Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Misshapen-like kinase 1
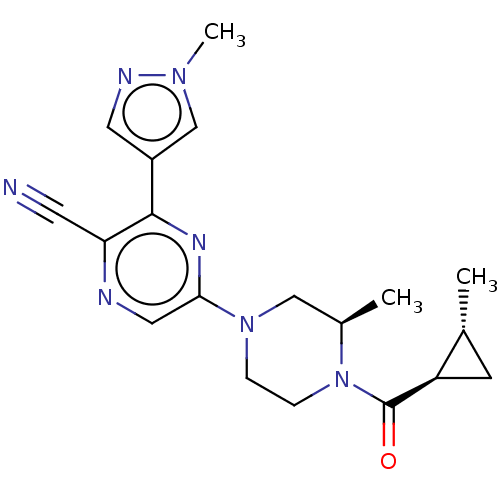
Ligand
BDBM50537742
Substrate
n/a
Meas. Tech.
ChEBML_1970494
IC50
25590±n/a nM
Citation
 Zhou, H; McGowan, MA; Lipford, K; Christopher, M; Fradera, X; Witter, D; Lesburg, CA; Li, C; Methot, JL; Lampe, J; Achab, A; Shaffer, L; Goldenblatt, P; Shah, S; Bass, A; Schroeder, G; Chen, D; Zeng, H; Augustin, MA; Katz, JD Discovery and optimization of heteroaryl piperazines as potent and selective PI3K? inhibitors. Bioorg Med Chem Lett 30:0 (2020) [PubMed] Article
Zhou, H; McGowan, MA; Lipford, K; Christopher, M; Fradera, X; Witter, D; Lesburg, CA; Li, C; Methot, JL; Lampe, J; Achab, A; Shaffer, L; Goldenblatt, P; Shah, S; Bass, A; Schroeder, G; Chen, D; Zeng, H; Augustin, MA; Katz, JD Discovery and optimization of heteroaryl piperazines as potent and selective PI3K? inhibitors. Bioorg Med Chem Lett 30:0 (2020) [PubMed] Article More Info.:
Target
Name:
Misshapen-like kinase 1
Synonyms:
B55 | GCK family kinase MiNK | MAP4K6 | MAPK/ERK kinase kinase kinase 6 | MEK kinase kinase 6 | MEKKK 6 | MINK | MINK1 | MINK1_HUMAN | Misshapen-like kinase 1 (MINK) | Misshapen/NIK-related kinase | Mitogen-activated protein kinase kinase kinase kinase 6 | Voltage-gated potassium channel subunit Kv7.1/Misshapen-like kinase 1 | YSK2 | ZC3
Type:
PROTEIN
Mol. Mass.:
149838.92
Organism:
Homo sapiens (Human)
Description:
ChEMBL_793919
Residue:
1332
Sequence:
MGDPAPARSLDDIDLSALRDPAGIFELVEVVGNGTYGQVYKGRHVKTGQLAAIKVMDVTEDEEEEIKQEINMLKKYSHHRNIATYYGAFIKKSPPGNDDQLWLVMEFCGAGSVTDLVKNTKGNALKEDCIAYICREILRGLAHLHAHKVIHRDIKGQNVLLTENAEVKLVDFGVSAQLDRTVGRRNTFIGTPYWMAPEVIACDENPDATYDYRSDIWSLGITAIEMAEGAPPLCDMHPMRALFLIPRNPPPRLKSKKWSKKFIDFIDTCLIKTYLSRPPTEQLLKFPFIRDQPTERQVRIQLKDHIDRSRKKRGEKEETEYEYSGSEEEDDSHGEEGEPSSIMNVPGESTLRREFLRLQQENKSNSEALKQQQQLQQQQQRDPEAHIKHLLHQRQRRIEEQKEERRRVEEQQRREREQRKLQEKEQQRRLEDMQALRREEERRQAEREQEYKRKQLEEQRQSERLQRQLQQEHAYLKSLQQQQQQQQLQKQQQQQLLPGDRKPLYHYGRGMNPADKPAWAREVEERTRMNKQQNSPLAKSKPGSTGPEPPIPQASPGPPGPLSQTPPMQRPVEPQEGPHKSLVAHRVPLKPYAAPVPRSQSLQDQPTRNLAAFPASHDPDPAIPAPTATPSARGAVIRQNSDPTSEGPGPSPNPPAWVRPDNEAPPKVPQRTSSIATALNTSGAGGSRPAQAVRARPRSNSAWQIYLQRRAERGTPKPPGPPAQPPGPPNASSNPDLRRSDPGWERSDSVLPASHGHLPQAGSLERNRVGVSSKPDSSPVLSPGNKAKPDDHRSRPGRPADFVLLKERTLDEAPRPPKKAMDYSSSSEEVESSEDDEEEGEGGPAEGSRDTPGGRSDGDTDSVSTMVVHDVEEITGTQPPYGGGTMVVQRTPEEERNLLHADSNGYTNLPDVVQPSHSPTENSKGQSPPSKDGSGDYQSRGLVKAPGKSSFTMFVDLGIYQPGGSGDSIPITALVGGEGTRLDQLQYDVRKGSVVNVNPTNTRAHSETPEIRKYKKRFNSEILCAALWGVNLLVGTENGLMLLDRSGQGKVYGLIGRRRFQQMDVLEGLNLLITISGKRNKLRVYYLSWLRNKILHNDPEVEKKQGWTTVGDMEGCGHYRVVKYERIKFLVIALKSSVEVYAWAPKPYHKFMAFKSFADLPHRPLLVDLTVEEGQRLKVIYGSSAGFHAVDVDSGNSYDIYIPVHIQSQITPHAIIFLPNTDGMEMLLCYEDEGVYVNTYGRIIKDVVLQWGEMPTSVAYICSNQIMGWGEKAIEIRSVETGHLDGVFMHKRAQRLKFLCERNDKVFFASVRSGGSSQVYFMTLNRNCIMNW