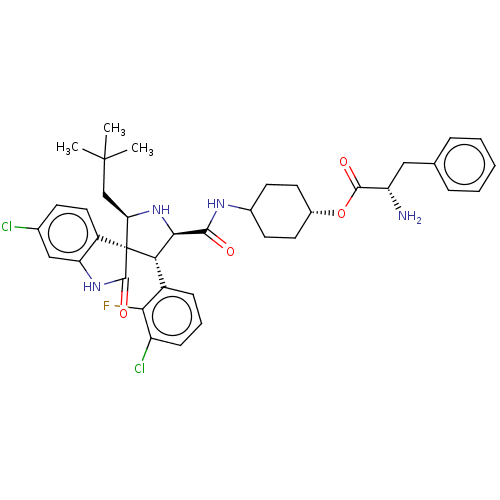
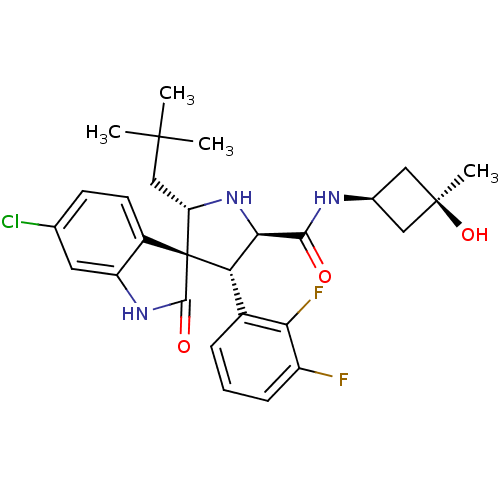
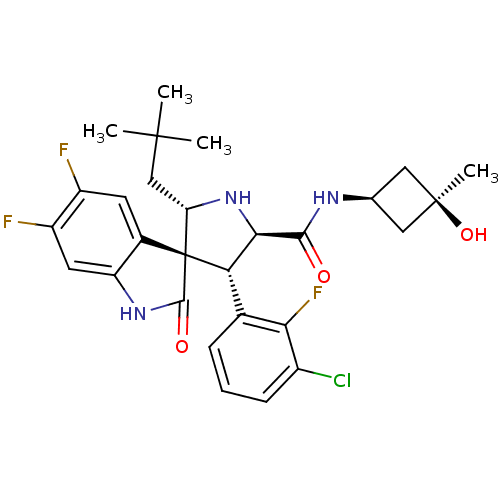
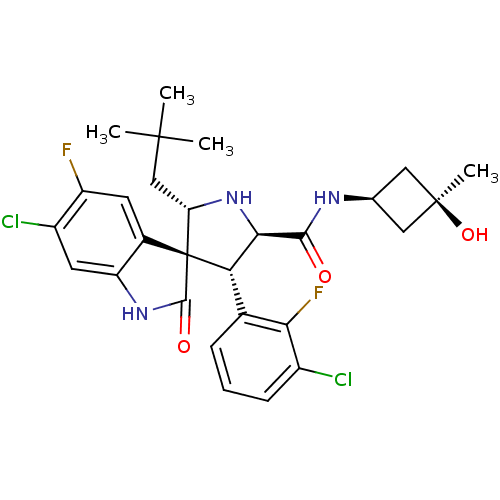
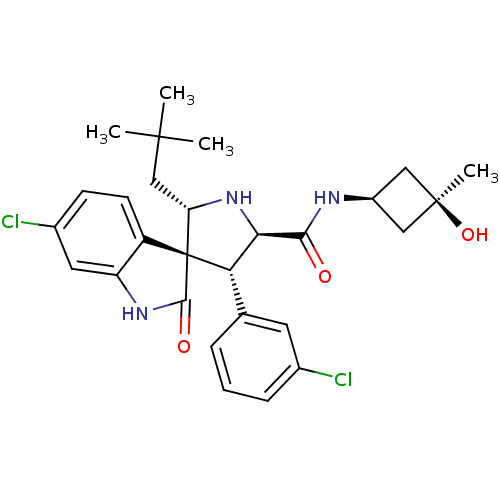
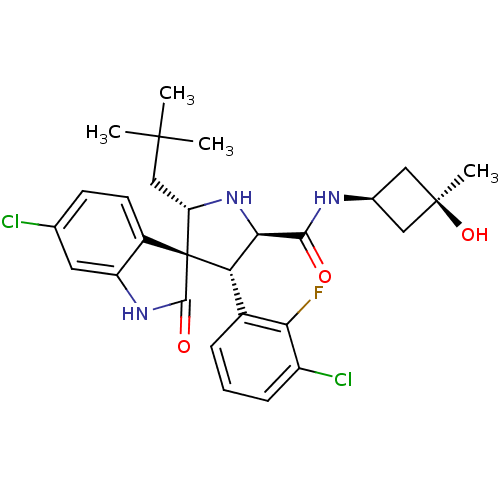
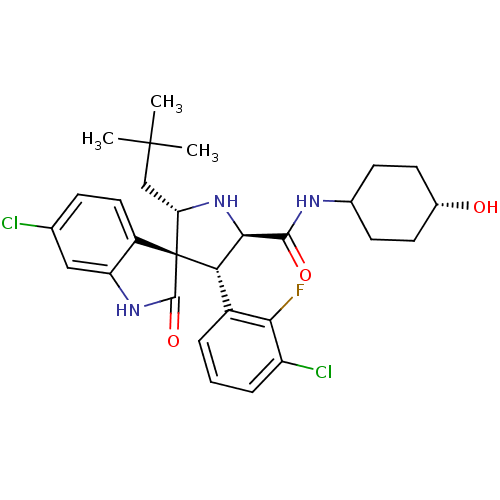
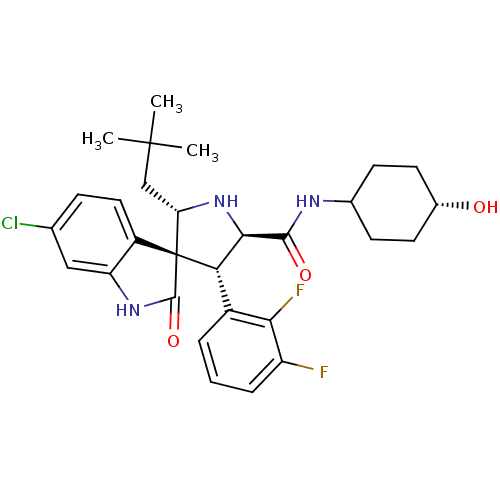
