Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Glycogen phosphorylase, liver form
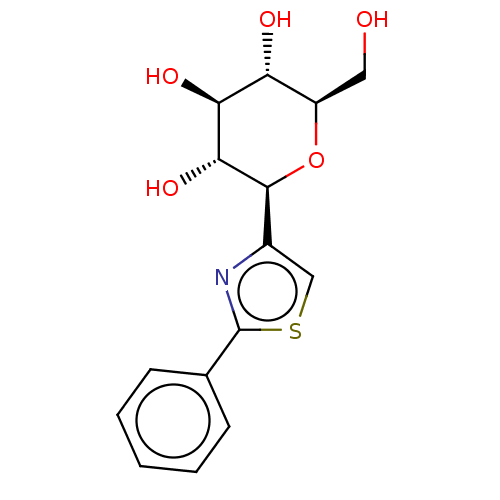
Ligand
BDBM50538947
Substrate
n/a
Meas. Tech.
ChEMBL_1977444 (CHEMBL4610579)
Ki
172200±n/a nM
Citation
 Kyriakis, E; Karra, AG; Papaioannou, O; Solovou, T; Skamnaki, VT; Liggri, PGV; Zographos, SE; Szennyes, E; Bokor, �; Kun, S; Psarra, AG; Soms�k, L; Leonidas, DD The architecture of hydrogen and sulfur ?-hole interactions explain differences in the inhibitory potency of C-?-d-glucopyranosyl thiazoles, imidazoles and an N-?-d glucopyranosyl tetrazole for human liver glycogen phosphorylase and offer new insights to structure-based design. Bioorg Med Chem 28:0 (2020) [PubMed] Article
Kyriakis, E; Karra, AG; Papaioannou, O; Solovou, T; Skamnaki, VT; Liggri, PGV; Zographos, SE; Szennyes, E; Bokor, �; Kun, S; Psarra, AG; Soms�k, L; Leonidas, DD The architecture of hydrogen and sulfur ?-hole interactions explain differences in the inhibitory potency of C-?-d-glucopyranosyl thiazoles, imidazoles and an N-?-d glucopyranosyl tetrazole for human liver glycogen phosphorylase and offer new insights to structure-based design. Bioorg Med Chem 28:0 (2020) [PubMed] Article More Info.:
Target
Name:
Glycogen phosphorylase, liver form
Synonyms:
Glycogen Phosphorylase (PYGL) | Glycogen Phosphorylase, liver form | Liver glycogen phosphorylase | PYGL | PYGL_HUMAN
Type:
Homodimer
Mol. Mass.:
97153.98
Organism:
Human
Description:
Dimers associate into a tetramer to form the enzymatically active phosphorylase A.
Residue:
847
Sequence:
MAKPLTDQEKRRQISIRGIVGVENVAELKKSFNRHLHFTLVKDRNVATTRDYYFALAHTVRDHLVGRWIRTQQHYYDKCPKRVYYLSLEFYMGRTLQNTMINLGLQNACDEAIYQLGLDIEELEEIEEDAGLGNGGLGRLAACFLDSMATLGLAAYGYGIRYEYGIFNQKIRDGWQVEEADDWLRYGNPWEKSRPEFMLPVHFYGKVEHTNTGTKWIDTQVVLALPYDTPVPGYMNNTVNTMRLWSARAPNDFNLRDFNVGDYIQAVLDRNLAENISRVLYPNDNFFEGKELRLKQEYFVVAATLQDIIRRFKASKFGSTRGAGTVFDAFPDQVAIQLNDTHPALAIPELMRIFVDIEKLPWSKAWELTQKTFAYTNHTVLPEALERWPVDLVEKLLPRHLEIIYEINQKHLDRIVALFPKDVDRLRRMSLIEEEGSKRINMAHLCIVGSHAVNGVAKIHSDIVKTKVFKDFSELEPDKFQNKTNGITPRRWLLLCNPGLAELIAEKIGEDYVKDLSQLTKLHSFLGDDVFLRELAKVKQENKLKFSQFLETEYKVKINPSSMFDVQVKRIHEYKRQLLNCLHVITMYNRIKKDPKKLFVPRTVIIGGKAAPGYHMAKMIIKLITSVADVVNNDPMVGSKLKVIFLENYRVSLAEKVIPATDLSEQISTAGTEASGTGNMKFMLNGALTIGTMDGANVEMAEEAGEENLFIFGMRIDDVAALDKKGYEAKEYYEALPELKLVIDQIDNGFFSPKQPDLFKDIINMLFYHDRFKVFADYEAYVKCQDKVSQLYMNPKAWNTMVLKNIAASGKFSSDRTIKEYAQNIWNVEPSDLKISLSNESNKVNGN