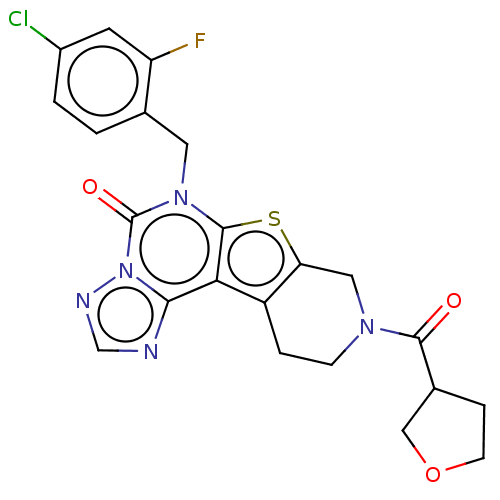
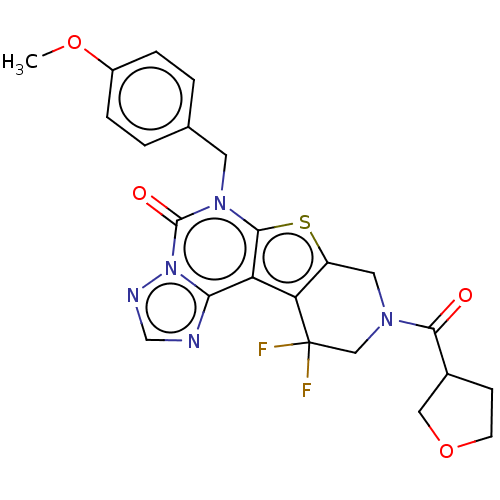
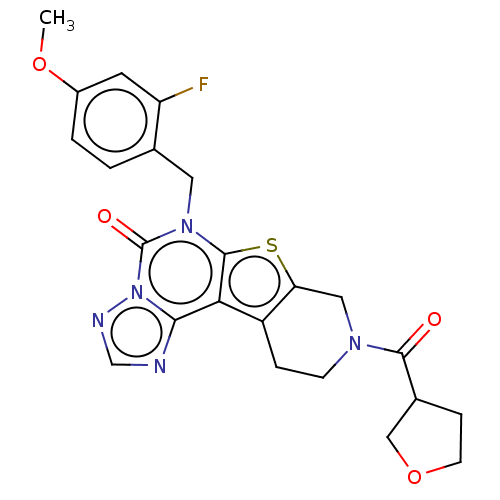
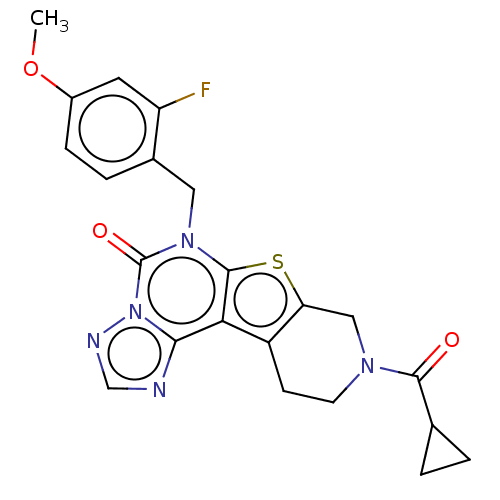
Report error Found 304 Enz. Inhib. hit(s) with all data for entry = 817
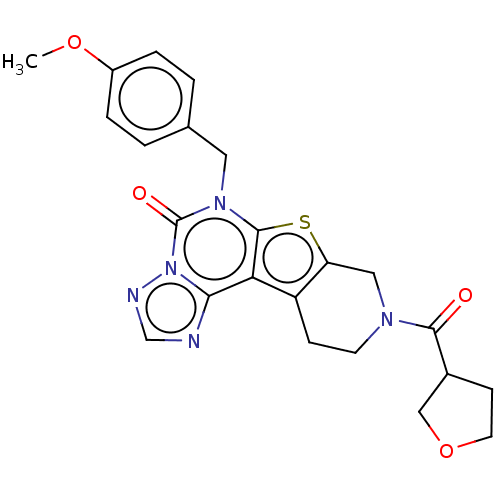
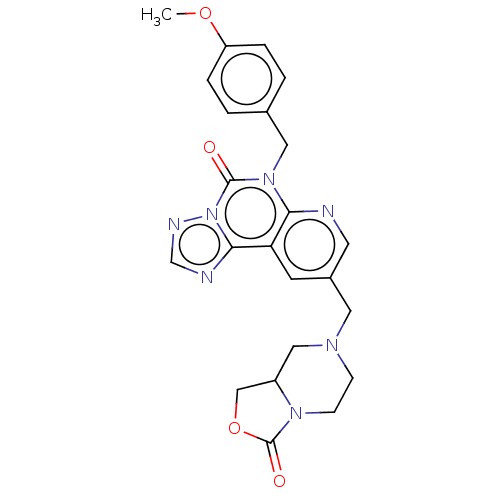
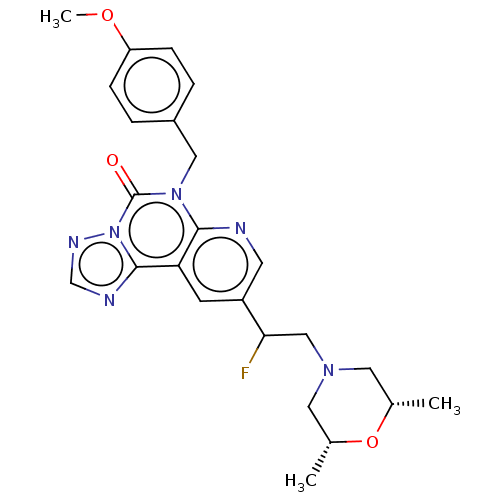