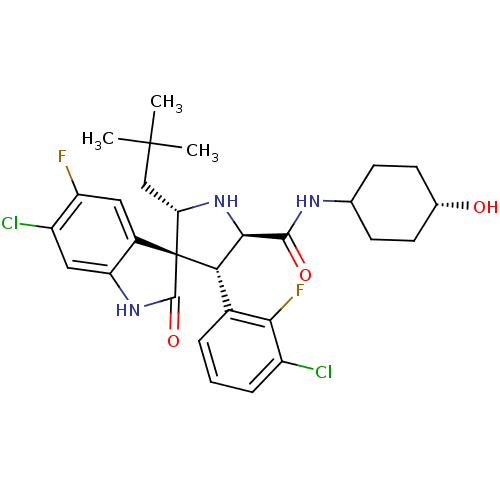
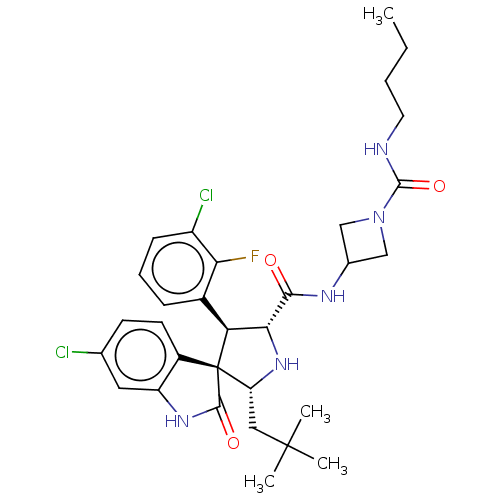
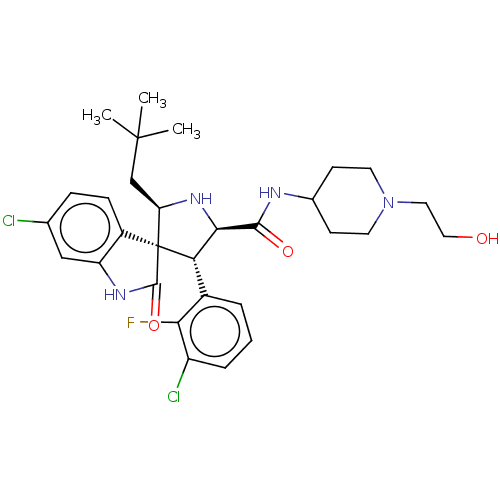
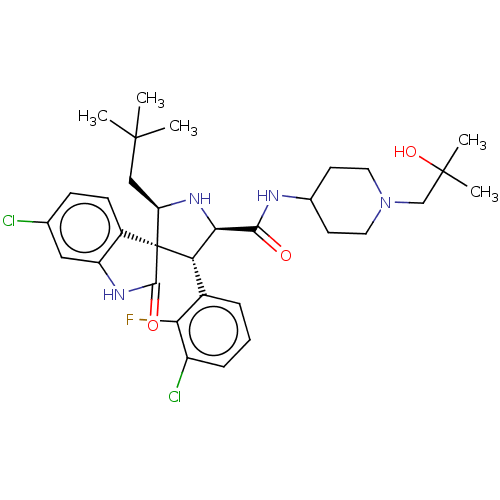
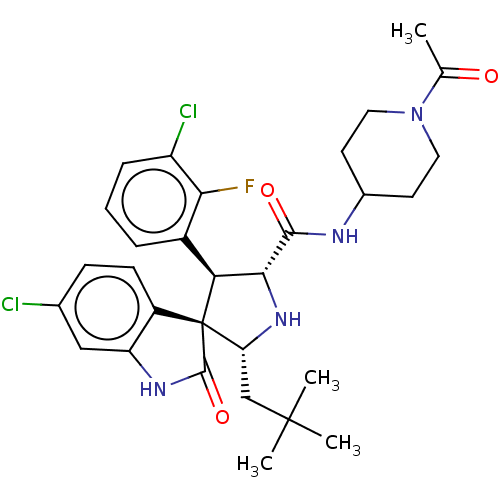
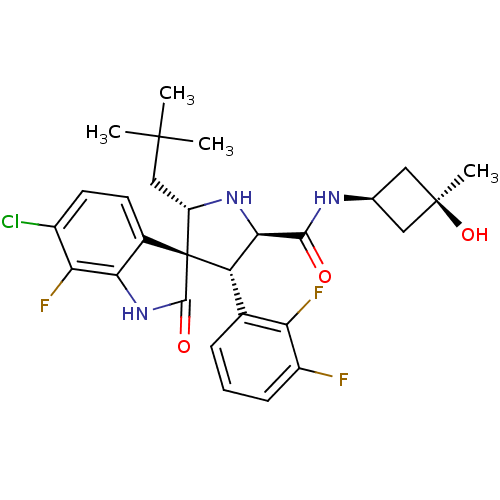
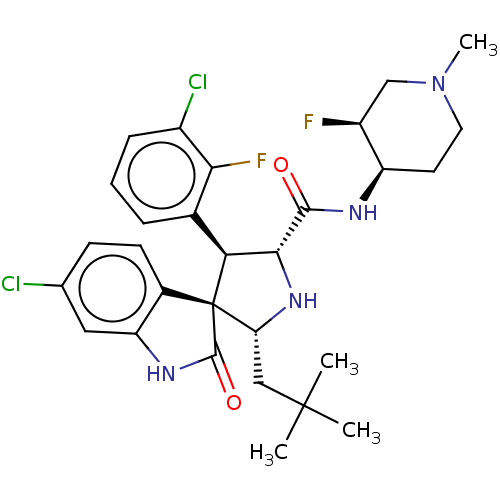
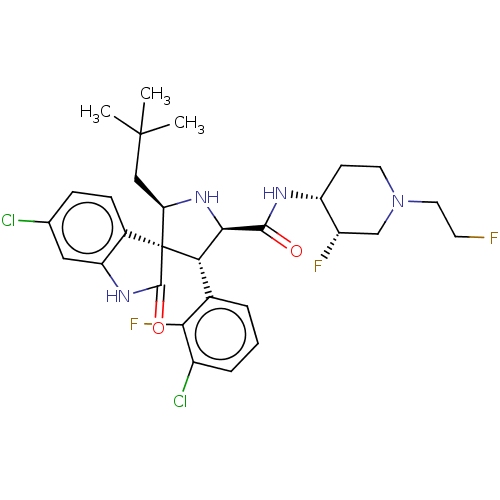
