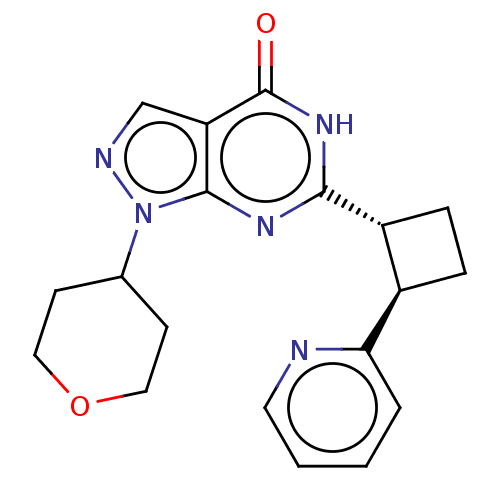
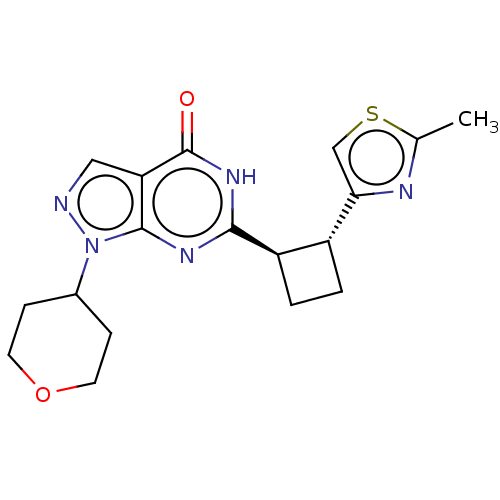
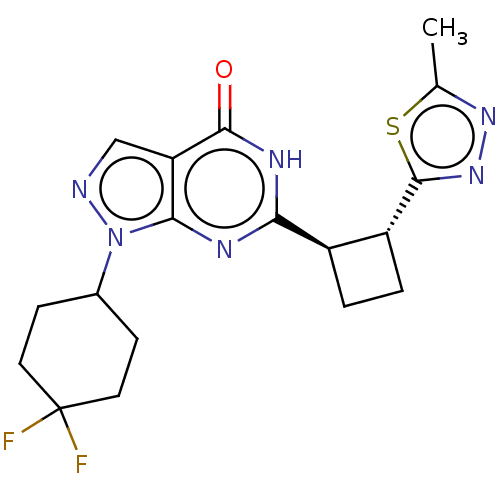
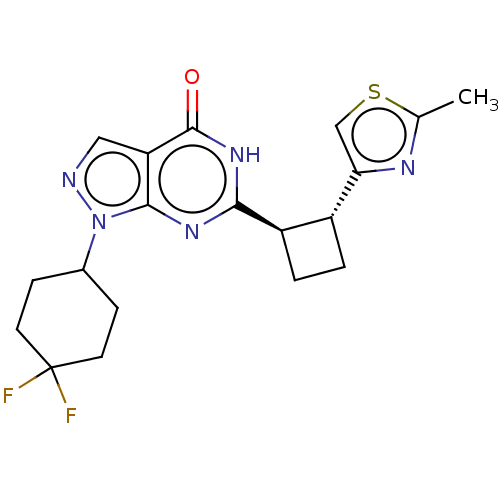
Report error Found 34 Enz. Inhib. hit(s) with all data for entry = 7939
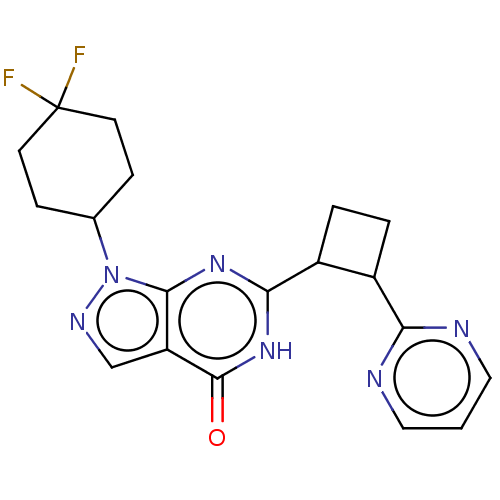
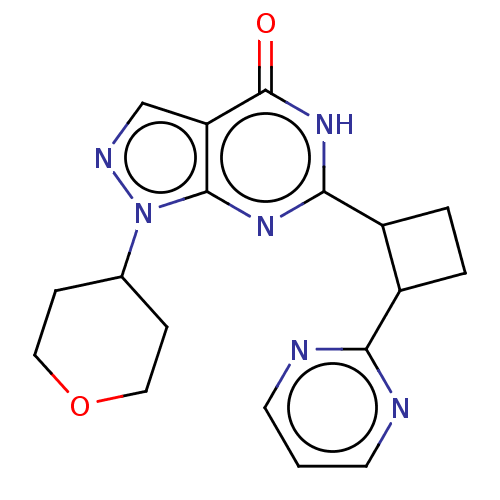
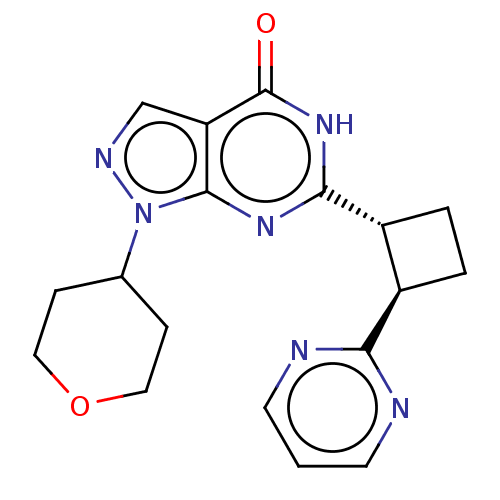
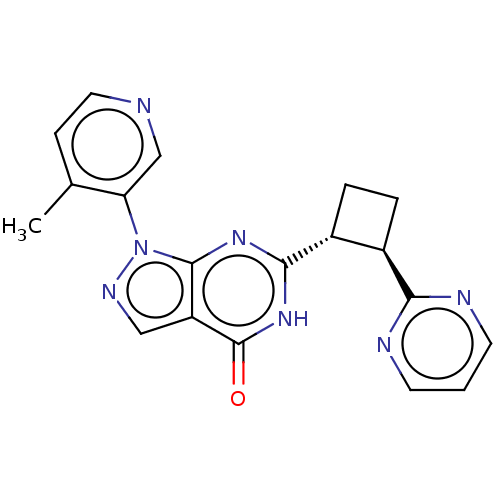
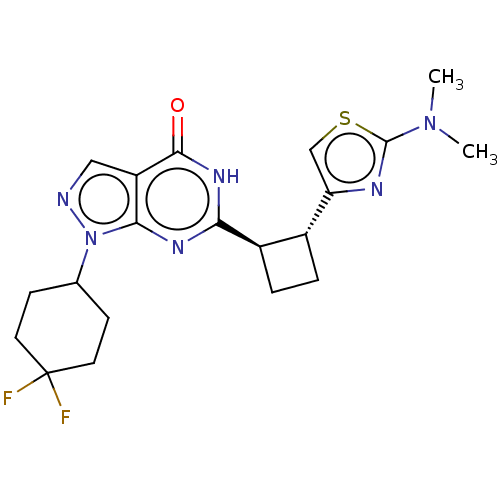
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 4nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
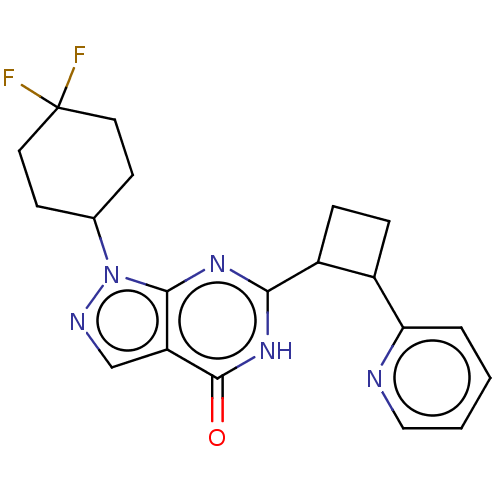
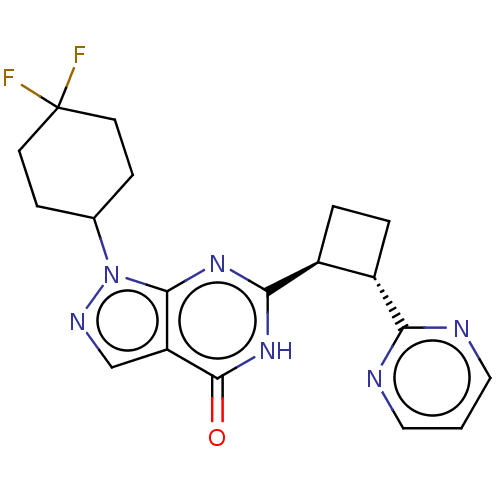
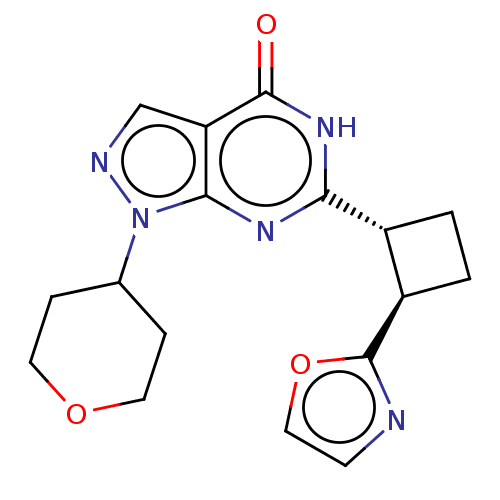
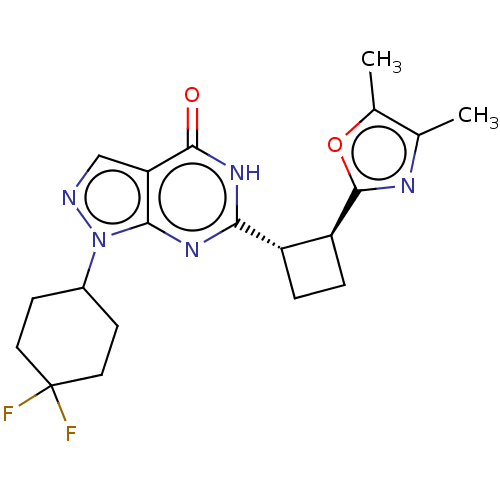
