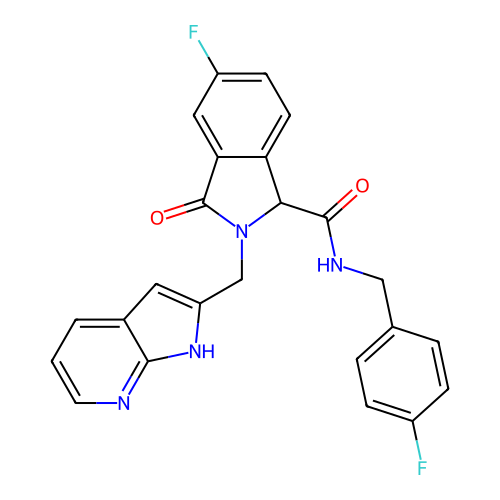
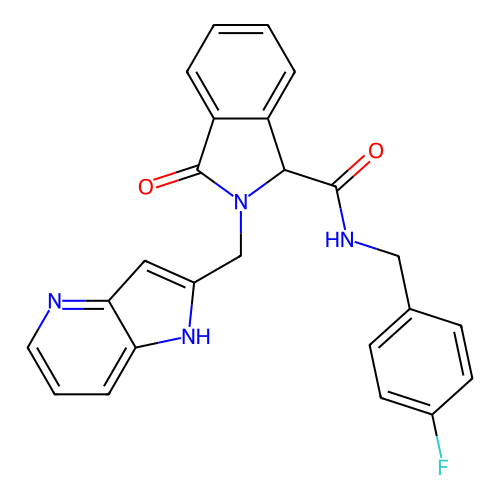
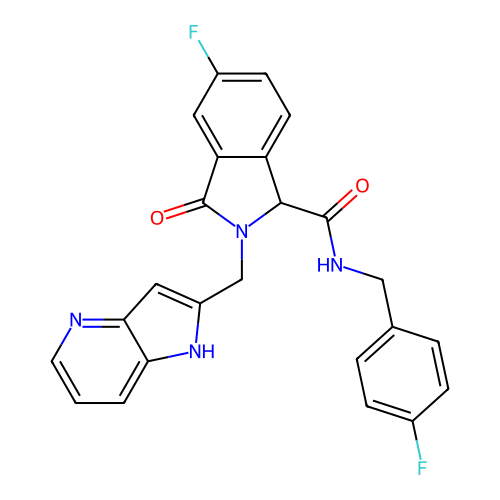
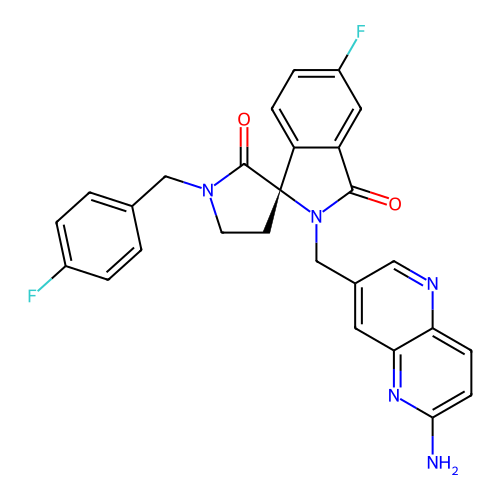
Report error Found 68 Enz. Inhib. hit(s) with all data for entry = 50021894
Affinity DataKd: 1nMAssay Description:Binding affinity to apo PRMT5 (1 to 637 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells assessed as dissociation consta...More data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of PRMT5 in human HCT-116 cells with MTAP KO assessed as decrease in SMDA level incubated for 48 hrs by immunofluorescence analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of PRMT5 in human HCT-116 cells with MTAP KO assessed as decrease in SMDA level incubated for 48 hrs by immunofluorescence analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of PRMT5 in human HCT-116 cells with MTAP KO assessed as decrease in SMDA level incubated for 48 hrs by immunofluorescence analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Inhibition of PRMT5 in human HCT-116 cells with MTAP KO assessed as decrease in SMDA level incubated for 48 hrs by immunofluorescence analysisMore data for this Ligand-Target Pair
Affinity DataKd: 6.30nMAssay Description:Binding affinity to apo PRMT5 (1 to 637 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using SAM as substrate assessed...More data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 14nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 23nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 24nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 27nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 29nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 31nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 44nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 51nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 57nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 60nMAssay Description:Inhibition of PRMT5 in human HCT-116 cells with MTAP KO assessed as decrease in SMDA level incubated for 48 hrs by immunofluorescence analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 66nMAssay Description:Inhibition of PRMT5 in human HCT-116 cells with MTAP KO assessed as decrease in SMDA level incubated for 48 hrs by immunofluorescence analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 68nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 73nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 76nMAssay Description:Inhibition of PRMT5 in human HCT-116 cells with MTAP KO assessed as decrease in SMDA level incubated for 48 hrs by immunofluorescence analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 99nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 107nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 109nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 140nMAssay Description:Inhibition of PRMT5 in human HCT-116 cells with MTAP KO assessed as decrease in SMDA level incubated for 48 hrs by immunofluorescence analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Inhibition of PRMT5 in human HCT-116 cells with MTAP KO assessed as decrease in SMDA level incubated for 48 hrs by immunofluorescence analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 240nMAssay Description:Inhibition of PRMT5 in human HCT-116 cells with MTAP KO assessed as decrease in SMDA level incubated for 48 hrs by immunofluorescence analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Inhibition of PRMT5 in human HCT-116 cells with MTAP KO assessed as decrease in SMDA level incubated for 48 hrs by immunofluorescence analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 370nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 380nMAssay Description:Inhibition of PRMT5 in human HCT-116 cells with MTAP KO assessed as decrease in SMDA level incubated for 48 hrs by immunofluorescence analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 380nMAssay Description:Inhibition of PRMT5 in human HCT-116 cells with MTAP KO assessed as decrease in SMDA level incubated for 48 hrs by immunofluorescence analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 510nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 580nMAssay Description:Inhibition of PRMT5 in human HCT-116 cells with MTAP KO assessed as decrease in SMDA level incubated for 48 hrs by immunofluorescence analysisMore data for this Ligand-Target Pair
Affinity DataKi: 610nMAssay Description:Inhibition of human TSPOMore data for this Ligand-Target Pair
Affinity DataIC50: 660nMAssay Description:Inhibition of PRMT5 in human HCT-116 cells with MTAP KO assessed as decrease in SMDA level incubated for 48 hrs by immunofluorescence analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 680nMAssay Description:Inhibition of PRMT5 in human HCT-116 cells with MTAP KO assessed as decrease in SMDA level incubated for 48 hrs by immunofluorescence analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 970nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMAssay Description:Inhibition of PRMT5 in human HCT-116 cells with MTAP KO assessed as decrease in SMDA level incubated for 48 hrs by immunofluorescence analysisMore data for this Ligand-Target Pair
Affinity DataKd: 1.30E+3nMAssay Description:Binding affinity to apo PRMT5 (1 to 637 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells assessed as dissociation consta...More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.67E+3nMAssay Description:Inhibition of PRMT5 (1 to 637 residues)/MEP504 (2 to 342 residues)(unknown origin) expressed in baculovirus infected Sf21 insect cells using histone ...More data for this Ligand-Target Pair
 3D Structure (crystal)
3D Structure (crystal)