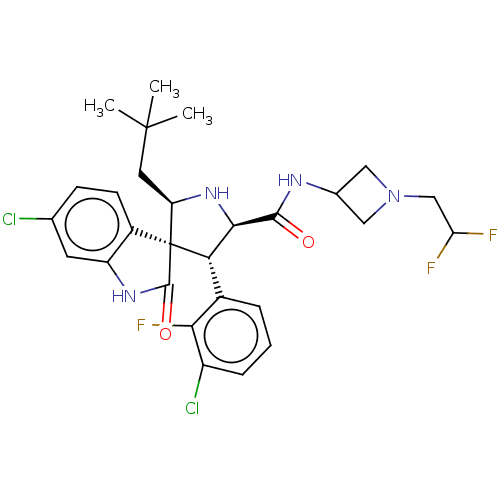
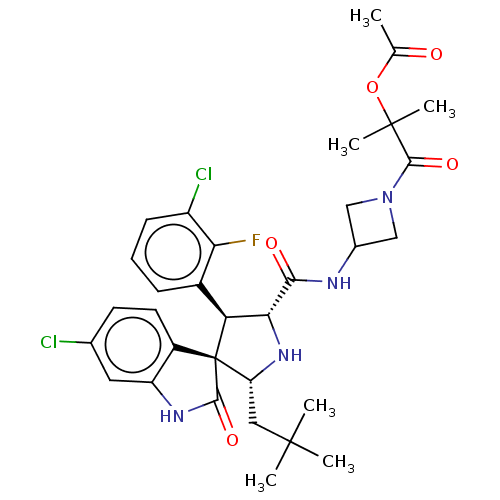
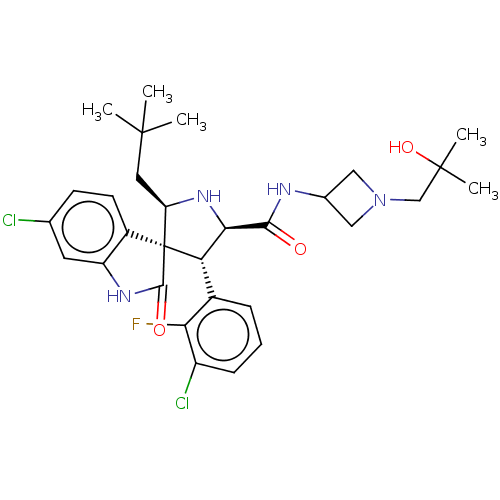
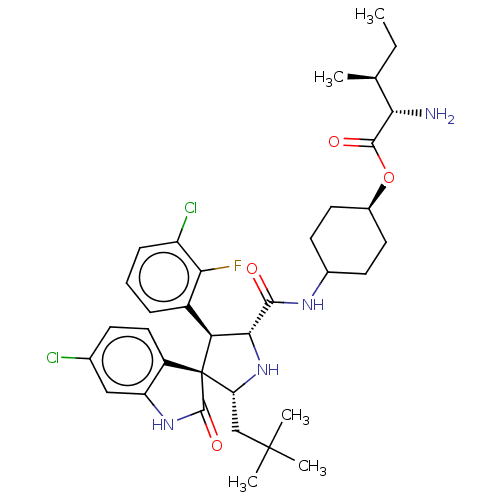
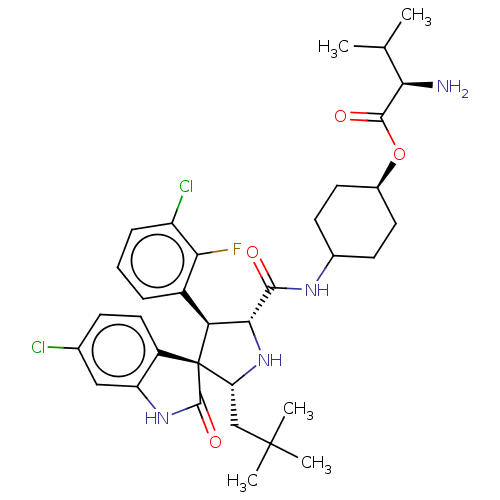
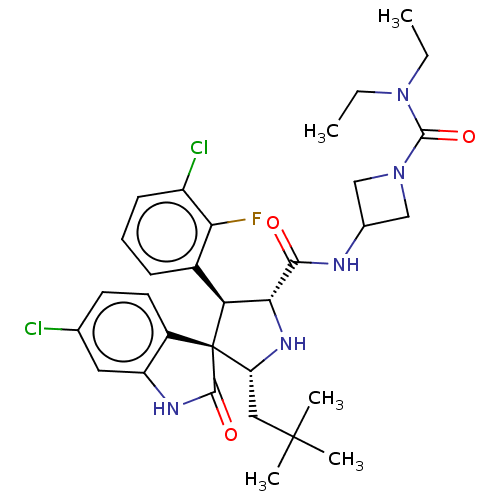
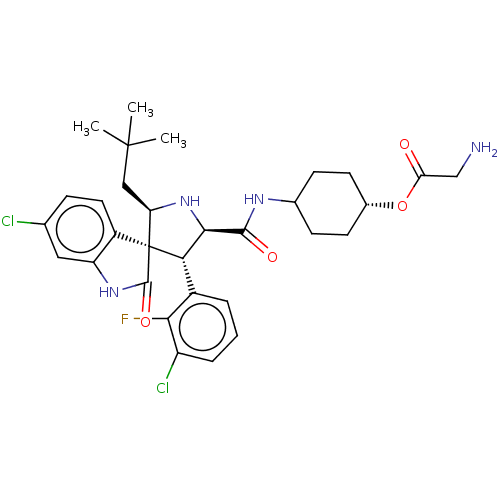
