Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Complement factor B
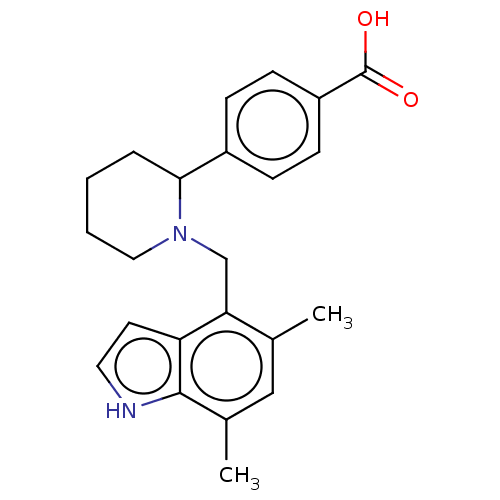
Ligand
BDBM160473
Substrate
n/a
Meas. Tech.
ChEMBL_1981492 (CHEMBL4614754)
IC50
2280±n/a nM
Citation
 Mainolfi, N; Ehara, T; Karki, RG; Anderson, K; Mac Sweeney, A; Liao, SM; Argikar, UA; Jendza, K; Zhang, C; Powers, J; Klosowski, DW; Crowley, M; Kawanami, T; Ding, J; April, M; Forster, C; Serrano-Wu, M; Capparelli, M; Ramqaj, R; Solovay, C; Cumin, F; Smith, TM; Ferrara, L; Lee, W; Long, D; Prentiss, M; De Erkenez, A; Yang, L; Liu, F; Sellner, H; Sirockin, F; Valeur, E; Erbel, P; Ostermeier, D; Ramage, P; Gerhartz, B; Schubart, A; Flohr, S; Gradoux, N; Feifel, R; Vogg, B; Wiesmann, C; Maibaum, J; Eder, J; Sedrani, R; Harrison, RA; Mogi, M; Jaffee, BD; Adams, CM Discovery of 4-((2 S,4 S)-4-Ethoxy-1-((5-methoxy-7-methyl-1 H-indol-4-yl)methyl)piperidin-2-yl)benzoic Acid (LNP023), a Factor B Inhibitor Specifically Designed To Be Applicable to Treating a Diverse Array of Complement Mediated Diseases J Med Chem 63:5697-5722 (2020) [PubMed] Article
Mainolfi, N; Ehara, T; Karki, RG; Anderson, K; Mac Sweeney, A; Liao, SM; Argikar, UA; Jendza, K; Zhang, C; Powers, J; Klosowski, DW; Crowley, M; Kawanami, T; Ding, J; April, M; Forster, C; Serrano-Wu, M; Capparelli, M; Ramqaj, R; Solovay, C; Cumin, F; Smith, TM; Ferrara, L; Lee, W; Long, D; Prentiss, M; De Erkenez, A; Yang, L; Liu, F; Sellner, H; Sirockin, F; Valeur, E; Erbel, P; Ostermeier, D; Ramage, P; Gerhartz, B; Schubart, A; Flohr, S; Gradoux, N; Feifel, R; Vogg, B; Wiesmann, C; Maibaum, J; Eder, J; Sedrani, R; Harrison, RA; Mogi, M; Jaffee, BD; Adams, CM Discovery of 4-((2 S,4 S)-4-Ethoxy-1-((5-methoxy-7-methyl-1 H-indol-4-yl)methyl)piperidin-2-yl)benzoic Acid (LNP023), a Factor B Inhibitor Specifically Designed To Be Applicable to Treating a Diverse Array of Complement Mediated Diseases J Med Chem 63:5697-5722 (2020) [PubMed] Article More Info.:
Target
Name:
Complement factor B
Synonyms:
3.4.21.47 | Bf | C3/C5 convertase | CFAB_MOUSE | Cfb | Complement factor B | Complement factor B Ba fragment | Complement factor B Bb fragment | H2-Bf
Type:
PROTEIN
Mol. Mass.:
85011.61
Organism:
Mouse
Description:
ChEMBL_119901
Residue:
761
Sequence:
MESPQLCLVLLVLGFSSGGVSATPVLEARPQVSCSLEGVEIKGGSFQLLQGGQALEYLCPSGFYPYPVQTRTCRSTGSWSDLQTRDQKIVQKAECRAIRCPRPQDFENGEFWPRSPFYNLSDQISFQCYDGYVLRGSANRTCQENGRWDGQTAICDDGAGYCPNPGIPIGTRKVGSQYRLEDIVTYHCSRGLVLRGSQKRKCQEGGSWSGTEPSCQDSFMYDSPQEVAEAFLSSLTETIEGADAEDGHSPGEQQKRKIVLDPSGSMNIYLVLDGSDSIGSSNFTGAKRCLTNLIEKVASYGVRPRYGLLTYATVPKVLVRVSDERSSDADWVTEKLNQISYEDHKLKSGTNTKRALQAVYSMMSWAGDAPPEGWNRTRHVIIIMTDGLHNMGGNPVTVIQDIRALLDIGRDPKNPREDYLDVYVFGVGPLVDSVNINALASKKDNEHHVFKVKDMEDLENVFYQMIDETKSLSLCGMVWEHKKGNDYHKQPWQAKISVTRPLKGHETCMGAVVSEYFVLTAAHCFMVDDQKHSIKVSVGGQRRDLEIEEVLFHPKYNINGKKAEGIPEFYDYDVALVKLKNKLKYGQTLRPICLPCTEGTTRALRLPQTATCKQHKEQLLPVKDVKALFVSEQGKSLTRKEVYIKNGDKKASCERDATKAQGYEKVKDASEVVTPRFLCTGGVDPYADPNTCKGDSGGPLIVHKRSRFIQVGVISWGVVDVCRDQRRQQLVPSYARDFHINLFQVLPWLKDKLKDEDLGFL