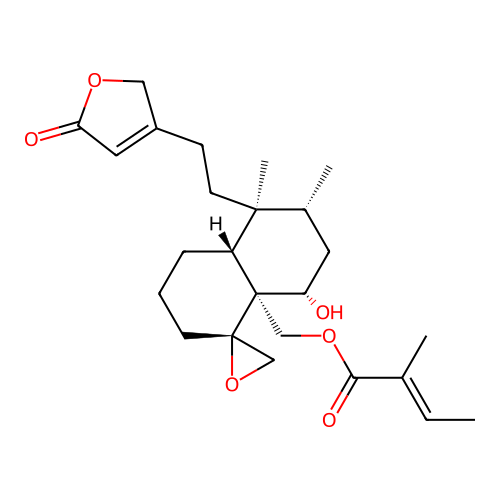
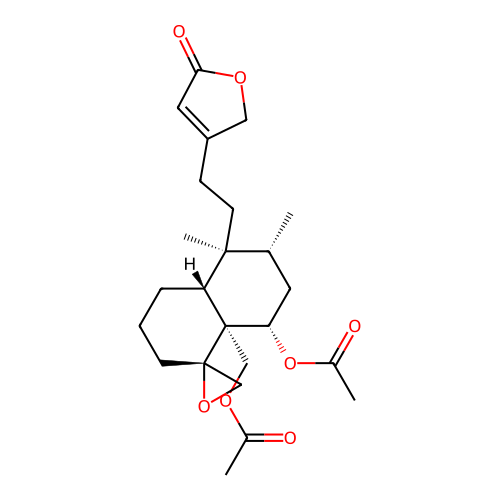
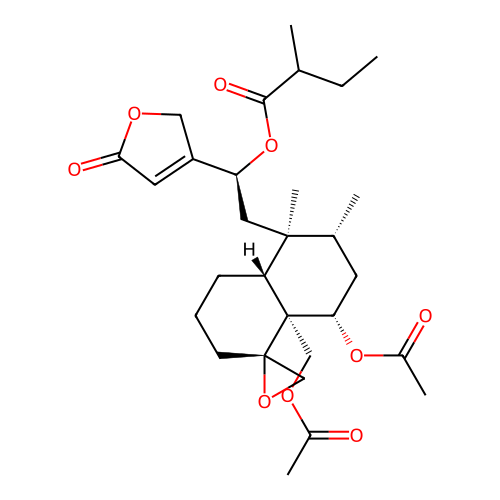
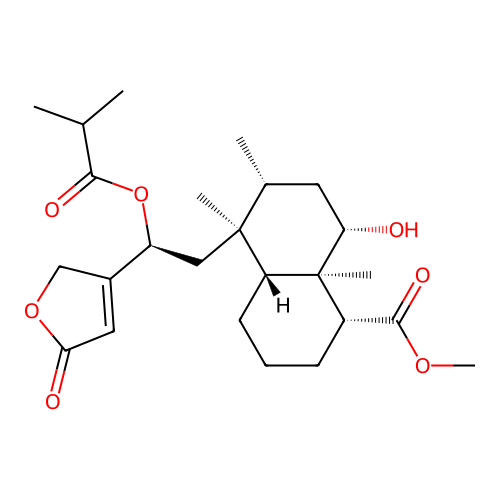
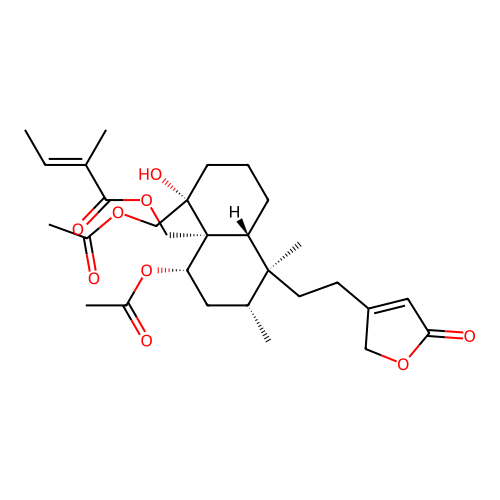
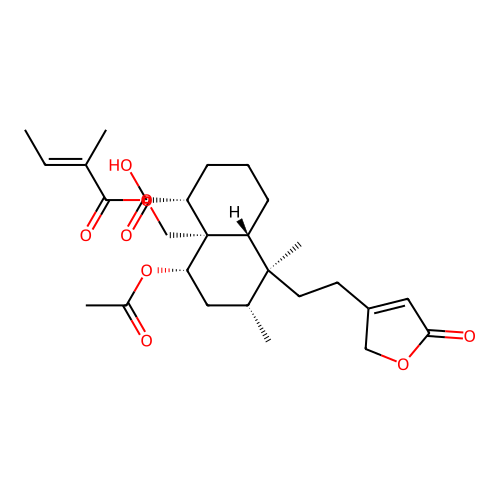
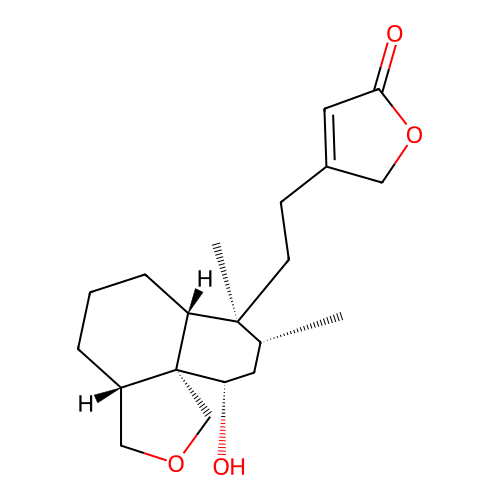
Report error Found 43 Enz. Inhib. hit(s) with all data for entry = 50021994
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataKd: 270nMAssay Description:Binding affinity to PPAR gamma LBD (203 to 476 residues) (unknown origin) extracted from Escherichia coli BL21 (DE3) cells assessed as dissociation c...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 320nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 750nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.26E+3nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.30E+3nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.43E+3nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 2.05E+3nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 2.20E+3nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 2.20E+3nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataKd: 3.38E+3nMAssay Description:Binding affinity to PPAR gamma LBD (203 to 476 residues) (unknown origin) extracted from Escherichia coli BL21 (DE3) cells assessed as dissociation c...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 5.23E+3nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 6.99E+3nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 8.18E+3nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 8.35E+3nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.02E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.04E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.15E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.15E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.16E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.19E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.20E