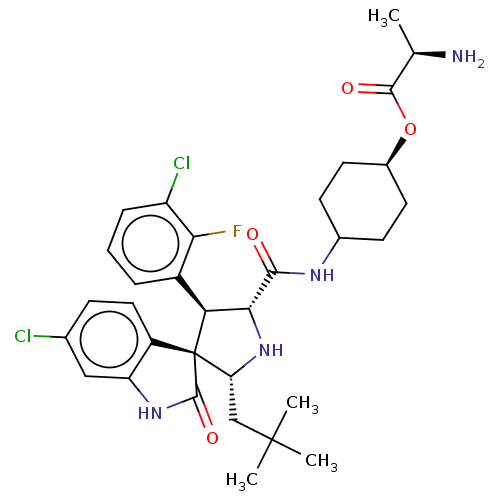
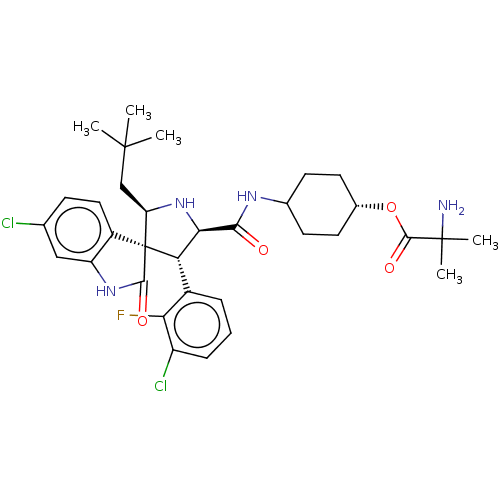
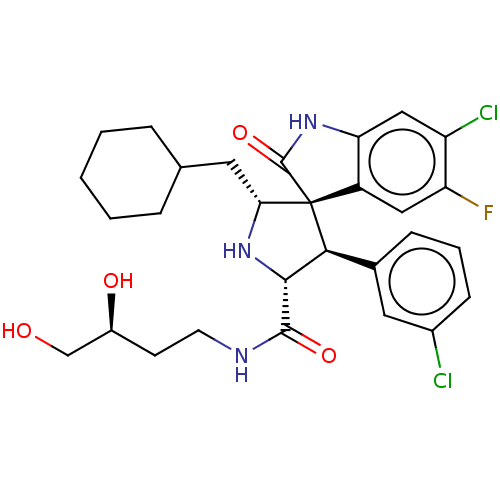
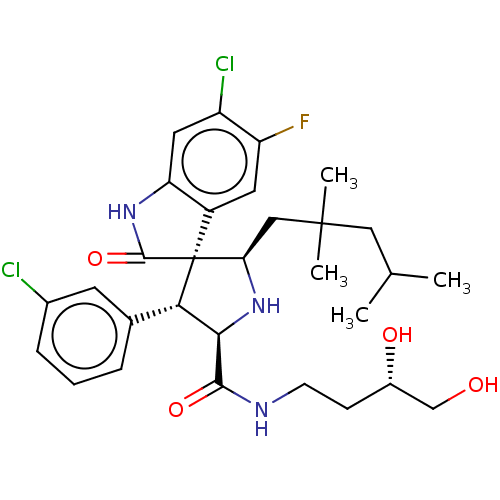
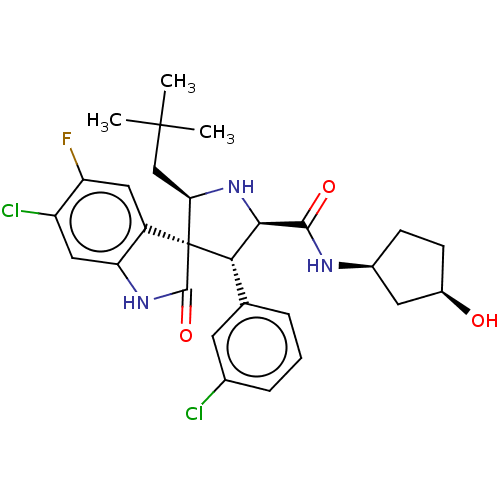
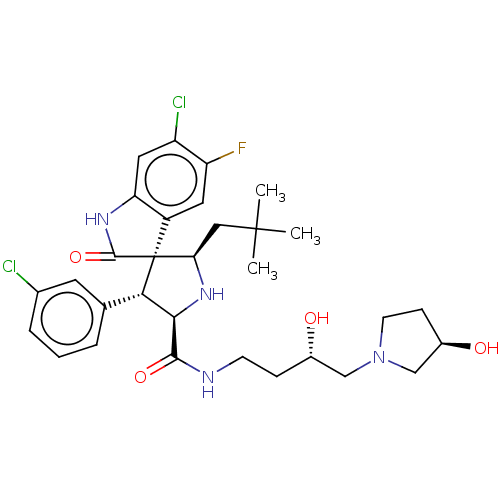
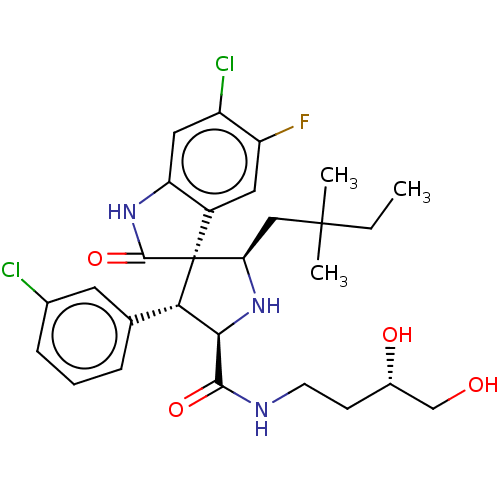
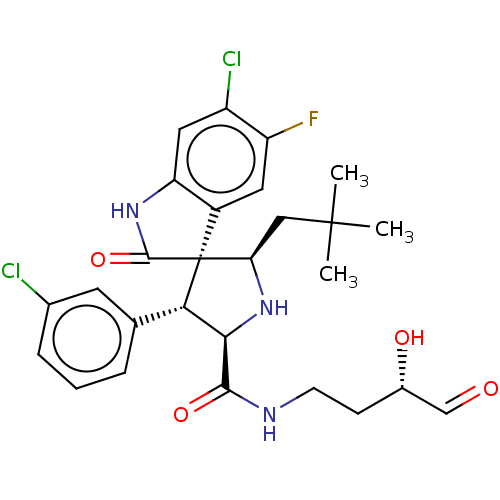
