Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
DNA gyrase subunit A/B
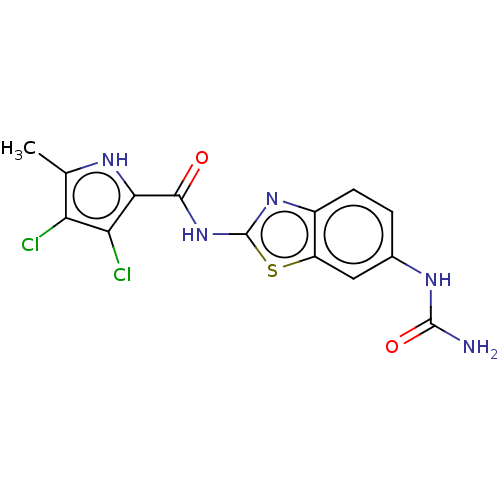
Ligand
BDBM50547863
Substrate
n/a
Meas. Tech.
ChEMBL_2019364 (CHEMBL4672942)
IC50
1300±n/a nM
Citation
 Skok, ?; Baran?okov�, M; Benek, O; Cruz, CD; Tammela, P; Toma?i?, T; Zidar, N; Ma?i?, LP; Zega, A; Stevenson, CEM; Mundy, JEA; Lawson, DM; Maxwell, A; Kikelj, D; Ila?, J Exploring the Chemical Space of Benzothiazole-Based DNA Gyrase B Inhibitors. ACS Med Chem Lett 11:2433-2440 (2020) [PubMed] Article
Skok, ?; Baran?okov�, M; Benek, O; Cruz, CD; Tammela, P; Toma?i?, T; Zidar, N; Ma?i?, LP; Zega, A; Stevenson, CEM; Mundy, JEA; Lawson, DM; Maxwell, A; Kikelj, D; Ila?, J Exploring the Chemical Space of Benzothiazole-Based DNA Gyrase B Inhibitors. ACS Med Chem Lett 11:2433-2440 (2020) [PubMed] Article More Info.:
Target
Name:
DNA gyrase subunit A/B
Synonyms:
DNA Gyrase
Type:
A2B2 tetramer
Mol. Mass.:
n/a
Description:
n/a
Components:
This complex has 2 components.
Component 1
Name:
DNA gyrase subunit A
Synonyms:
DNA gyrase subunit A (gyrA) | GYRA_STAAU | gyrA
Type:
Enzyme Subunit
Mol. Mass.:
99588.82
Organism:
Staphylococcus aureus
Description:
n/a
Residue:
889
Sequence:
MAELPQSRINERNITSEMRESFLDYAMSVIVARALPDVRDGLKPVHRRILYGLNEQGMTPDKSYKKSARIVGDVMGKYHPHGDSSIYEAMVRMAQDFSYRYPLVDGQGNFGSMDGDGAAAMRYTEARMTKITLELLRDINKDTIDFIDNYDGNEREPSVLPARFPNLLANGASGIAVGMATNIPPHNLTELINGVLSLSKNPDISIAELMEDIEGPDFPTAGLILGKSGIRRAYETGRGSIQMRSRAVIEERGGGRQRIVVTEIPFQVNKARMIEKIAELVRDKKIDGITDLRDETSLRTGVRVVIDVRKDANASVILNNLYKQTPLQTSFGVNMIALVNGRPKLINLKEALVHYLEHQKTVVRRRTQYNLRKAKDRAHILEGLRIALDHIDEIISTIRESDTDKVAMESLQQRFKLSEKQAQAILDMRLRRLTGLERDKIEAEYNELLNYISELEAILADEEVLLQLVRDELTEIRDRFGDDRRTEIQLGGFEDLEDEDLIPEEQIVITLSHNNYIKRLPVSTYRAQNRGGRGVQGMNTLEEDFVSQLVTLSTHDHVLFFTNKGRVYKLKGYEVPELSRQSKGIPVVNAIELENDEVISTMIAVKDLESEDNFLVFATKRGVVKRSALSNFSRINRNGKIAISFREDDELIAVRLTSGQEDILIGTSHASLIRFPESTLRPLGRTATGVKGITLREGDEVVGLDVAHANSVDEVLVVTENGYGKRTPVNDYRLSNRGGKGIKTATITERNGNVVCITTVTGEEDLMIVTNAGVIIRLDVADISQNGRAAQGVRLIRLGDDQFVSTVAKVKEDAEDETNEDEQSTSTVSEDGTEQQREAVVNDETPGNAIHTEVIDSEENDEDGRIEVRQDFMDRVEEDIQQSLDEDEE
Component 2
Name:
DNA gyrase subunit B
Synonyms:
DNA gyrase | DNA gyrase subunit B (DNA gyraseB) | DNA gyrase subunit B (gyrB) | GYRB_STAAU | gyrB
Type:
Enzyme Subunit
Mol. Mass.:
72530.91
Organism:
Staphylococcus aureus
Description:
n/a
Residue:
644
Sequence:
MVTALSDVNNTDNYGAGQIQVLEGLEAVRKRPGMYIGSTSERGLHHLVWEIVDNSIDEALAGYANQIEVVIEKDNWIKVTDNGRGIPVDIQEKMGRPAVEVILTVLHAGGKFGGGGYKVSGGLHGVGSSVVNALSQDLEVYVHRNETIYHQAYKKGVPQFDLKEVGTTDKTGTVIRFKADGEIFTETTVYNYETLQQRIRELAFLNKGIQITLRDERDEENVREDSYHYEGGIKSYVELLNENKEPIHDEPIYIHQSKDDIEVEIAIQYNSGYATNLLTYANNIHTYEGGTHEDGFKRALTRVLNSYGLSSKIMKEEKDRLSGEDTREGMTAIISIKHGDPQFEGQTKTKLGNSEVRQVVDKLFSEHFERFLYENPQVARTVVEKGIMAARARVAAKKAREVTRRKSALDVASLPGKLADCSSKSPEECEIFLVEGDSAGGSTKSGRDSRTQAILPLRGKILNVEKARLDRILNNNEIRQMITAFGTGIGGDFDLAKARYHKIVIMTDADVDGAHIRTLLLTFFYRFMRPLIEAGYVYIAQPPLYKLTQGKQKYYVYNDRELDKLKSELNPTPKWSIARYKGLGEMNADQLWETTMNPEHRALLQVKLEDAIEADQTFEMLMGDVVENRRQFIEDNAVYANLDF