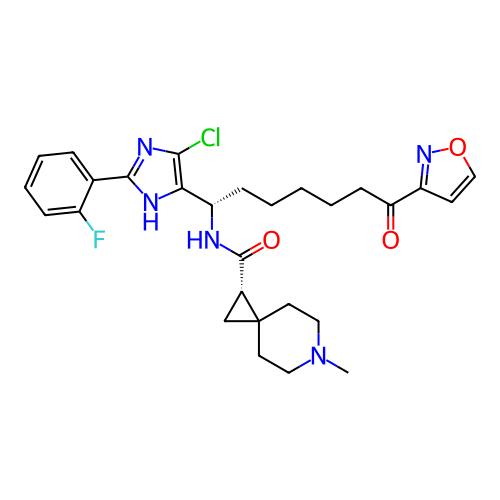
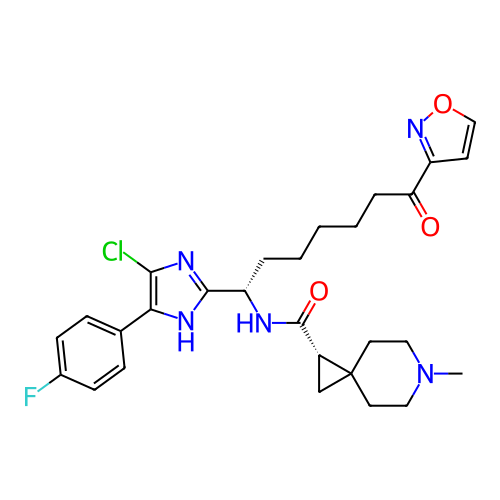
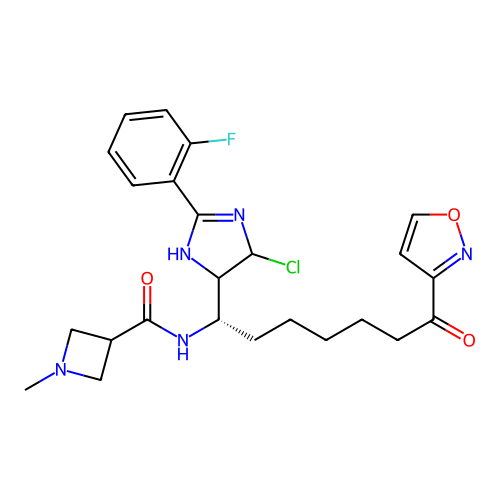
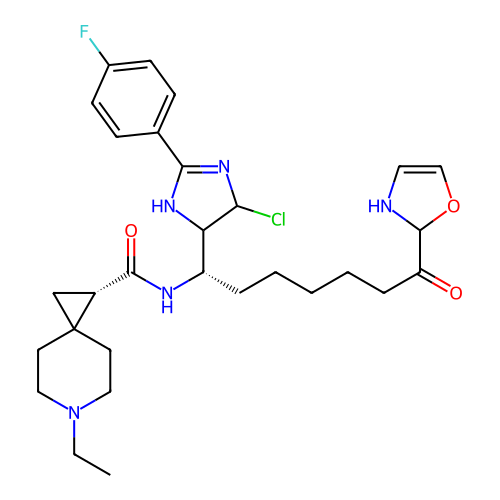
Report error Found 967 Enz. Inhib. hit(s) with all data for entry = 13005
Affinity DataIC50: 0.0350nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0460nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0460nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0490nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0660nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0680nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0830nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0870nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0930nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.100nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.100nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.110nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.110nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.120nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.130nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.130nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.130nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.140nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.140nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.140nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.140nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.140nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.150nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.160nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.160nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.160nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.170nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.180nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.180nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.180nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.180nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.180nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.180nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.180nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.180nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.190nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.190nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.190nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.220nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.230nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.230nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.240nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.240nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.240nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.240nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.25nMAssay Description:The HDAC 1, 2 assays employed buffer A, which contained 20 mM HEPES, pH 8.0, 1 mM MgCl2, 137 mM NaCl, 2.7 mM KCl, 0.05% BSA. The HDAC3/SMRT assay emp...More data for this Ligand-Target Pair