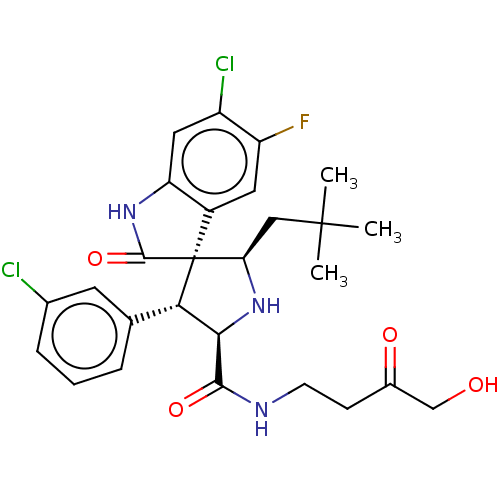
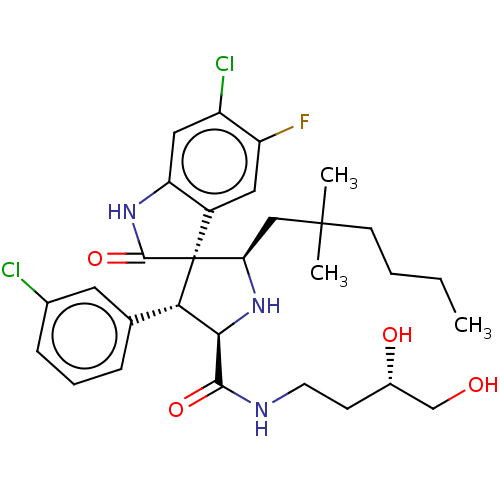
Report error Found 185 Enz. Inhib. hit(s) with all data for entry = 7054
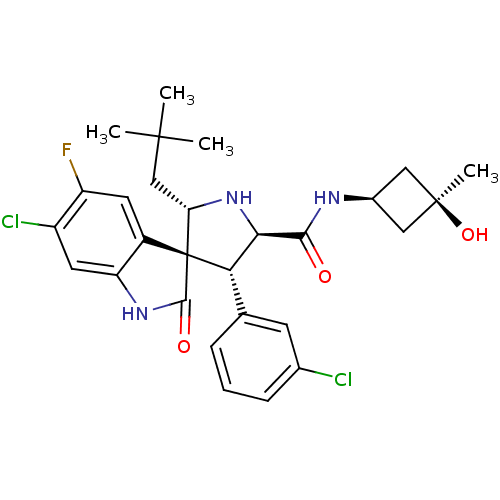
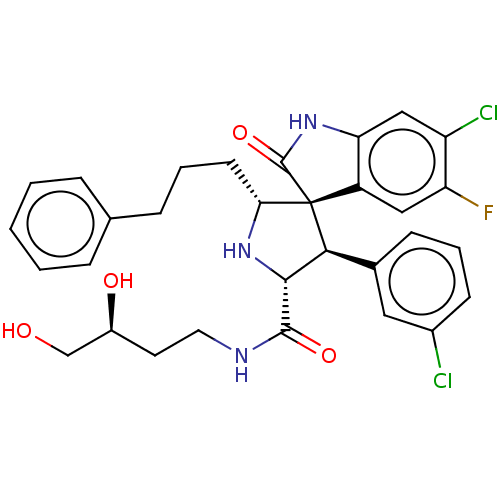
Affinity DataIC50: 100nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
