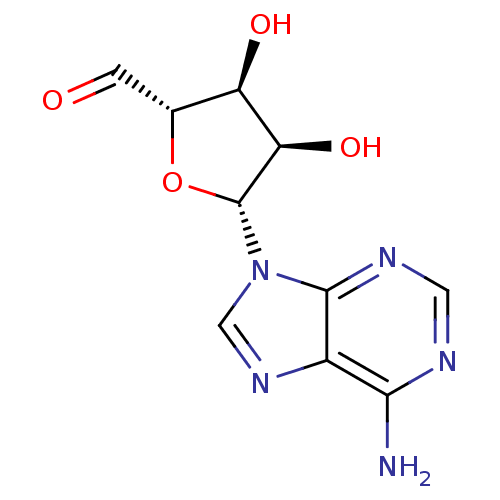
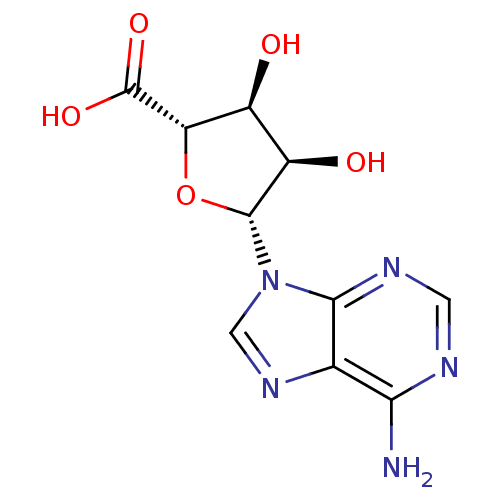
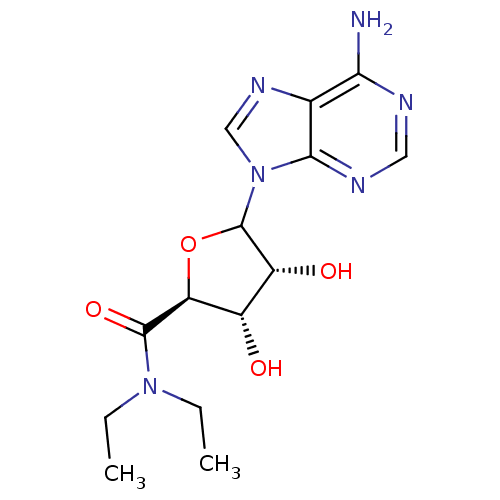
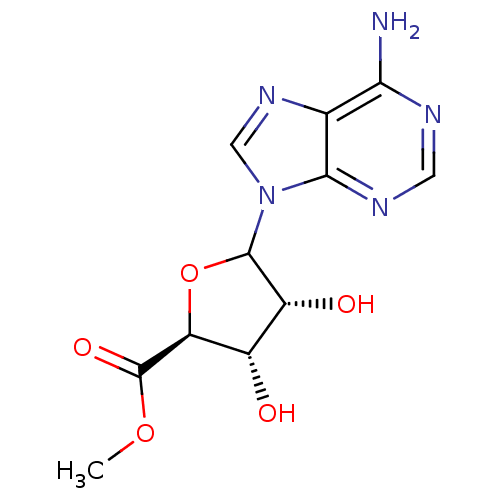
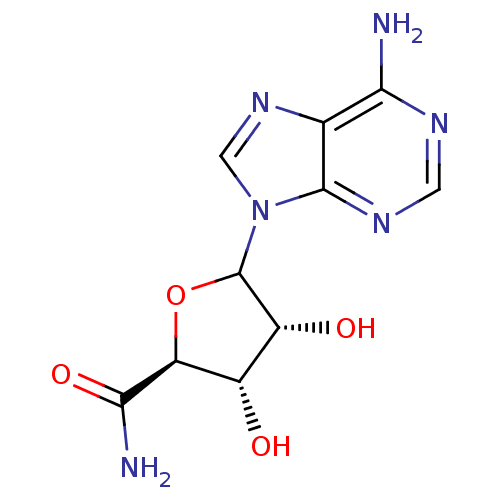
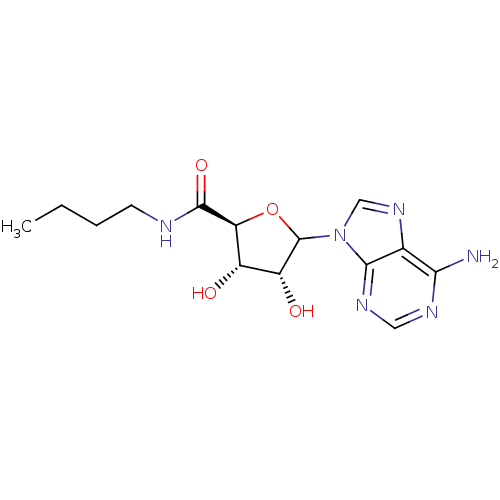
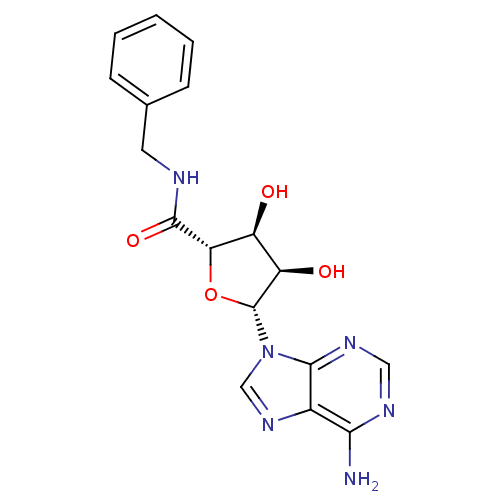
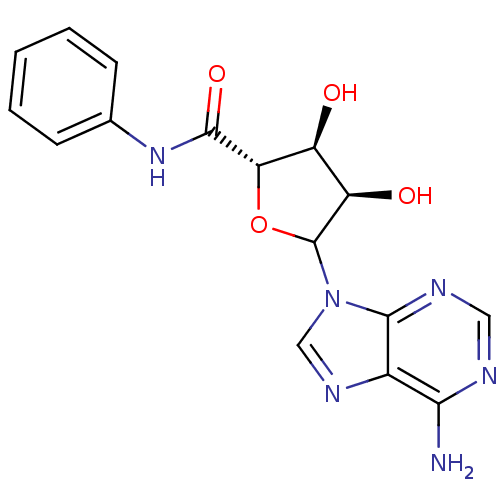
Report error Found 14 Enz. Inhib. hit(s) with all data for entry = 50036458
Affinity DataKi: 39nMAssay Description:Inhibition of S-adenosyl-L-homocysteine hydrolase, inactivation rate constant.More data for this Ligand-Target Pair
Affinity DataKi: 320nMAssay Description:Inhibition of S-adenosyl-L-homocysteine hydrolase, inactivation rate constant.More data for this Ligand-Target Pair
Affinity DataKi: 360nMAssay Description:Inhibition of S-adenosyl-L-homocysteine hydrolase, inactivation rate constant.More data for this Ligand-Target Pair
Affinity DataKi: 410nMAssay Description:Inhibition of S-adenosyl-L-homocysteine hydrolase, inactivation rate constant.More data for this Ligand-Target Pair
Affinity DataKi: 570nMAssay Description:Inhibition of S-adenosyl-L-homocysteine hydrolase, inactivation rate constant.More data for this Ligand-Target Pair
Affinity DataKi: 570nMAssay Description:Inhibition of S-adenosyl-L-homocysteine hydrolase, inactivation rate constant.More data for this Ligand-Target Pair
