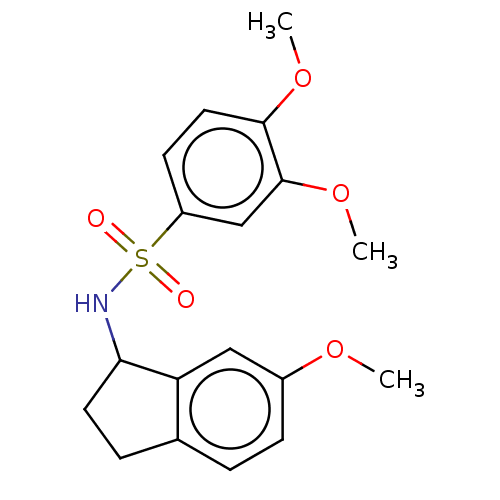
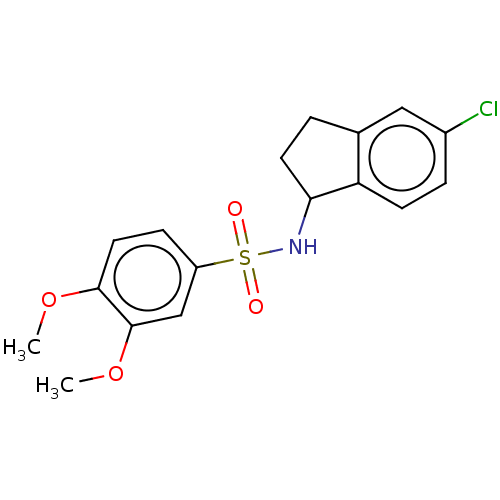
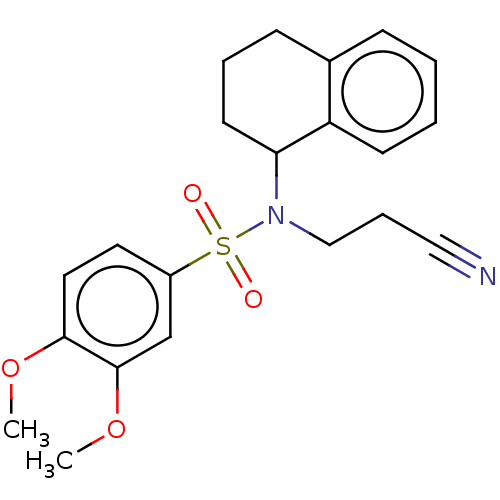
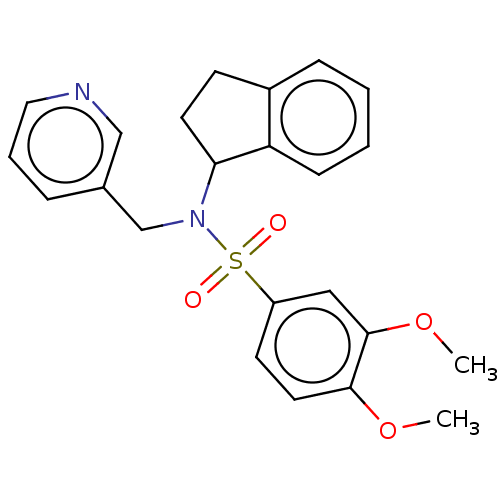
Report error Found 44 Enz. Inhib. hit(s) with all data for entry = 50008249
Affinity DataIC50: 20nMAssay Description:Relative inhibition of phosphodiesterase 4 activity and Rolipram binding to PDE4More data for this Ligand-Target Pair
Affinity DataIC50: 500nMAssay Description:Inhibition of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 500nMAssay Description:Inhibition of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 600nMAssay Description:Inhibition of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+3nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+3nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:Binding activity towards Rolipram binding site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 3.50E+3nMAssay Description:Inhibition of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+3nMAssay Description:Inhibition of Rolipram binding to phosphodiesterase 4 at 20 uMMore data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+3nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 4.30E+3nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 4.30E+3nMAssay Description:Binding activity towards Rolipram binding site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 4.50E+3nMAssay Description:Binding activity towards Rolipram binding site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 5.20E+3nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 5.30E+3nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+3nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+3nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 6.40E+3nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 7.80E+3nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 9.00E+3nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 9.00E+3nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Binding activity towards Rolipram binding site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+4nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMAssay Description:Binding activity towards Rolipram binding site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 1.76E+4nMAssay Description:Inhibition of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+4nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+4nMAssay Description:Binding activity towards Rolipram binding site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+4nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 3.50E+4nMAssay Description:Binding activity towards Rolipram binding site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Binding activity towards Rolipram binding site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Binding activity towards Rolipram binding site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:In vitro binding activity towards catalytic site of phosphodiesterase 4More data for this Ligand-Target Pair
Affinity DataIC50: 1.12E+5nMAssay Description:Binding activity towards Rolipram binding site of phosphodiesterase 4More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1A/1B/1C(Human)
Chiroscience
Curated by ChEMBL
Chiroscience
Curated by ChEMBL
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of phosphodiesterase 1 at 20 uMMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of phosphodiesterase 5 at 20 uMMore data for this Ligand-Target Pair
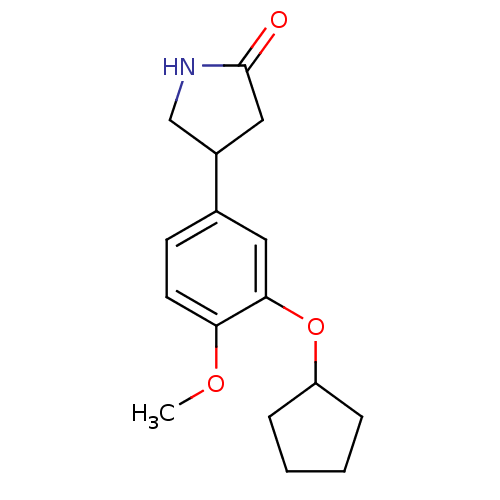
 3D Structure (crystal)
3D Structure (crystal)