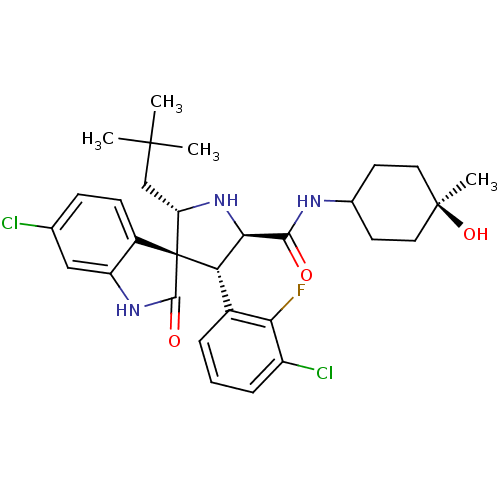
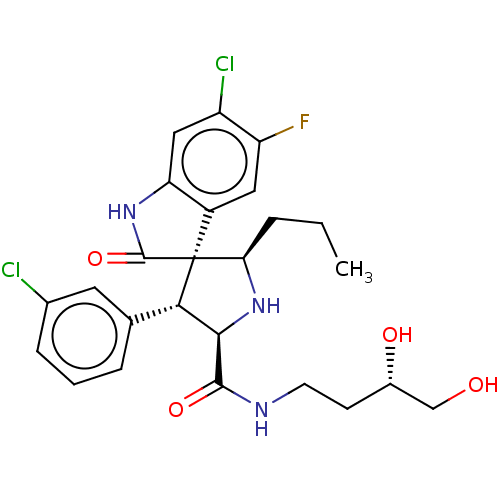
Affinity DataIC50: 100nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
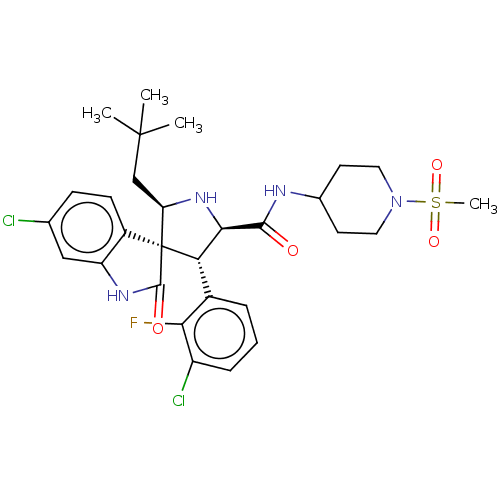
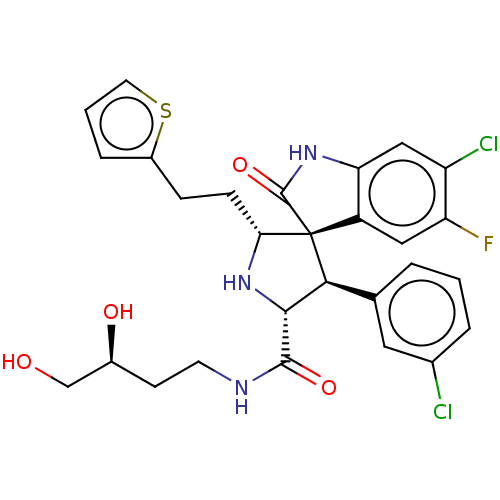
Affinity DataIC50: 100nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