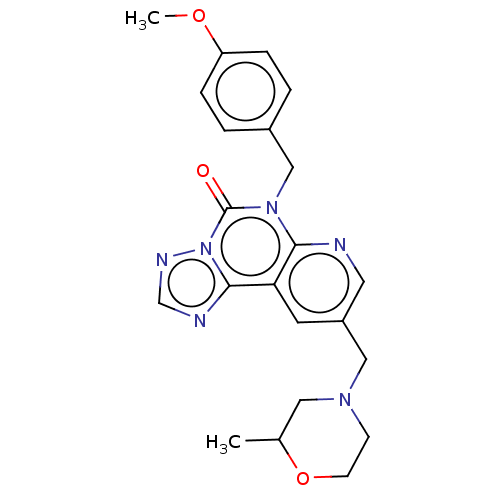
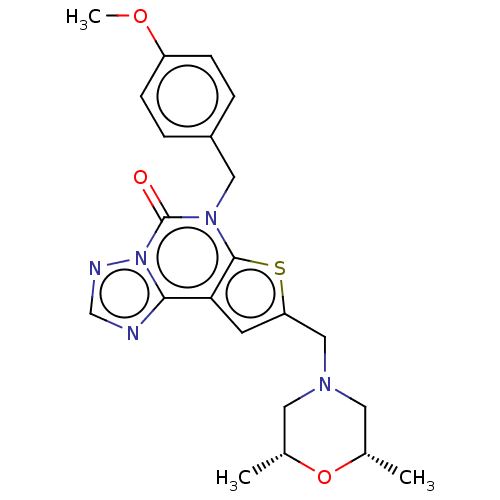
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
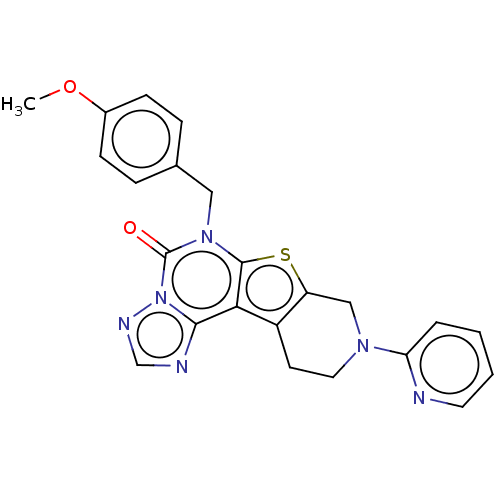
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
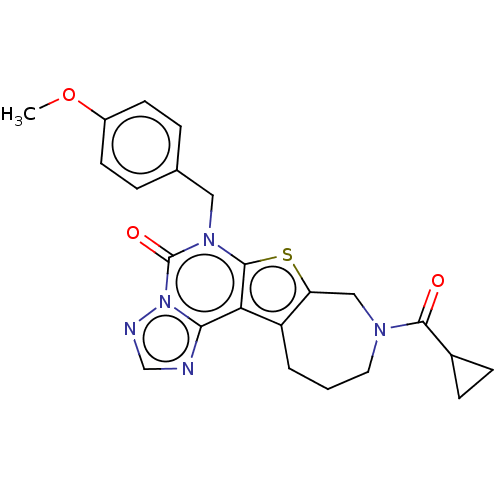
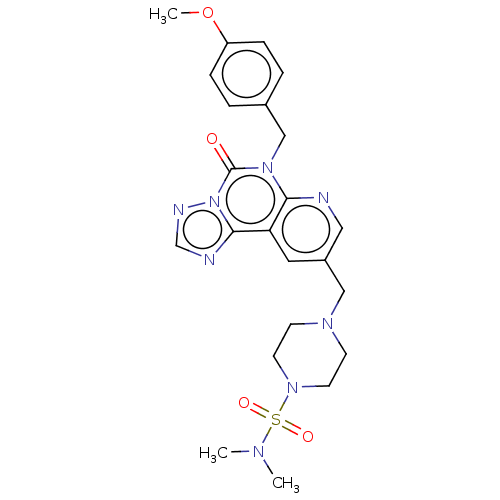
