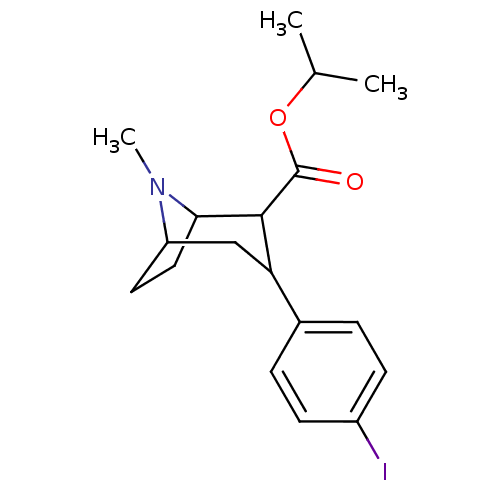
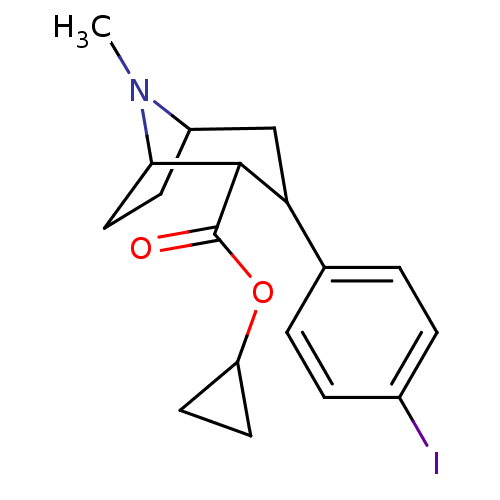
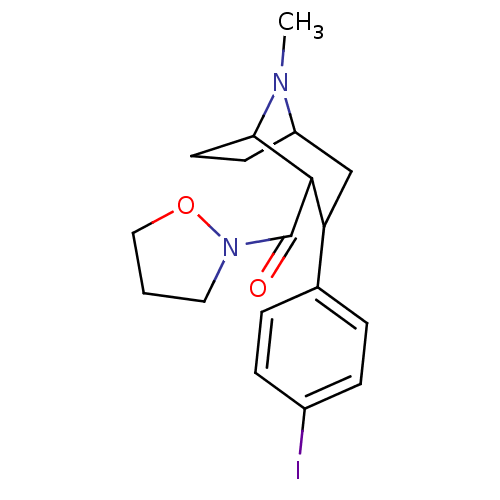
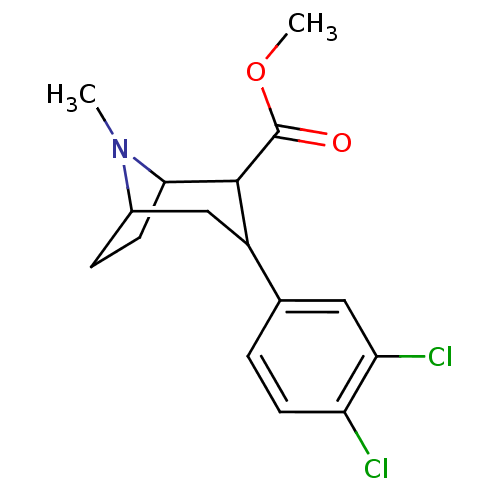
Report error Found 237 Enz. Inhib. hit(s) with all data for entry = 50036367
Affinity DataIC50: 0.370nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 0.430nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 0.610nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 0.75nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 0.790nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 0.810nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 0.850nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 0.960nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 1.5nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 1.5nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataKi: 1.70nMAssay Description:Inhibition of [3H]- 5-HT uptake in Serotonin transporter using serotonin uptake assay in rat midbrain tissueMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of [3H]DA uptake by rat striatal dopamine transporterMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 2.10nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 2.60nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataKi: 2.80nMAssay Description:Inhibition of [3H]DA uptake by rat striatal dopamine transporterMore data for this Ligand-Target Pair
Affinity DataIC50: 2.90nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 3.10nMAssay Description:Binding affinity at the Serotonin transporter in rat midbrain by inhibition of 0.5 nM [3H]-paroxetine bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 3.30nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 3.70nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataKi: 3.70nMAssay Description:Inhibition of [3H]DA uptake by rat striatal dopamine transporterMore data for this Ligand-Target Pair
Affinity DataIC50: 3.90nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 4.20nMAssay Description:Binding affinity at the Serotonin transporter in rat midbrain by inhibition of 0.5 nM [3H]-paroxetine bindingMore data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Inhibition of [3H]- 5-HT uptake in Serotonin transporter using serotonin uptake assay in rat midbrain tissueMore data for this Ligand-Target Pair
Affinity DataKi: 5.30nMAssay Description:Inhibition of [3H]DA uptake by rat striatal dopamine transporterMore data for this Ligand-Target Pair
Affinity DataIC50: 5.5nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataKi: 5.90nMAssay Description:Inhibition of [3H]NE uptake by rat frontal cortex Norepinephrine transporterMore data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Inhibition of [3H]DA uptake by rat striatal dopamine transporterMore data for this Ligand-Target Pair
Affinity DataIC50: 6.5nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 6.60nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataKi: 6.90nMAssay Description:Inhibition of [3H]DA uptake by rat striatal dopamine transporterMore data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Inhibition of [3H]DA uptake by rat striatal dopamine transporterMore data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataKi: 7.5nMAssay Description:Inhibition of [3H]NE uptake by rat frontal cortex Norepinephrine transporterMore data for this Ligand-Target Pair
Affinity DataIC50: 8.20nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataKi: 8.40nMAssay Description:Inhibition of [3H]NE uptake by rat frontal cortex Norepinephrine transporterMore data for this Ligand-Target Pair
Affinity DataKi: 9.10nMAssay Description:Inhibition of [3H]DA uptake by rat striatal dopamine transporterMore data for this Ligand-Target Pair
Affinity DataIC50: 9.40nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Binding affinity at the Serotonin transporter in rat midbrain by inhibition of 0.5 nM [3H]-paroxetine bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Binding affinity at the Dopamine transporter in rat striata by inhibition of 0.5 nM [3H]WIN-35428 bindingMore data for this Ligand-Target Pair