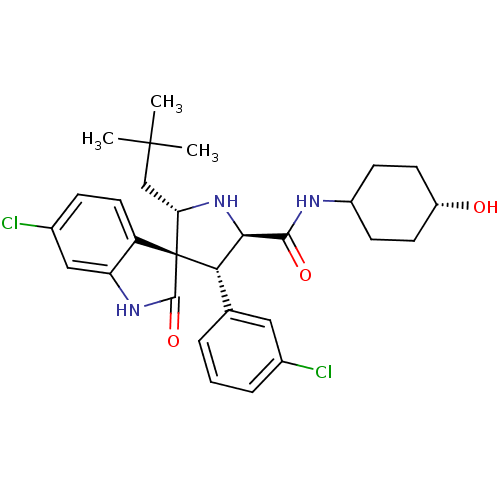
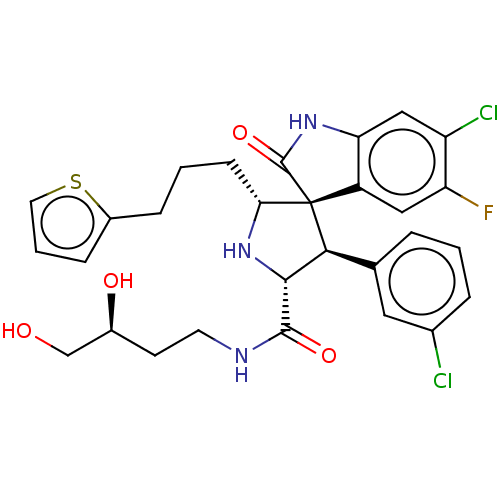
Affinity DataIC50: 100nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 500nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 500nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 500nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 500nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 500nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.5Assay Description:The binding affinity of the MDM2 inhibitors was determined using an optimized, sensitive and quantitative fluorescence polarization-based (FP-based) ...More data for this Ligand-Target Pair