Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Epidermal growth factor receptor [T790M,L858R]
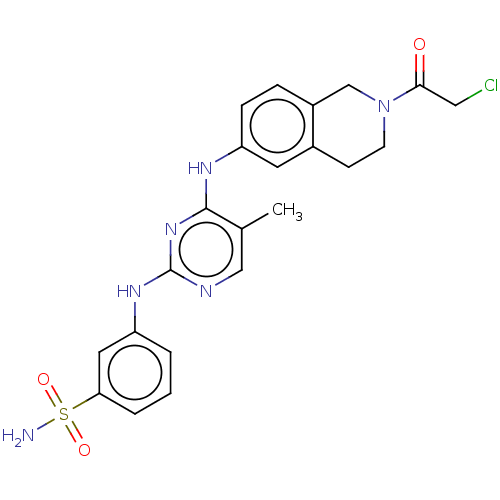
Ligand
BDBM397293
Substrate
n/a
Meas. Tech.
Omnia Assay
IC50
<10±n/a nM
Citation
 Singh, J; Petter, RC; Tester, RW; Kluge, AF; Mazdiyasni, H; Westlin, III, WF; Niu, D; Qiao, L Substituted 2,4-diaminopyrimidines as kinase inhibitors US Patent US9987276 Publication Date 6/5/2018
Singh, J; Petter, RC; Tester, RW; Kluge, AF; Mazdiyasni, H; Westlin, III, WF; Niu, D; Qiao, L Substituted 2,4-diaminopyrimidines as kinase inhibitors US Patent US9987276 Publication Date 6/5/2018 More Info.:
Target
Name:
Epidermal growth factor receptor [T790M,L858R]
Synonyms:
EGFR | EGFR (L858R, T790M) | EGFR_HUMAN | ERBB | ERBB1 | Epidermal growth factor receptor (EGFR) (T790M/L858R) | Epidermal growth factor receptor (EGFR)(T790M/L858R) | Epidermal growth factor receptor (T790M)(L858R) | Epidermal growth factor receptor (T790M, L858R) | Epidermal growth factor receptor (T790M/L858R) | Epidermal growth factor receptor(L858R, T790M) | HER1
Type:
n/a
Mol. Mass.:
134353.72
Organism:
Homo sapiens (Human)
Description:
P00533[T790M,L858R]
Residue:
1210
Sequence:
MRPSGTAGAALLALLAALCPASRALEEKKVCQGTSNKLTQLGTFEDHFLSLQRMFNNCEVVLGNLEITYVQRNYDLSFLKTIQEVAGYVLIALNTVERIPLENLQIIRGNMYYENSYALAVLSNYDANKTGLKELPMRNLQEILHGAVRFSNNPALCNVESIQWRDIVSSDFLSNMSMDFQNHLGSCQKCDPSCPNGSCWGAGEENCQKLTKIICAQQCSGRCRGKSPSDCCHNQCAAGCTGPRESDCLVCRKFRDEATCKDTCPPLMLYNPTTYQMDVNPEGKYSFGATCVKKCPRNYVVTDHGSCVRACGADSYEMEEDGVRKCKKCEGPCRKVCNGIGIGEFKDSLSINATNIKHFKNCTSISGDLHILPVAFRGDSFTHTPPLDPQELDILKTVKEITGFLLIQAWPENRTDLHAFENLEIIRGRTKQHGQFSLAVVSLNITSLGLRSLKEISDGDVIISGNKNLCYANTINWKKLFGTSGQKTKIISNRGENSCKATGQVCHALCSPEGCWGPEPRDCVSCRNVSRGRECVDKCNLLEGEPREFVENSECIQCHPECLPQAMNITCTGRGPDNCIQCAHYIDGPHCVKTCPAGVMGENNTLVWKYADAGHVCHLCHPNCTYGCTGPGLEGCPTNGPKIPSIATGMVGALLLLLVVALGIGLFMRRRHIVRKRTLRRLLQERELVEPLTPSGEAPNQALLRILKETEFKKIKVLGSGAFGTVYKGLWIPEGEKVKIPVAIKELREATSPKANKEILDEAYVMASVDNPHVCRLLGICLTSTVQLIMQLMPFGCLLDYVREHKDNIGSQYLLNWCVQIAKGMNYLEDRRLVHRDLAARNVLVKTPQHVKITDFGRAKLLGAEEKEYHAEGGKVPIKWMALESILHRIYTHQSDVWSYGVTVWELMTFGSKPYDGIPASEISSILEKGERLPQPPICTIDVYMIMVKCWMIDADSRPKFRELIIEFSKMARDPQRYLVIQGDERMHLPSPTDSNFYRALMDEEDMDDVVDADEYLIPQQGFFSSPSTSRTPLLSSLSATSNNSTVACIDRNGLQSCPIKEDSFLQRYSSDPTGALTEDSIDDTFLPVPEYINQSVPKRPAGSVQNPVYHNQPLNPAPSRDPHYQDPHSTAVGNPEYLNTVQPTCVNSTFDSPAHWAQKGSHQISLDNPDYQQDFFPKEAKPNGIFKGSTAENAEYLRVAPQSSEFIGA