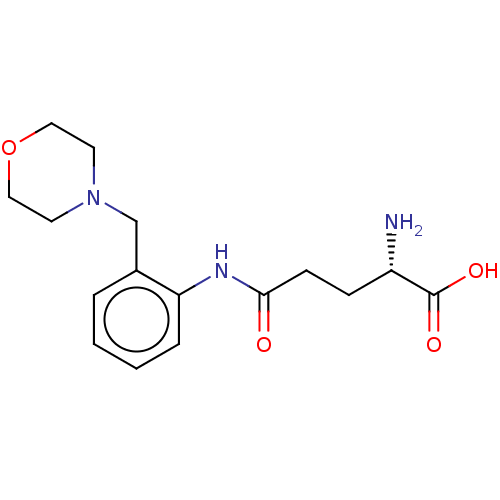
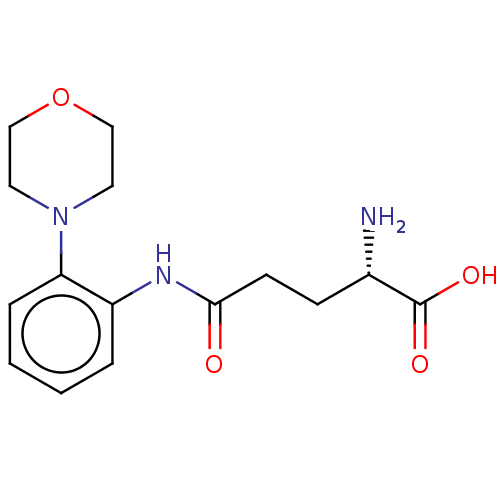
Report error Found 8 Enz. Inhib. hit(s) with all data for entry = 1455
Affinity DataIC50: 3.12E+5nMAssay Description:The glutamine uptake assays in HEK293 cells were carried out in 96 well plates (CulturPlate-96, Perkin Elmer). Cells were plated at a density of 35,0...More data for this Ligand-Target Pair
Affinity DataIC50: 4.36E+5nMAssay Description:The glutamine uptake assays in HEK293 cells were carried out in 96 well plates (CulturPlate-96, Perkin Elmer). Cells were plated at a density of 35,0...More data for this Ligand-Target Pair
Affinity DataIC50: 6.64E+5nMAssay Description:The glutamine uptake assays in HEK293 cells were carried out in 96 well plates (CulturPlate-96, Perkin Elmer). Cells were plated at a density of 35,0...More data for this Ligand-Target Pair
Affinity DataIC50: 6.77E+5nMAssay Description:The glutamine uptake assays in HEK293 cells were carried out in 96 well plates (CulturPlate-96, Perkin Elmer). Cells were plated at a density of 35,0...More data for this Ligand-Target Pair
Affinity DataIC50: 6.97E+5nMAssay Description:The glutamine uptake assays in HEK293 cells were carried out in 96 well plates (CulturPlate-96, Perkin Elmer). Cells were plated at a density of 35,0...More data for this Ligand-Target Pair
Affinity DataIC50: 7.76E+5nMAssay Description:The glutamine uptake assays in HEK293 cells were carried out in 96 well plates (CulturPlate-96, Perkin Elmer). Cells were plated at a density of 35,0...More data for this Ligand-Target Pair
Affinity DataIC50: 8.32E+5nMAssay Description:The glutamine uptake assays in HEK293 cells were carried out in 96 well plates (CulturPlate-96, Perkin Elmer). Cells were plated at a density of 35,0...More data for this Ligand-Target Pair
Affinity DataIC50: 9.54E+5nMAssay Description:The glutamine uptake assays in HEK293 cells were carried out in 96 well plates (CulturPlate-96, Perkin Elmer). Cells were plated at a density of 35,0...More data for this Ligand-Target Pair