Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Dipeptidyl peptidase 4
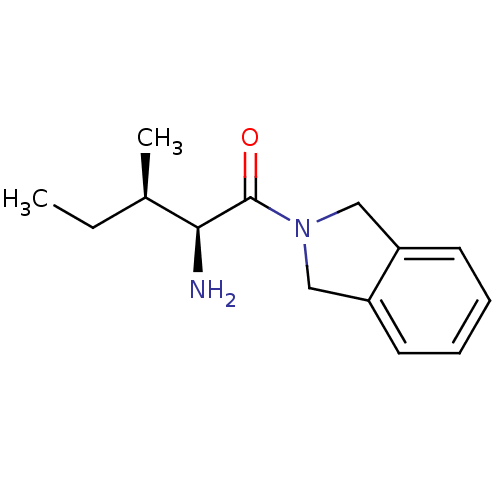
Ligand
BDBM50225437
Substrate
n/a
Meas. Tech.
ChEBML_1717759
IC50
90000±n/a nM
Citation
 Rea, D; Van Elzen, R; De Winter, H; Van Goethem, S; Landuyt, B; Luyten, W; Schoofs, L; Van Der Veken, P; Augustyns, K; De Meester, I; F�l�p, V; Lambeir, AM Crystal structure of Porphyromonas gingivalis dipeptidyl peptidase 4 and structure-activity relationships based on inhibitor profiling. Eur J Med Chem 139:482-491 (2017) [PubMed] Article
Rea, D; Van Elzen, R; De Winter, H; Van Goethem, S; Landuyt, B; Luyten, W; Schoofs, L; Van Der Veken, P; Augustyns, K; De Meester, I; F�l�p, V; Lambeir, AM Crystal structure of Porphyromonas gingivalis dipeptidyl peptidase 4 and structure-activity relationships based on inhibitor profiling. Eur J Med Chem 139:482-491 (2017) [PubMed] Article More Info.:
Target
Name:
Dipeptidyl peptidase 4
Synonyms:
ADABP | ADCP2 | Adenosine deaminase complexing protein 2 | CD26 | CD_antigen=CD26 | DPP IV | DPP4 | DPP4_HUMAN | DPPIV | Dipeptidyl peptidase 4 (DDP-IV) | Dipeptidyl peptidase 4 (DPP IV) | Dipeptidyl peptidase 4 (DPP-4) | Dipeptidyl peptidase 4 (DPP4) | Dipeptidyl peptidase 4 (DPPIV) | Dipeptidyl peptidase 4 membrane form | Dipeptidyl peptidase 4 soluble form | Dipeptidyl peptidase IV | Dipeptidyl peptidase IV (DDP-4) | Dipeptidyl peptidase IV (DDP-IV) | Dipeptidyl peptidase IV (DPP IV) | Dipeptidyl peptidase IV membrane form | Dipeptidyl peptidase IV soluble form | Dipeptidyl peptidase-IV (DPP-4) | Dipeptidyl peptidase-IV (DPP-IV) | T-cell activation antigen CD26 | TP103
Type:
Enzyme
Mol. Mass.:
88271.01
Organism:
Homo sapiens (Human)
Description:
P27487
Residue:
766
Sequence:
MKTPWKVLLGLLGAAALVTIITVPVVLLNKGTDDATADSRKTYTLTDYLKNTYRLKLYSLRWISDHEYLYKQENNILVFNAEYGNSSVFLENSTFDEFGHSINDYSISPDGQFILLEYNYVKQWRHSYTASYDIYDLNKRQLITEERIPNNTQWVTWSPVGHKLAYVWNNDIYVKIEPNLPSYRITWTGKEDIIYNGITDWVYEEEVFSAYSALWWSPNGTFLAYAQFNDTEVPLIEYSFYSDESLQYPKTVRVPYPKAGAVNPTVKFFVVNTDSLSSVTNATSIQITAPASMLIGDHYLCDVTWATQERISLQWLRRIQNYSVMDICDYDESSGRWNCLVARQHIEMSTTGWVGRFRPSEPHFTLDGNSFYKIISNEEGYRHICYFQIDKKDCTFITKGTWEVIGIEALTSDYLYYISNEYKGMPGGRNLYKIQLSDYTKVTCLSCELNPERCQYYSVSFSKEAKYYQLRCSGPGLPLYTLHSSVNDKGLRVLEDNSALDKMLQNVQMPSKKLDFIILNETKFWYQMILPPHFDKSKKYPLLLDVYAGPCSQKADTVFRLNWATYLASTENIIVASFDGRGSGYQGDKIMHAINRRLGTFEVEDQIEAARQFSKMGFVDNKRIAIWGWSYGGYVTSMVLGSGSGVFKCGIAVAPVSRWEYYDSVYTERYMGLPTPEDNLDHYRNSTVMSRAENFKQVEYLLIHGTADDNVHFQQSAQISKALVDVGVDFQAMWYTDEDHGIASSTAHQHIYTHMSHFIKQCFSLP