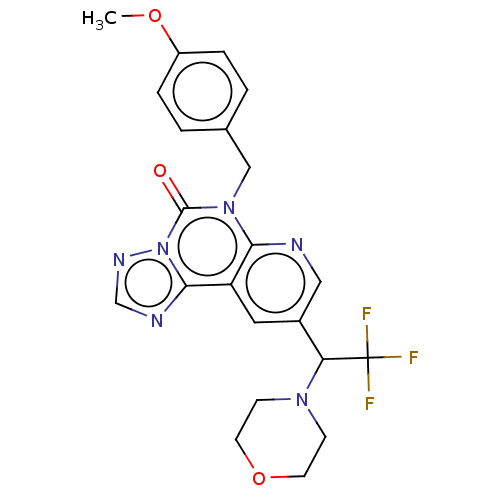
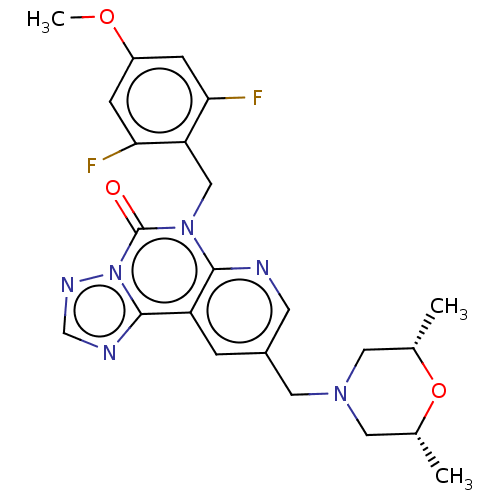
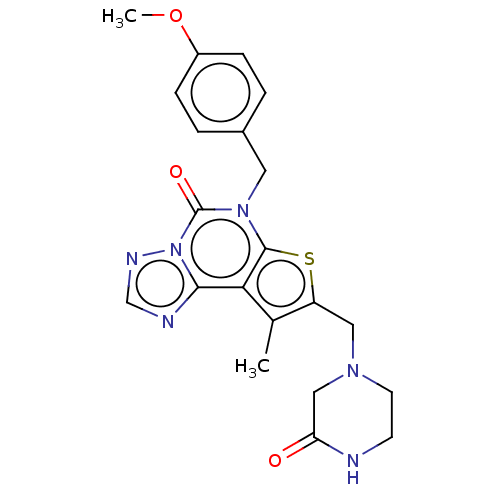
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
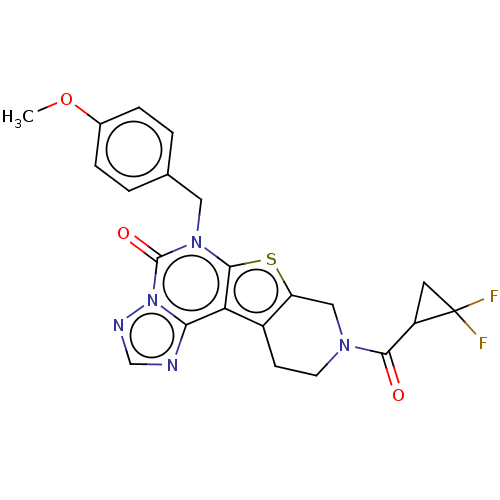
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
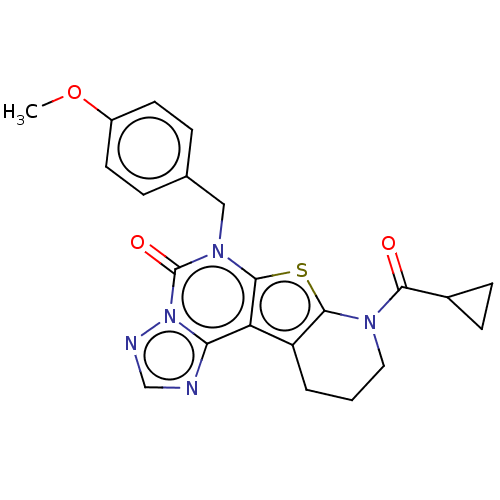
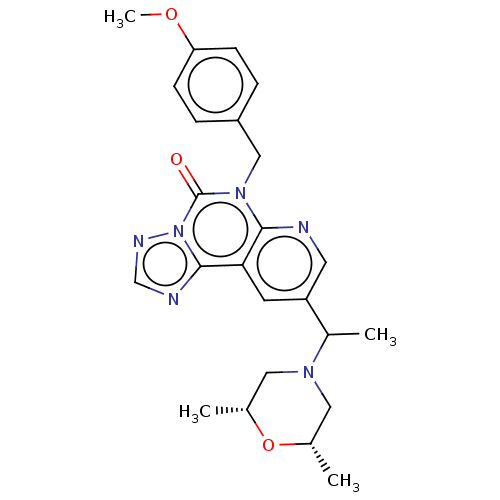
