Report error Found 4 Enz. Inhib. hit(s) with all data for entry = 11263
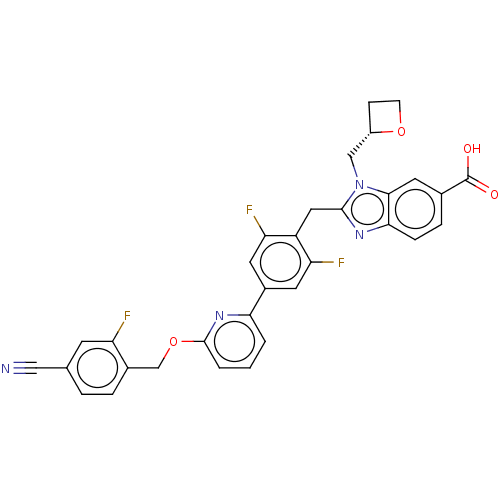
Affinity DataIC50: 5.41E+3nMAssay Description:PDE10A1 enzyme activities are measured with a yittrium silicate based scintillation proximity assay (SPA) that detects radioactive nucleotide monopho...More data for this Ligand-Target Pair
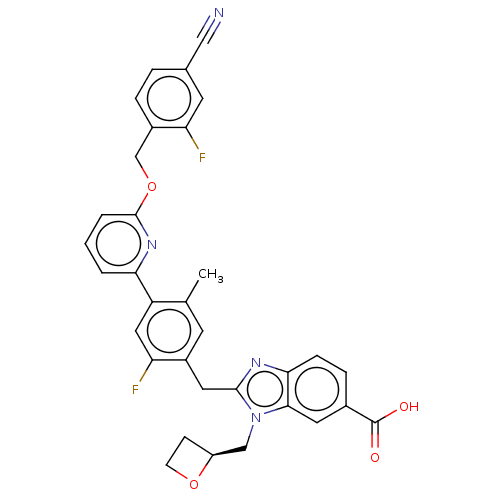
Affinity DataIC50: 7.43E+3nMAssay Description:PDE10A1 enzyme activities are measured with a yittrium silicate based scintillation proximity assay (SPA) that detects radioactive nucleotide monopho...More data for this Ligand-Target Pair
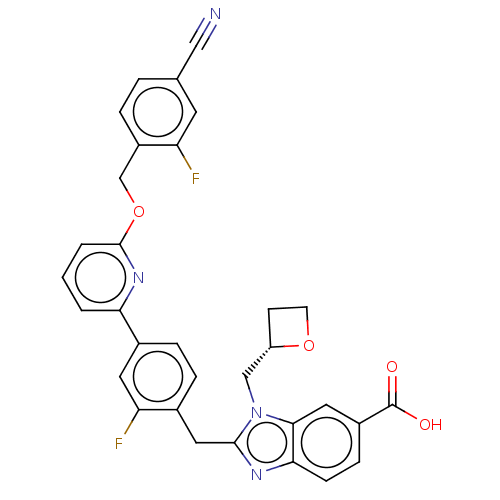
Affinity DataIC50: 1.00E+4nMAssay Description:PDE10A1 enzyme activities are measured with a yittrium silicate based scintillation proximity assay (SPA) that detects radioactive nucleotide monopho...More data for this Ligand-Target Pair
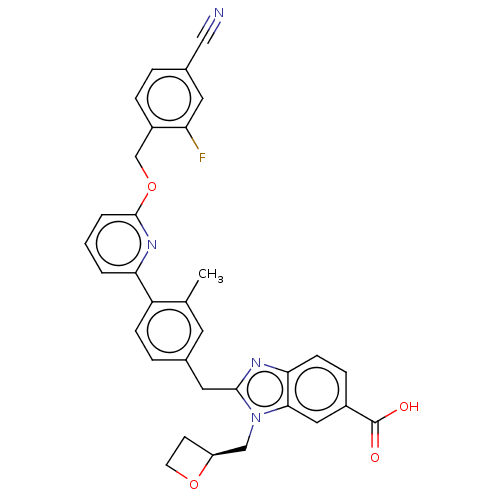
Affinity DataIC50: 1.00E+4nMAssay Description:PDE10A1 enzyme activities are measured with a yittrium silicate based scintillation proximity assay (SPA) that detects radioactive nucleotide monopho...More data for this Ligand-Target Pair