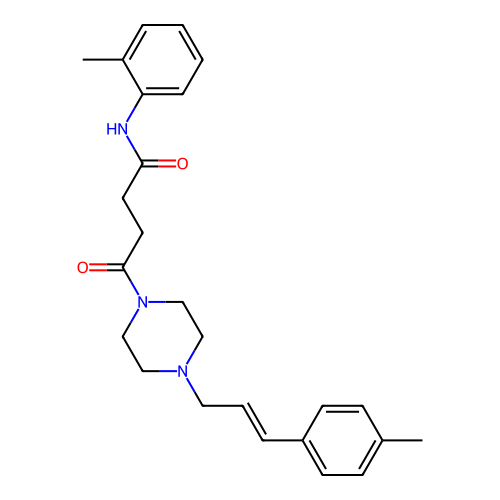
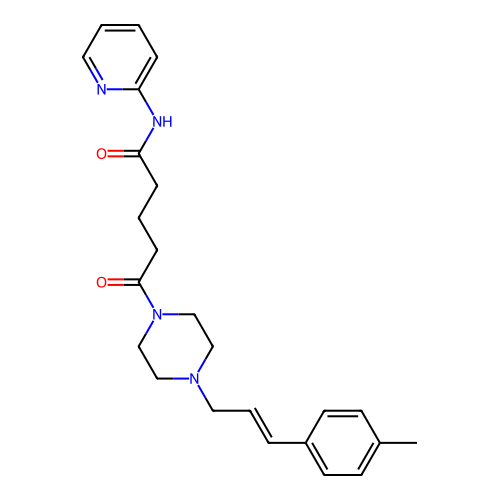
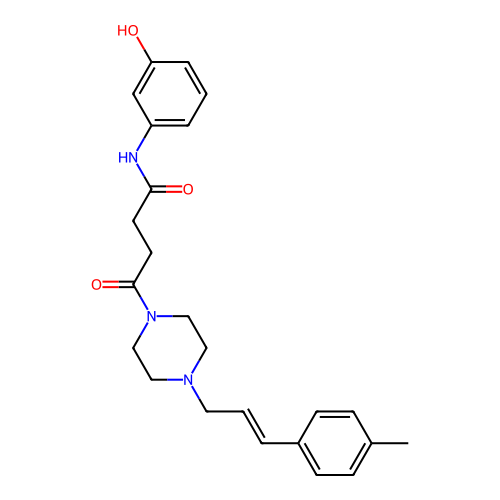
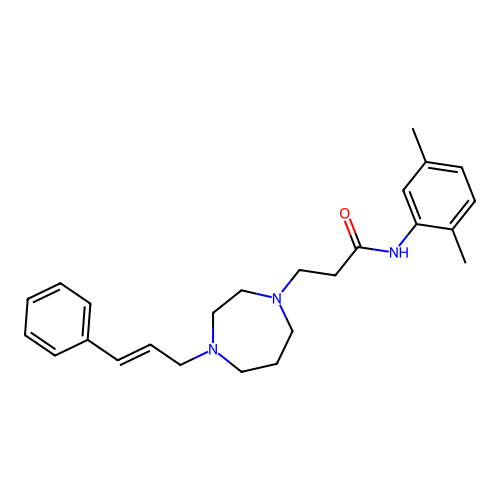
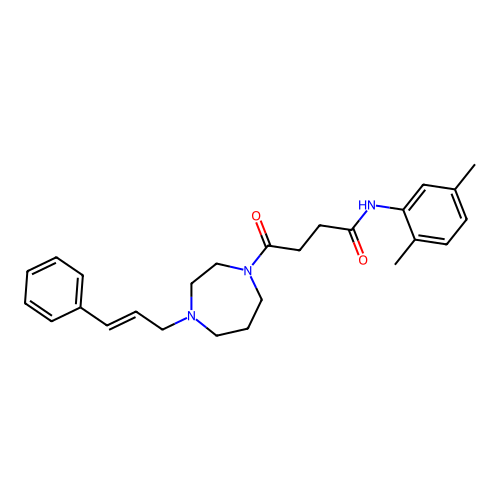
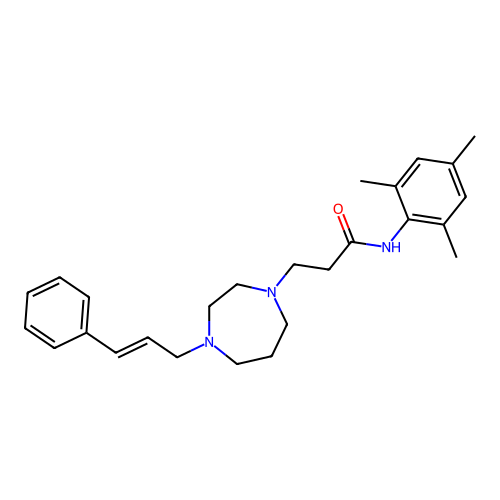
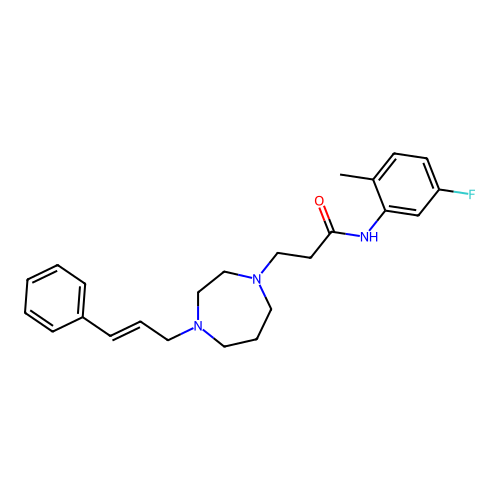
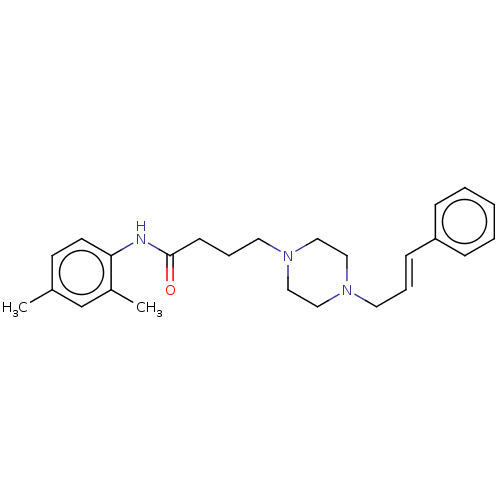
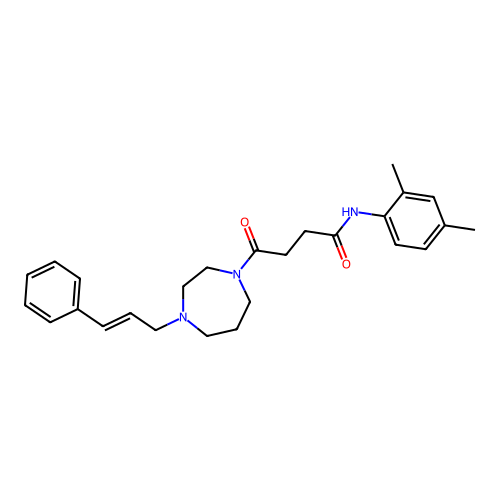
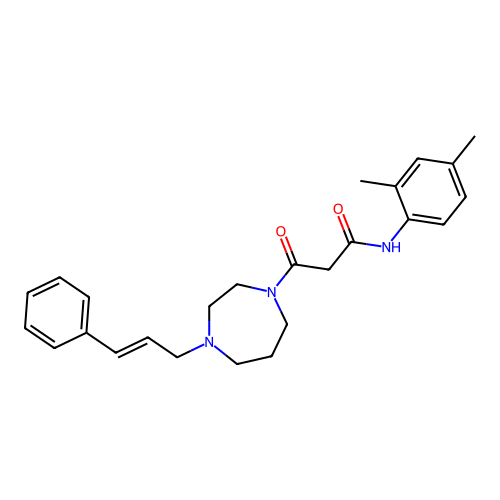
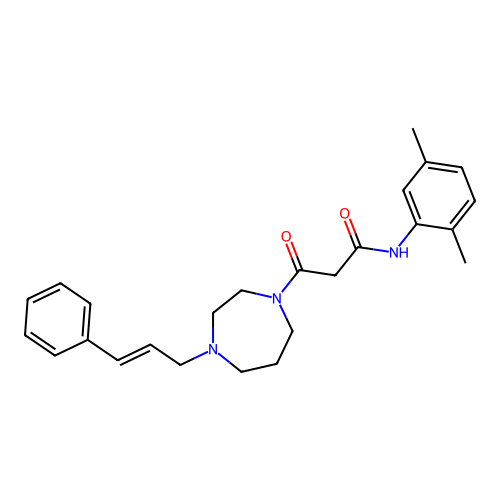
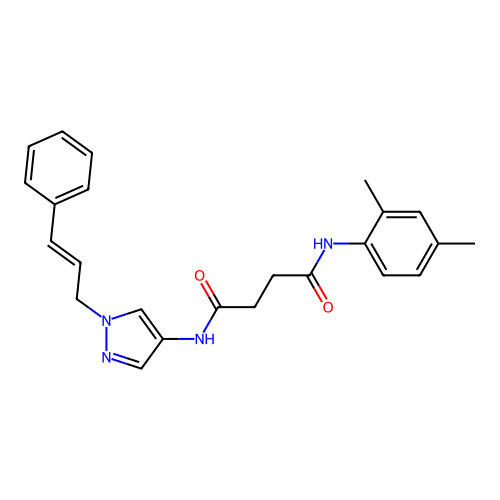
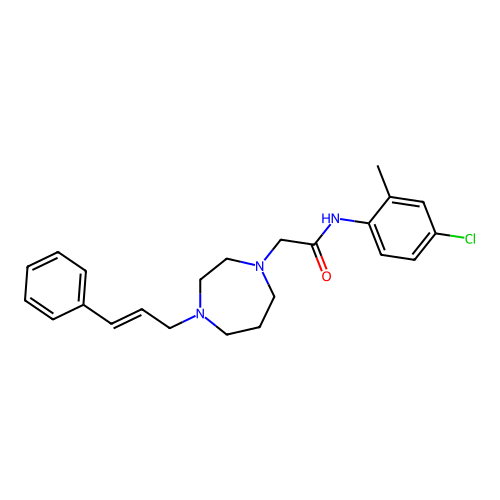
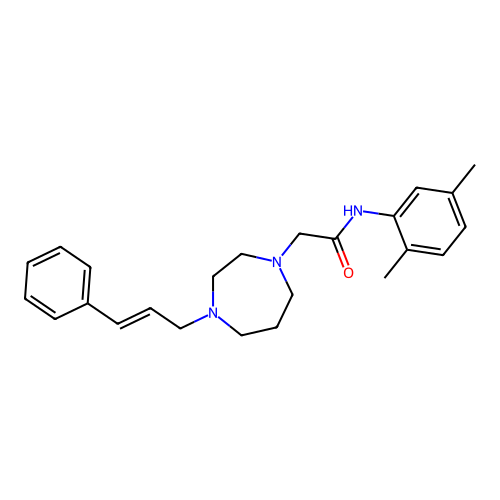
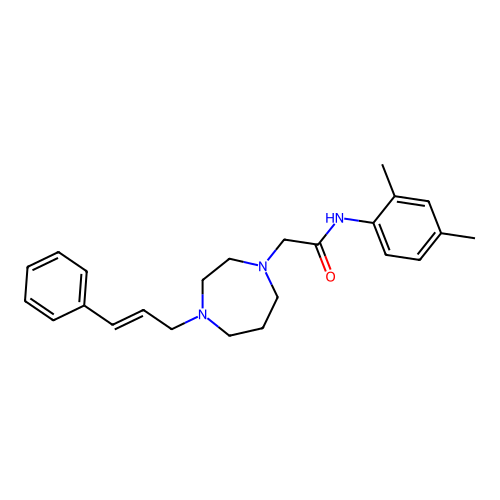
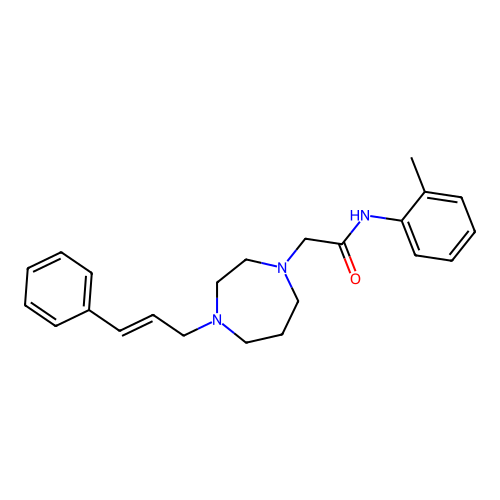
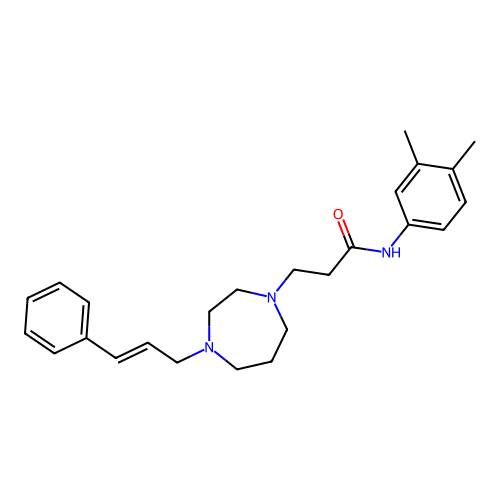
Report error Found 52 Enz. Inhib. hit(s) with all data for entry = 50021057
Affinity DataIC50: 1.15E+4nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 1.73E+4nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 3.10E+4nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 3.86E+4nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 4.03E+4nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 8.80E+4nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 1.32E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 1.73E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 1.93E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataKd: 1.98E+5nMAssay Description:Binding affinity to tyrosinase (unknown origin) assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.04E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 2.17E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 2.34E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 2.45E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 2.49E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 2.54E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 2.55E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 3.38E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 3.78E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 3.93E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 4.65E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 4.65E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 5.36E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 5.55E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 6.40E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 6.53E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 6.81E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 7.32E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 7.45E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase monophenolase activity using L-tyrosine as substrate preincubated for 10 mins followed by substrate addition and me...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+5nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-dopamine as substrate preincubated for 10 mins followed by substrate addition and meas...More data for this Ligand-Target Pair