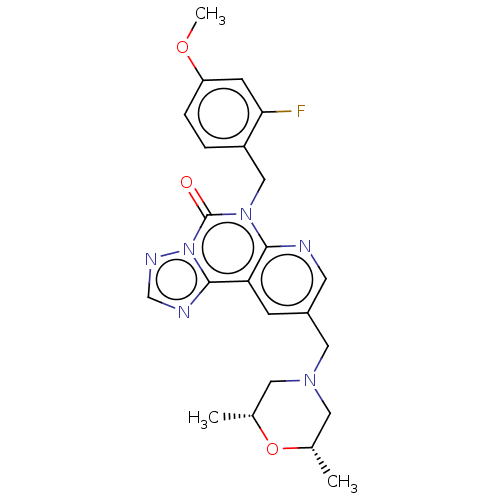
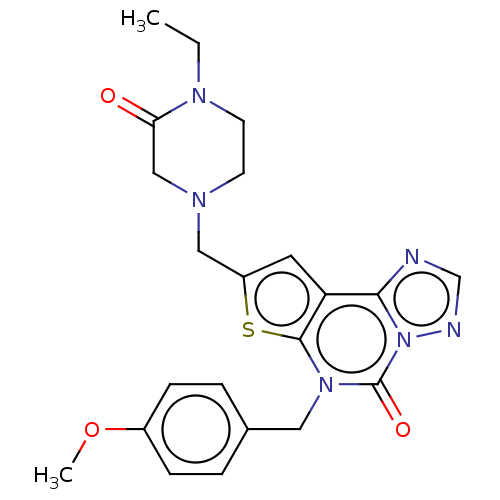
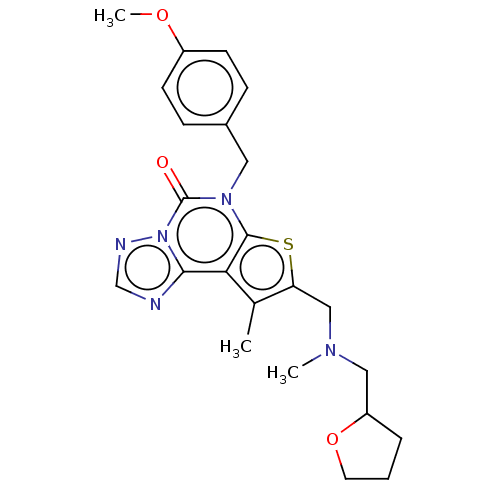
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
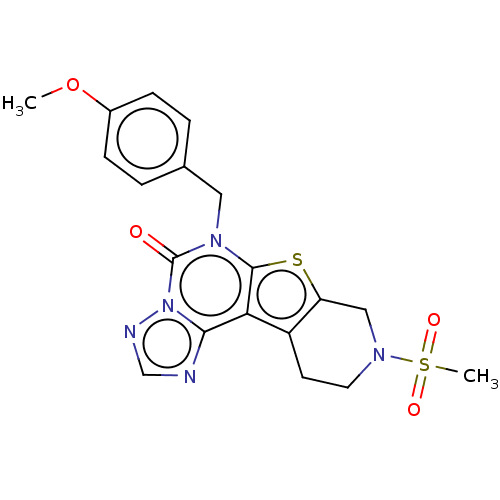
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
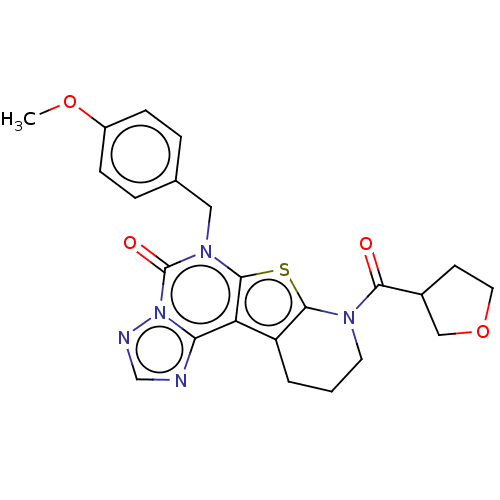