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.22E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.25E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.30E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.30E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.31E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.32E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.33E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.50E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.51E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataKd: 1.60E+4nMAssay Description:Binding affinity to PPAR gamma LBD (203 to 476 residues) (unknown origin) extracted from Escherichia coli BL21 (DE3) cells assessed as dissociation c...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataKd: 1.77E+4nMAssay Description:Binding affinity to PPAR gamma LBD (203 to 476 residues) (unknown origin) extracted from Escherichia coli BL21 (DE3) cells assessed as dissociation c...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 1.95E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Antagonist activity at PPAR gamma LBD (203 to 476 residues) (unknown origin) expressed in HEK293T cells co-transfected with pCMV-GAL4-PPARgamma-LBD-S...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataKd: 7.95E+4nMAssay Description:Binding affinity to PPAR gamma LBD (203 to 476 residues) (unknown origin) extracted from Escherichia coli BL21 (DE3) cells assessed as dissociation c...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Sun Yat-sen University
Curated by ChEMBL
Sun Yat-sen University
Curated by ChEMBL
Affinity DataKd: 1.27E+5nMAssay Description:Binding affinity to PPAR gamma LBD (203 to 476 residues) (unknown origin) extracted from Escherichia coli BL21 (DE3) cells assessed as dissociation c...More data for this Ligand-Target Pair
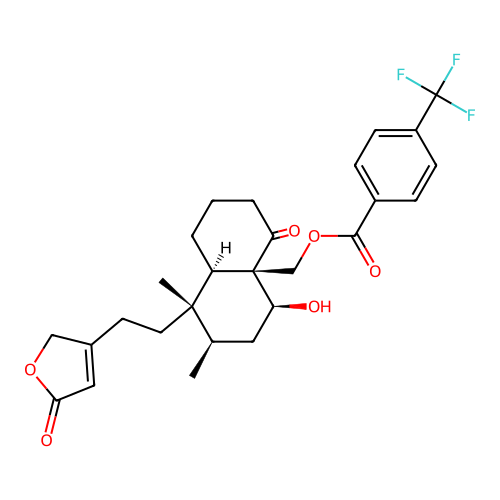
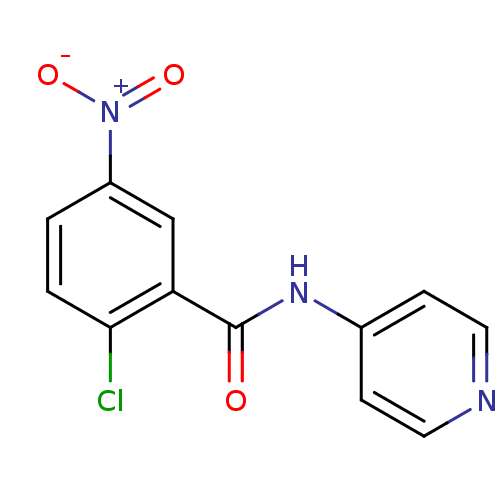
 3D Structure (crystal)
3D Structure (crystal)