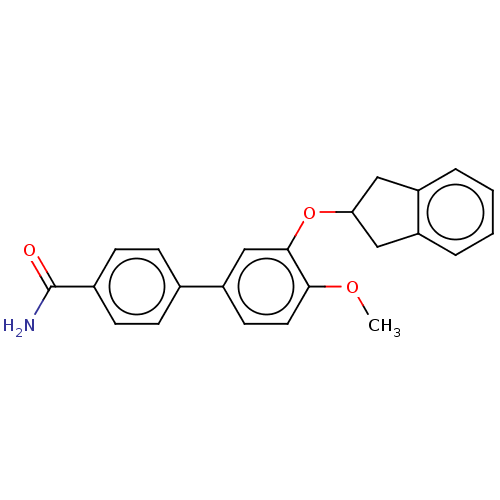
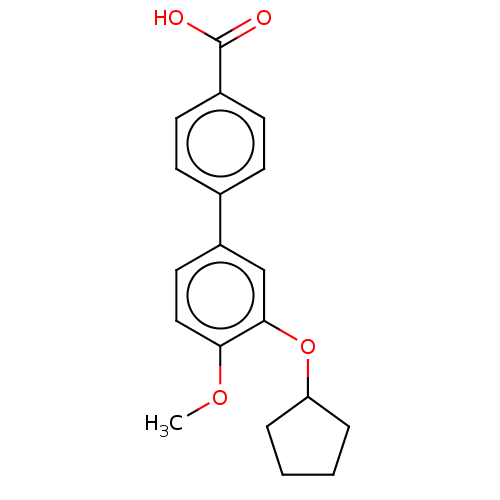
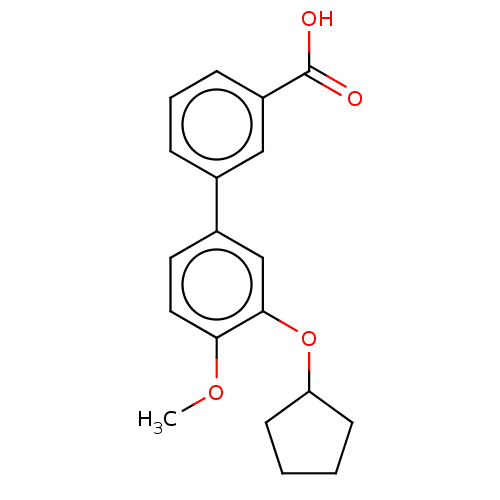
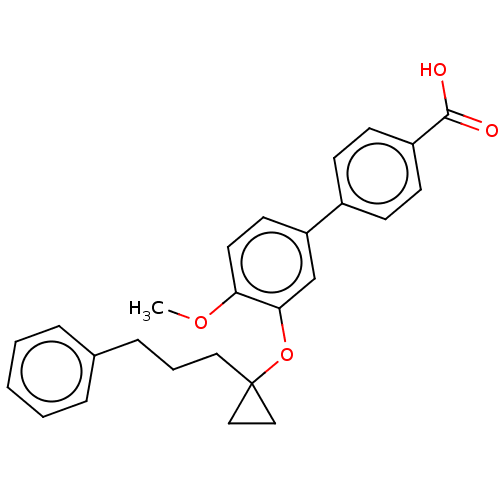
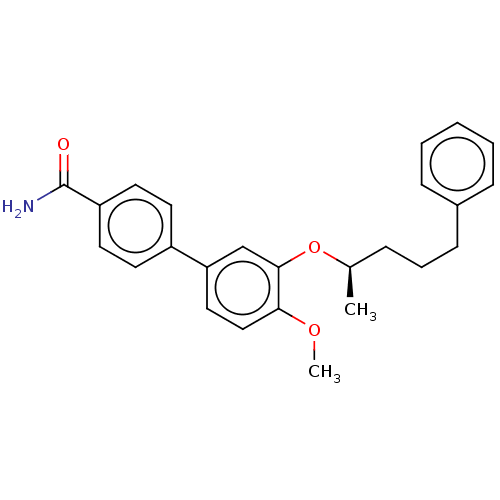
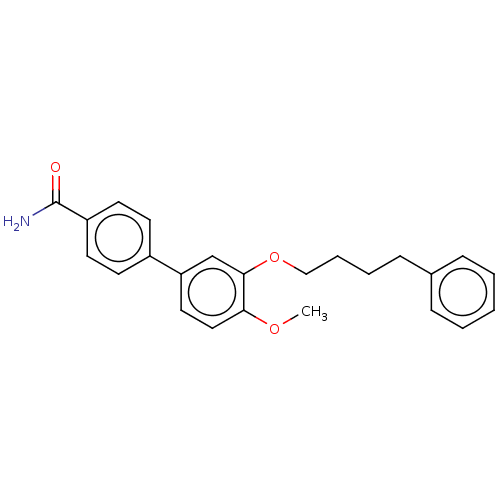
Report error Found 29 Enz. Inhib. hit(s) with all data for entry = 50003427
Affinity DataIC50: 3nMAssay Description:Displacement of [3H]rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Displacement of [3H]rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Displacement of [3H]rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Displacement of [3H]rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:Displacement of [3H]rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Displacement of [3H]rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Displacement of [3H]rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 81nMAssay Description:Inhibition of Phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 110nMAssay Description:Inhibition of Phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 111nMAssay Description:Displacement of [3H]rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 129nMAssay Description:Displacement of [3H]rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:Inhibition of Phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:Displacement of [3H]rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:Inhibition of Phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 140nMAssay Description:Inhibition of Phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 160nMAssay Description:Inhibition of Phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Inhibition of Phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 323nMAssay Description:Displacement of [3H]rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 410nMAssay Description:Inhibition of Phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 520nMAssay Description:Inhibition of Phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 653nMAssay Description:Displacement of [3H]rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 670nMAssay Description:Inhibition of Phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 1.42E+3nMAssay Description:Inhibition of Phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 1.64E+3nMAssay Description:Inhibition of Phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 1.68E+3nMAssay Description:Displacement of [3H]rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+3nMAssay Description:Displacement of [3H]rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 1.94E+3nMAssay Description:Inhibition of Phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 3.45E+3nMAssay Description:Inhibition of Phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 2.03E+4nMAssay Description:Inhibition of Phosphodiesterase 4More data for this Ligand-Target Pair
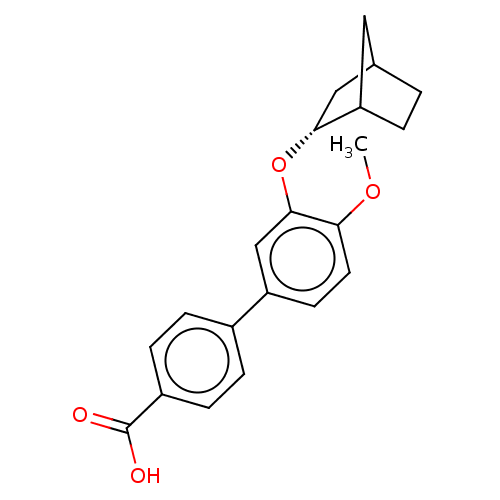
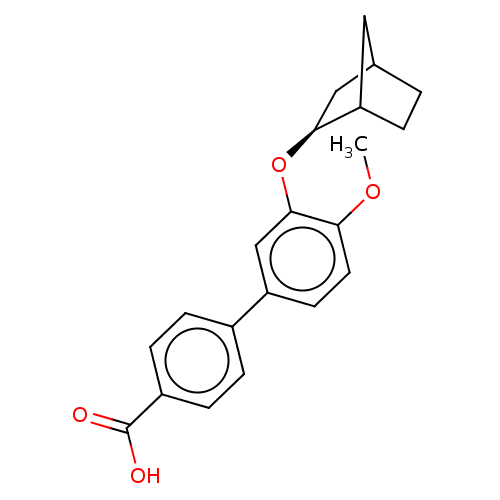
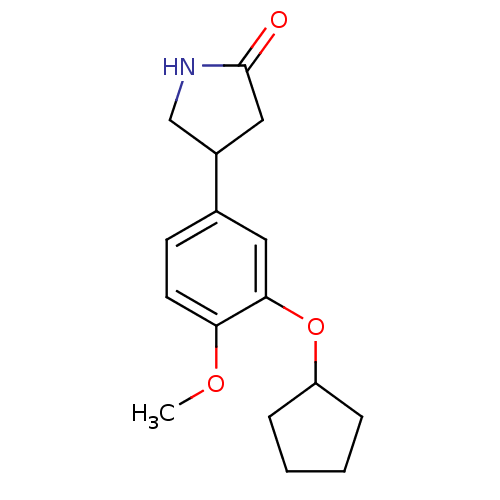
 3D Structure (crystal)
3D Structure (crystal)