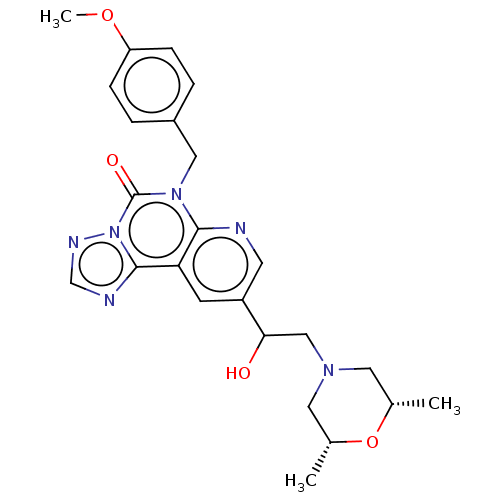
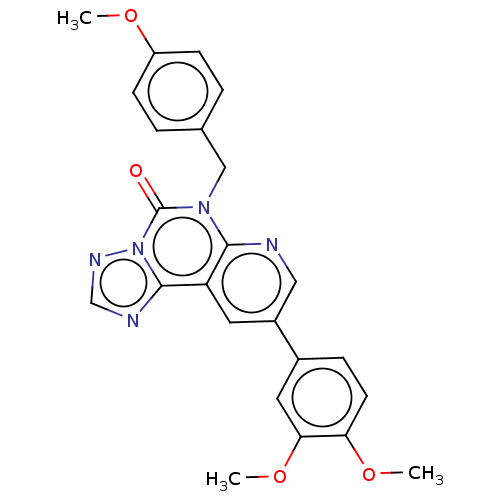
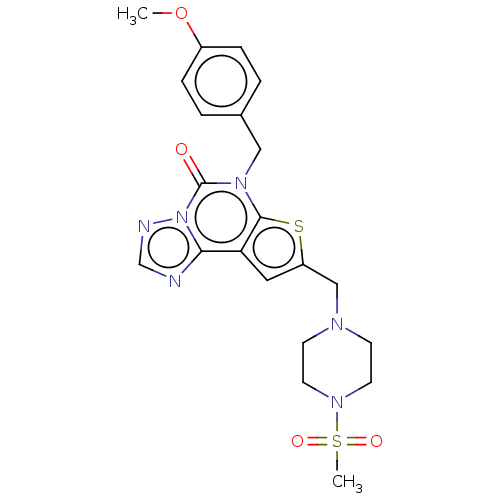
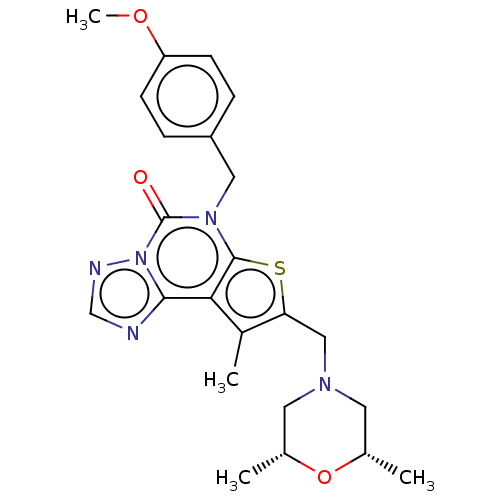
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
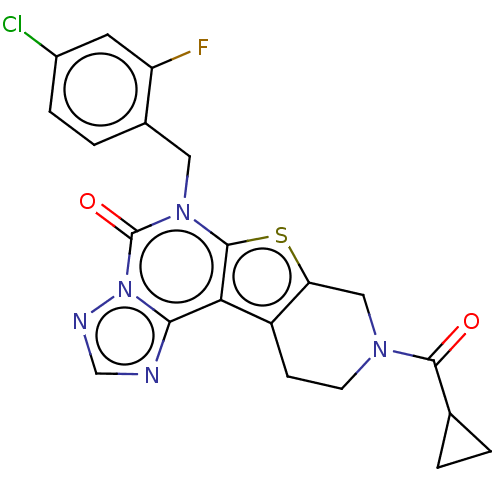
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair
TargetDual specificity calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B(Human)
Dart Neuroscience (Cayman)
US Patent
Dart Neuroscience (Cayman)
US Patent
Affinity DataIC50: 100nMT: 2°CAssay Description:PDE1B inhibition was determined by an IMAP TR-FRET assay. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, Calmodulin, cAMP or c...More data for this Ligand-Target Pair