Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Metabotropic glutamate receptor 1
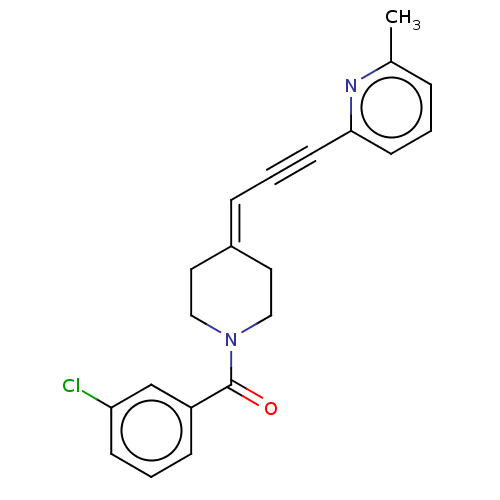
Ligand
BDBM50529980
Substrate
n/a
Meas. Tech.
ChEMBL_1911025 (CHEMBL4413471)
Ki
>1000±n/a nM
Citation
 Graziani, D; Caligari, S; Callegari, E; De Toma, C; Longhi, M; Frigerio, F; Dilernia, R; Menegon, S; Pinzi, L; Pirona, L; Tazzari, V; Valsecchi, AE; Vistoli, G; Rastelli, G; Riva, C Evaluation of Amides, Carbamates, Sulfonamides, and Ureas of 4-Prop-2-ynylidenecycloalkylamine as Potent, Selective, and Bioavailable Negative Allosteric Modulators of Metabotropic Glutamate Receptor 5. J Med Chem 62:1246-1273 (2019) [PubMed] Article
Graziani, D; Caligari, S; Callegari, E; De Toma, C; Longhi, M; Frigerio, F; Dilernia, R; Menegon, S; Pinzi, L; Pirona, L; Tazzari, V; Valsecchi, AE; Vistoli, G; Rastelli, G; Riva, C Evaluation of Amides, Carbamates, Sulfonamides, and Ureas of 4-Prop-2-ynylidenecycloalkylamine as Potent, Selective, and Bioavailable Negative Allosteric Modulators of Metabotropic Glutamate Receptor 5. J Med Chem 62:1246-1273 (2019) [PubMed] Article More Info.:
Target
Name:
Metabotropic glutamate receptor 1
Synonyms:
GRM1_RAT | Gprc1a | Grm1 | Metabotropic Glutamate 1a | Metabotropic glutamate receptor | Metabotropic glutamate receptor 1 | Mglur1 | metabotropic glutamate 1 | metabotropic glutamate 1/5-D | metabotropic glutamate 1/DA | metabotropic glutamate receptor 1 isoform alpha precursor
Type:
Enzyme Catalytic Domain
Mol. Mass.:
133240.47
Organism:
RAT
Description:
metabotropic glutamate 1/2 0 RAT::P23385
Residue:
1199
Sequence:
MVRLLLIFFPMIFLEMSILPRMPDRKVLLAGASSQRSVARMDGDVIIGALFSVHHQPPAEKVPERKCGEIREQYGIQRVEAMFHTLDKINADPVLLPNITLGSEIRDSCWHSSVALEQSIEFIRDSLISIRDEKDGLNRCLPDGQTLPPGRTKKPIAGVIGPGSSSVAIQVQNLLQLFDIPQIAYSATSIDLSDKTLYKYFLRVVPSDTLQARAMLDIVKRYNWTYVSAVHTEGNYGESGMDAFKELAAQEGLCIAHSDKIYSNAGEKSFDRLLRKLRERLPKARVVVCFCEGMTVRGLLSAMRRLGVVGEFSLIGSDGWADRDEVIEGYEVEANGGITIKLQSPEVRSFDDYFLKLRLDTNTRNPWFPEFWQHRFQCRLPGHLLENPNFKKVCTGNESLEENYVQDSKMGFVINAIYAMAHGLQNMHHALCPGHVGLCDAMKPIDGRKLLDFLIKSSFVGVSGEEVWFDEKGDAPGRYDIMNLQYTEANRYDYVHVGTWHEGVLNIDDYKIQMNKSGMVRSVCSEPCLKGQIKVIRKGEVSCCWICTACKENEFVQDEFTCRACDLGWWPNAELTGCEPIPVRYLEWSDIESIIAIAFSCLGILVTLFVTLIFVLYRDTPVVKSSSRELCYIILAGIFLGYVCPFTLIAKPTTTSCYLQRLLVGLSSAMCYSALVTKTNRIARILAGSKKKICTRKPRFMSAWAQVIIASILISVQLTLVVTLIIMEPPMPILSYPSIKEVYLICNTSNLGVVAPVGYNGLLIMSCTYYAFKTRNVPANFNEAKYIAFTMYTTCIIWLAFVPIYFGSNYKIITTCFAVSLSVTVALGCMFTPKMYIIIAKPERNVRSAFTTSDVVRMHVGDGKLPCRSNTFLNIFRRKKPGAGNANSNGKSVSWSEPGGRQAPKGQHVWQRLSVHVKTNETACNQTAVIKPLTKSYQGSGKSLTFSDASTKTLYNVEEEDNTPSAHFSPPSSPSMVVHRRGPPVATTPPLPPHLTAEETPLFLADSVIPKGLPPPLPQQQPQQPPPQQPPQQPKSLMDQLQGVVTNFGSGIPDFHAVLAGPGTPGNSLRSLYPPPPPPQHLQMLPLHLSTFQEESISPPGEDIDDDSERFKLLQEFVYEREGNTEEDELEEEEDLPTASKLTPEDSPALTPPSPFRDSVASGSSVPSSPVSESVLCTPPNVTYASVILRDYKQSSSTL