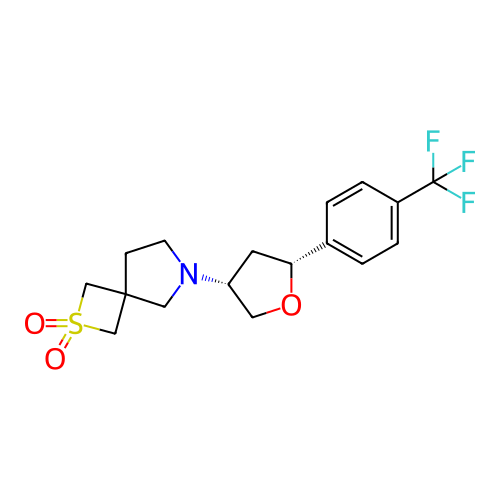
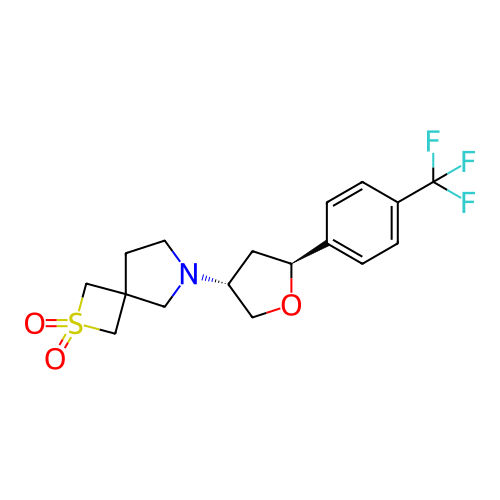
Report error Found 253 Enz. Inhib. hit(s) with all data for entry = 12498
Affinity DataKi: 2.80nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 6.5nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 6.70nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 6.90nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 9.60nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 9.90nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:DHCR7: The radioligand binding assay was prepared by adding assay buffer diluted hEBP-DHCR7 membrane at 66.7 μg/ml×150 μl/well into the 96-well compo...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:DHCR7: The radioligand binding assay was prepared by adding assay buffer diluted hEBP-DHCR7 membrane at 66.7 μg/ml×150 μl/well into the 96-well compo...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 14.8nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:DHCR7: The radioligand binding assay was prepared by adding assay buffer diluted hEBP-DHCR7 membrane at 66.7 μg/ml×150 μl/well into the 96-well compo...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 15.8nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:DHCR7: The radioligand binding assay was prepared by adding assay buffer diluted hEBP-DHCR7 membrane at 66.7 μg/ml×150 μl/well into the 96-well compo...More data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:DHCR7: The radioligand binding assay was prepared by adding assay buffer diluted hEBP-DHCR7 membrane at 66.7 μg/ml×150 μl/well into the 96-well compo...More data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 17.5nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:DHCR7: The radioligand binding assay was prepared by adding assay buffer diluted hEBP-DHCR7 membrane at 66.7 μg/ml×150 μl/well into the 96-well compo...More data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 18.7nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 19nMAssay Description:DHCR7: The radioligand binding assay was prepared by adding assay buffer diluted hEBP-DHCR7 membrane at 66.7 μg/ml×150 μl/well into the 96-well compo...More data for this Ligand-Target Pair
Affinity DataKi: 19nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:DHCR7: The radioligand binding assay was prepared by adding assay buffer diluted hEBP-DHCR7 membrane at 66.7 μg/ml×150 μl/well into the 96-well compo...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:DHCR7: The radioligand binding assay was prepared by adding assay buffer diluted hEBP-DHCR7 membrane at 66.7 μg/ml×150 μl/well into the 96-well compo...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:SPA: Determination of equilibrium dissociation constant Kd of radioligand: Membrane prepared as described above was pre-incubated with PVT-WGA SPA be...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:DHCR7: The radioligand binding assay was prepared by adding assay buffer diluted hEBP-DHCR7 membrane at 66.7 μg/ml×150 μl/well into the 96-well compo...More data for this Ligand-Target Pair