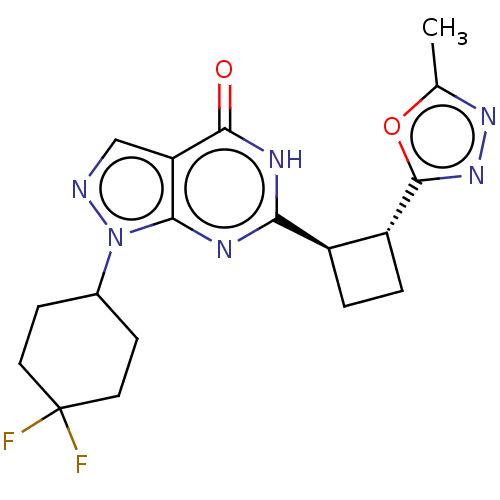
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 4nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
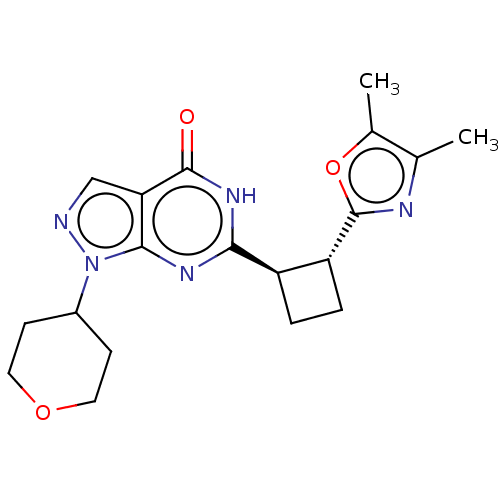
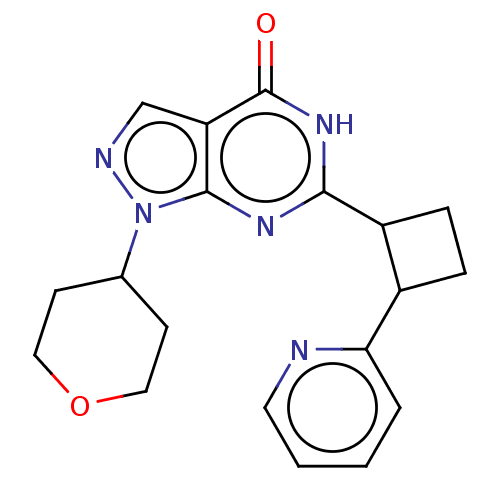
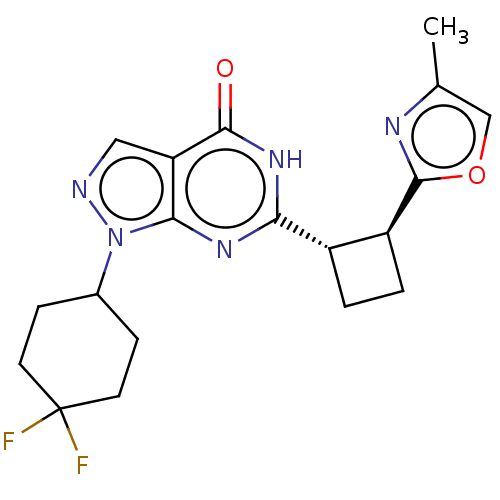
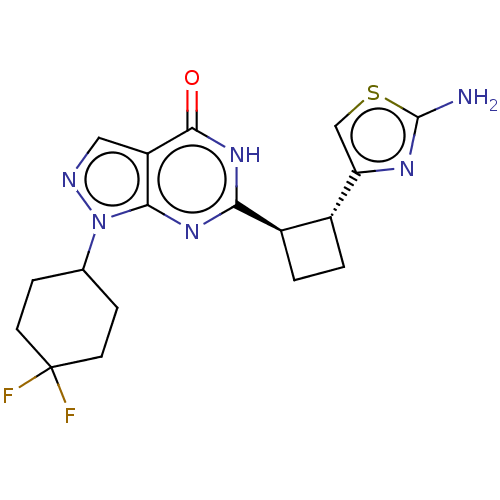
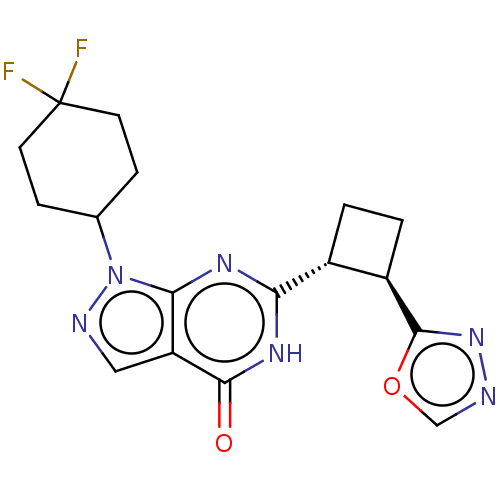
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 5nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
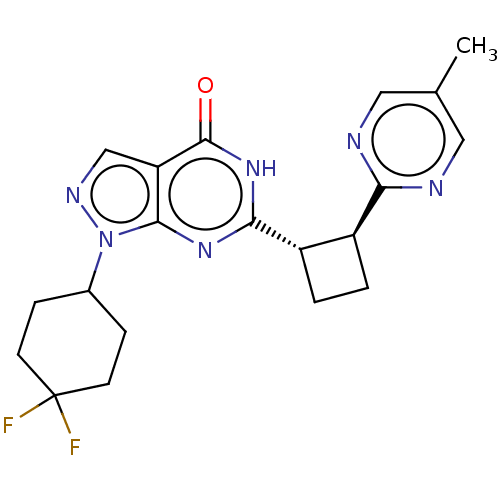
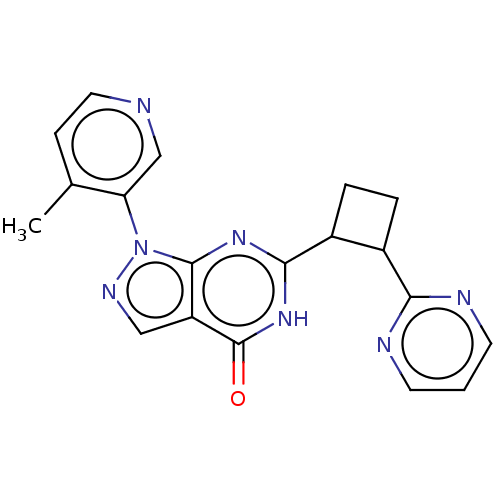
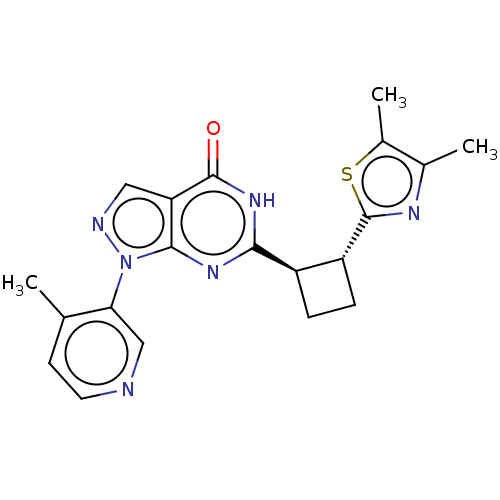
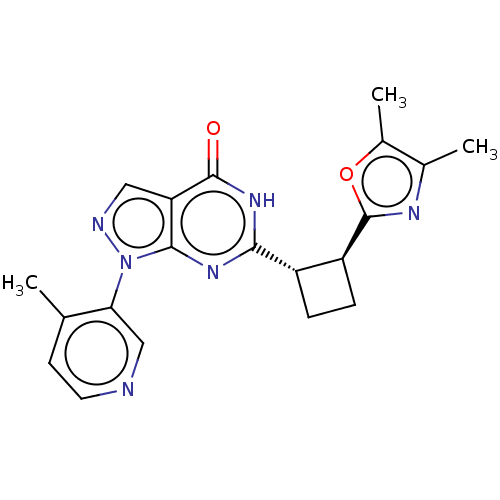
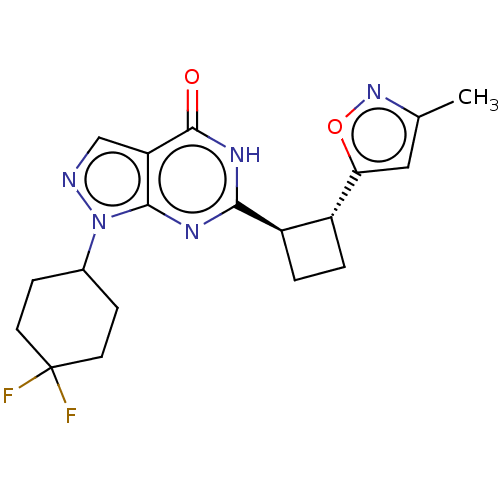
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 5nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 5nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 7nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 7nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 7nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 7nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 11nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 11nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 13nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 16nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 19nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 21nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 23nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 23nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 31nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 31nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 32nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 58nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 60nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 80nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 85nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 218nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 233nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 242nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 304nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 360nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 388nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 433nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 450nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 473nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair
TargetIsoform PDE9A2 of High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (PDE9A2)(Human)
Boehringer Ingelheim International
US Patent
Boehringer Ingelheim International
US Patent
Affinity DataIC50: 1.35E+3nMT: 2°CAssay Description:The PDE9A2 enzymatic activity assay was run as scintillation proximity assay (SPA), in general according to the protocol of the manufacturer (GE Heal...More data for this Ligand-Target Pair