Affinity DataKi: 710nMAssay Description:Inhibition of S-adenosyl-L-homocysteine hydrolase, inactivation rate constant.More data for this Ligand-Target Pair
Affinity DataKi: 770nMAssay Description:Inhibition of S-adenosyl-L-homocysteine hydrolase, inactivation rate constant.More data for this Ligand-Target Pair
Affinity DataKi: 1.70E+3nMAssay Description:Inhibition of S-adenosyl-L-homocysteine hydrolase, inactivation rate constant.More data for this Ligand-Target Pair
Affinity DataKi: 1.93E+3nMAssay Description:Inhibition of S-adenosyl-L-homocysteine hydrolase, inactivation rate constant.More data for this Ligand-Target Pair
Affinity DataKi: 2.60E+3nMAssay Description:Inhibition of S-adenosyl-L-homocysteine hydrolase, inactivation rate constant.More data for this Ligand-Target Pair
Affinity DataKi: 1.98E+4nMAssay Description:Inhibition of S-adenosyl-L-homocysteine hydrolase, inactivation rate constant.More data for this Ligand-Target Pair
Affinity DataKi: 4.34E+4nMAssay Description:Inhibition of S-adenosyl-L-homocysteine hydrolase, inactivation rate constant.More data for this Ligand-Target Pair
Affinity DataKi: 4.85E+4nMAssay Description:Inhibition of S-adenosyl-L-homocysteine hydrolase, inactivation rate constant.More data for this Ligand-Target Pair